

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
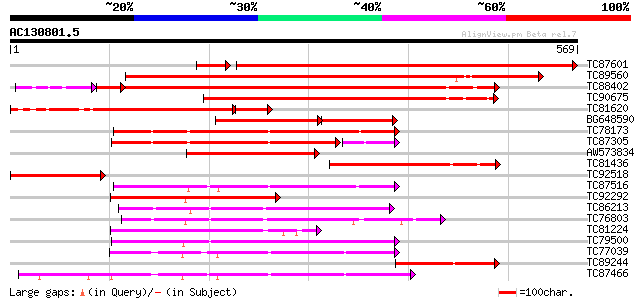
Query= AC130801.5 + phase: 0
(569 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta ... 592 0.0
TC89560 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta ... 626 e-180
TC88402 similar to GP|20804521|dbj|BAB92215. putative CRK1 prote... 478 e-146
TC90675 similar to GP|20804521|dbj|BAB92215. putative CRK1 prote... 351 4e-97
TC81620 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent pr... 232 2e-73
BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 ... 180 4e-71
TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 hom... 218 6e-57
TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control ... 201 7e-55
AW573834 similar to PIR|D86320|D86 hypothetical protein AAF27112... 208 4e-54
TC81436 similar to PIR|A96571|A96571 hypothetical protein F8L10.... 193 2e-49
TC92518 similar to GP|14532508|gb|AAK63982.1 AT3g05050/T12H1_1 {... 191 5e-49
TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2... 181 9e-46
TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase ... 170 2e-42
TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein... 166 2e-41
TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold... 166 2e-41
TC81224 homologue to GP|17064846|gb|AAL32577.1 putative protein ... 164 9e-41
TC79500 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabido... 157 8e-39
TC77039 GP|313148|emb|CAA48472.1| protein kinase {Medicago sativ... 154 7e-38
TC89244 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent pr... 153 2e-37
TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK... 149 4e-36
>TC87601 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta vulgaris},
partial (50%)
Length = 1493
Score = 592 bits (1526), Expect(2) = 0.0
Identities = 283/342 (82%), Positives = 307/342 (89%)
Frame = +2
Query: 228 LHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGV 287
LHRDIKGSNLLIDNEGILKIADFGLASFFDP +PMT+RVVTLWYRPPELLLGATDYGV
Sbjct: 125 LHRDIKGSNLLIDNEGILKIADFGLASFFDPTRRHPMTNRVVTLWYRPPELLLGATDYGV 304
Query: 288 GIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPRE 347
G+DLWSAGCILGELL GKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPRE
Sbjct: 305 GVDLWSAGCILGELLYGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPRE 484
Query: 348 PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYP 407
PYKRCIR+ FK FPPSALPL+D LLAIDP+ER +ASDALRSEFF TEPYACDPSSLPKYP
Sbjct: 485 PYKRCIRDVFKDFPPSALPLVDTLLAIDPIERRSASDALRSEFFNTEPYACDPSSLPKYP 664
Query: 408 PSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLI 467
P+KEMD KRRDDE RR RAA+KA+ DG KKHRTR+R++KA PAPEANAELQ NIDRRRLI
Sbjct: 665 PTKEMDVKRRDDEARRSRAAAKARADGGKKHRTRDRAVKAAPAPEANAELQYNIDRRRLI 844
Query: 468 THANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSGP 527
THANAKSKSEKFPPPH+DGQLGFPLG+S+HIDPD +P D+S S +YT+SKEPFQA+SGP
Sbjct: 845 THANAKSKSEKFPPPHEDGQLGFPLGASNHIDPDIVPHDVSLDSMSYTFSKEPFQAYSGP 1024
Query: 528 IGNAADIGVSKRKKYTAGEAWDLLKPHTGAPKDKFKGKAIIA 569
IGN G +KRKK TA +A DL KP+ G KDK KGK IIA
Sbjct: 1025IGNTTSNGFTKRKKNTANDALDLSKPYKGTHKDKGKGKKIIA 1150
Score = 61.2 bits (147), Expect(2) = 0.0
Identities = 28/34 (82%), Positives = 31/34 (90%)
Frame = +3
Query: 188 MEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEH 221
M HDLAGLAASP I+FTE Q+KCYM+QLLSGLEH
Sbjct: 3 MVHDLAGLAASPDIKFTEPQVKCYMHQLLSGLEH 104
>TC89560 similar to GP|7671528|emb|CAB89490.1 CRK1 protein {Beta vulgaris},
partial (64%)
Length = 1266
Score = 626 bits (1614), Expect = e-180
Identities = 308/422 (72%), Positives = 353/422 (82%), Gaps = 3/422 (0%)
Frame = +2
Query: 117 TYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSR 176
TYSNVYKA D++TGKVVALKKVRFDNLEPES+KFMAREI+ILRRLDHPNV+KL+GLVTSR
Sbjct: 2 TYSNVYKARDTLTGKVVALKKVRFDNLEPESVKFMAREILILRRLDHPNVVKLEGLVTSR 181
Query: 177 MSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSN 236
MSCSLYLVF+YM HDLAGLA +P I+FTE Q+KCYM+QL SGLEHCHNR VLHRDIKGSN
Sbjct: 182 MSCSLYLVFEYMAHDLAGLATNPAIKFTEPQVKCYMHQLFSGLEHCHNRHVLHRDIKGSN 361
Query: 237 LLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGC 296
LLIDN+G+LKIADFGLASFFDP++ +PMTSRVVTLWYRPPELLLGAT+YGVG+DLWSAGC
Sbjct: 362 LLIDNDGVLKIADFGLASFFDPDHKHPMTSRVVTLWYRPPELLLGATEYGVGVDLWSAGC 541
Query: 297 ILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRET 356
IL ELL GKPIMPGRTEVEQLHKI+KLCGSPS++YWKKSKLP+AT+FKP++ YKRCI ET
Sbjct: 542 ILAELLAGKPIMPGRTEVEQLHKIFKLCGSPSEDYWKKSKLPHATIFKPQQSYKRCIAET 721
Query: 357 FKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKR 416
FK FPPS+LPLI+ LLAIDP ER TA+ AL SEFFTT+PYACDPSSLPKYPPSKEMDAK
Sbjct: 722 FKNFPPSSLPLIETLLAIDPDERLTATAALHSEFFTTKPYACDPSSLPKYPPSKEMDAKL 901
Query: 417 RDDEVRRQRAASKAQVDGSKKHRTRERSMKA---MPAPEANAELQSNIDRRRLITHANAK 473
RD+E RR RAA ++ ++ + S + P N + Q ID LITHANAK
Sbjct: 902 RDEEARRLRAAGRSNA*WCQEITST*ASKEGDFPFQMPMLNCK-QILID-GGLITHANAK 1075
Query: 474 SKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSGPIGNAAD 533
SKSEKFPPPHQDG LG+PLGSS H+DP +D+ F+ST + K WSGP+ +
Sbjct: 1076SKSEKFPPPHQDGDLGYPLGSSQHMDPVFDASDVPFSSTNLSLPKADIHTWSGPLVDPGS 1255
Query: 534 IG 535
IG
Sbjct: 1256IG 1261
>TC88402 similar to GP|20804521|dbj|BAB92215. putative CRK1 protein {Oryza
sativa (japonica cultivar-group)}, partial (46%)
Length = 2184
Score = 478 bits (1229), Expect(3) = e-146
Identities = 237/383 (61%), Positives = 298/383 (76%), Gaps = 3/383 (0%)
Frame = +1
Query: 112 KIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQG 171
K+G+GTYS VYKA D K+VALK+VRFDNL+PES+KFMAREI ILRRLDHPN+IKL+G
Sbjct: 991 KLGKGTYSTVYKARDVTNQKIVALKRVRFDNLDPESVKFMAREIHILRRLDHPNIIKLEG 1170
Query: 172 LVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRD 231
L+TS S SLYLVF+YMEHDL GLA++P I+F+E Q+KCYM+QLLSGL+HCH+ VLHRD
Sbjct: 1171 LITSETSRSLYLVFEYMEHDLTGLASNPSIKFSEPQLKCYMHQLLSGLDHCHSHGVLHRD 1350
Query: 232 IKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDL 291
IKGSNLLIDN G+LKIADFGLA+ FD + P+TSRVVTLWYRPPELLLGA YGV +DL
Sbjct: 1351 IKGSNLLIDNNGVLKIADFGLANVFDAHLNIPLTSRVVTLWYRPPELLLGANHYGVAVDL 1530
Query: 292 WSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKR 351
WS GCIL EL G+PI+PG+TEVEQLH+I+KLCGSPS++YW K +LP++T+FKP Y+R
Sbjct: 1531 WSTGCILRELYTGRPILPGKTEVEQLHRIFKLCGSPSEDYWLKLRLPHSTVFKPPHHYRR 1710
Query: 352 CIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKE 411
C+ +TFK + +AL LI+ LL++DP R TA+ AL+SEFFT+EP CDPSSLPKYPPSKE
Sbjct: 1711 CVADTFKEYSSTALKLIETLLSVDPSNRGTAAAALKSEFFTSEPLPCDPSSLPKYPPSKE 1890
Query: 412 MDAKRRDDEVRRQRA-ASKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHA 470
+DAK RD+ RRQ A K Q GS +E+ +A + NA+L ++I ++ ++
Sbjct: 1891 IDAKMRDEATRRQGAVGDKEQRSGSAVR--QEKGPRAAVLTKDNADLGASIQQKH---YS 2055
Query: 471 NAKSKSEKFPP--PHQDGQLGFP 491
K++SE P H G G+P
Sbjct: 2056 ITKNRSELSYPHREHVSGTQGYP 2124
Score = 42.7 bits (99), Expect(3) = e-146
Identities = 17/29 (58%), Positives = 24/29 (82%)
Frame = +3
Query: 88 LTAVCGEALTGWIPRKADTFEKIDKIGQG 116
L++ G+A+ GWIPR A+TFE++ KIGQG
Sbjct: 918 LSSGAGDAIKGWIPRSANTFERLHKIGQG 1004
Score = 38.5 bits (88), Expect(3) = e-146
Identities = 27/81 (33%), Positives = 40/81 (49%)
Frame = +2
Query: 7 RQASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKPNP 66
R SS + G R+K AS+ SNG + + + +N +KKK + + +P
Sbjct: 695 RLNSSKRVEGGGDVRVK---ASSNEKKSNGSGQ-LYDDQNGRKKKIE---KNELTVIDHP 853
Query: 67 RLSNPPKHLRGEQVAAGWPSW 87
PK + EQVAAGWP+W
Sbjct: 854 GFGRVPKAIEAEQVAAGWPAW 916
>TC90675 similar to GP|20804521|dbj|BAB92215. putative CRK1 protein {Oryza
sativa (japonica cultivar-group)}, partial (33%)
Length = 1069
Score = 351 bits (901), Expect = 4e-97
Identities = 170/297 (57%), Positives = 230/297 (77%), Gaps = 1/297 (0%)
Frame = +2
Query: 195 LAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLAS 254
L ++P I+F+E QIKCYM QLLSGL+HCH+R +LHRDIKGSNLLIDN+GILKIADFGLA+
Sbjct: 8 LVSAPGIKFSEPQIKCYMKQLLSGLDHCHSRGILHRDIKGSNLLIDNKGILKIADFGLAN 187
Query: 255 FFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEV 314
FFDPN P+TSRVVTLWYRPPELLLG+++YGV +D+WS GCILGEL G+PI+PG+TEV
Sbjct: 188 FFDPNRSAPLTSRVVTLWYRPPELLLGSSNYGVAVDMWSTGCILGELYCGRPILPGKTEV 367
Query: 315 EQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAI 374
EQLH+I+KLCGSPS +YW+K +LP++T F+P Y+ C+++TFK P +A+ L++ LL++
Sbjct: 368 EQLHRIFKLCGSPSVDYWRKLRLPHSTAFRPPHHYRNCVKDTFKEVPSAAVRLMETLLSL 547
Query: 375 DPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRA-ASKAQVD 433
DP R TA+ AL++EFF+ EP ACDPS+LPKYPP+KE+DAK R++ R++ A K Q
Sbjct: 548 DPRLRGTAATALKNEFFSAEPVACDPSTLPKYPPTKEIDAKMREESTRKEGALGGKEQKV 727
Query: 434 GSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGF 490
GS+ +E+ ++ P+ANA S+I ++ N S++E F PH++ GF
Sbjct: 728 GSRVR--QEKEPQSFVLPKANA--VSHISMQQGQHFPNMTSRNESF-NPHREPASGF 883
>TC81620 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent protein
kinase-like {Arabidopsis thaliana}, partial (30%)
Length = 810
Score = 232 bits (592), Expect(2) = 2e-73
Identities = 123/228 (53%), Positives = 160/228 (69%)
Frame = +3
Query: 1 MGCVIGRQASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSR 60
MGC +G+ A G+GA+ R TA+ G N VE++ Q+ ++ + +
Sbjct: 72 MGCALGKPA----GAGARHRR-----RDNTATANGGN--NAVEVQEKQEAEAPTACELPA 218
Query: 61 RSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSN 120
PRL++ L Q + WP WL V G+A+ W PR+A+TFEK+ KIG+GTYSN
Sbjct: 219 PVSLLPRLNS----LAATQQS--WPPWLMEVAGDAIRDWTPRRANTFEKLAKIGKGTYSN 380
Query: 121 VYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCS 180
VYKA D +TGK+VALKKVR DNL+ ES+KFMAREI++LR+LDHPNVIKL+GLVTSR+S S
Sbjct: 381 VYKAKDLVTGKIVALKKVRIDNLDAESVKFMAREILVLRKLDHPNVIKLEGLVTSRISSS 560
Query: 181 LYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVL 228
LYLVF+YMEHDLAGL A ++F+ Q+KCYM QLLSGLEHCH+RR +
Sbjct: 561 LYLVFEYMEHDLAGLIAGLGVKFSLPQVKCYMKQLLSGLEHCHSRRCI 704
Score = 62.4 bits (150), Expect(2) = 2e-73
Identities = 29/37 (78%), Positives = 33/37 (88%)
Frame = +1
Query: 227 VLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNP 263
VLHRDIKGSNLLID+EGILKIADFGLA+F+D +P
Sbjct: 700 VLHRDIKGSNLLIDDEGILKIADFGLATFYDSKQKHP 810
>BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 [imported] -
Arabidopsis thaliana, partial (40%)
Length = 805
Score = 180 bits (457), Expect(2) = 4e-71
Identities = 82/108 (75%), Positives = 94/108 (86%)
Frame = +1
Query: 207 QIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTS 266
QIKCYM QLLSGLEHCH+R V+HRDIKGSNLL++NEGILK+ADFGLA+F + P+TS
Sbjct: 85 QIKCYMKQLLSGLEHCHSRGVMHRDIKGSNLLVNNEGILKVADFGLANFTNSGKKQPLTS 264
Query: 267 RVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEV 314
RVVTLWYRPPELLLG+TDYG +DLWS GC+ ELLVGKPI+ GRTEV
Sbjct: 265 RVVTLWYRPPELLLGSTDYGPSVDLWSVGCVFAELLVGKPILKGRTEV 408
Score = 106 bits (265), Expect(2) = 4e-71
Identities = 48/76 (63%), Positives = 62/76 (81%)
Frame = +2
Query: 314 VEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLA 373
VEQLHKI+KLCGSP DEYWKK++LP+ATLFKP++PY C+RETFK F +++ L+ LL+
Sbjct: 431 VEQLHKIFKLCGSPPDEYWKKTRLPHATLFKPQQPYDSCLRETFKDFSATSVNLLQNLLS 610
Query: 374 IDPVERETASDALRSE 389
I+P +R TAS AL E
Sbjct: 611 IEPNKRGTASSALSLE 658
>TC78173 SP|P24923|CC21_MEDSA Cell division control protein 2 homolog 1 (EC
2.7.1.-) (Fragment). [Alfalfa] {Medicago sativa},
complete
Length = 1268
Score = 218 bits (554), Expect = 6e-57
Identities = 119/291 (40%), Positives = 181/291 (61%), Gaps = 4/291 (1%)
Frame = +2
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
+ +EK++KIG+GTY VYKA D +T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 131 EQYEKVEKIGEGTYGVVYKARDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 307
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + +SP Q+K ++ Q+L G+ +C
Sbjct: 308 RNIVRLQDVVHSDKR--LYLVFEYLDLDLKKHMDSSPEFIKDPRQVKMFLYQMLCGIAYC 481
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 482 HSHRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 658
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E+ +P+ PG +E+++L KI+++ G+P+++ W + LP+
Sbjct: 659 SRHYSTPVDVWSVGCIFAEMANRRPLFPGDSEIDELFKIFRILGTPNEDTWPGVTSLPDF 838
Query: 341 TLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
PR P K + P+ L L++ +L +DP +R TA A+ E+F
Sbjct: 839 KSTFPRWPSKD-LATVVPNLEPAGLDLLNSMLCLDPTKRITARSAVEHEYF 988
>TC87305 homologue to SP|Q05006|CC22_MEDSA Cell division control protein 2
homolog 2 (EC 2.7.1.-). [Alfalfa] {Medicago sativa},
complete
Length = 1378
Score = 201 bits (511), Expect(2) = 7e-55
Identities = 103/233 (44%), Positives = 155/233 (66%), Gaps = 3/233 (1%)
Frame = +2
Query: 103 KADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRL 161
+ + +EK++KIG+GTY VYKA D T + +ALKK+R + E E + A REI +L+ +
Sbjct: 98 RMEQYEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEM 274
Query: 162 DHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLE 220
H N+++LQ +V S LYLVF+Y++ DL + +SP + QIK ++ Q+L G+
Sbjct: 275 QHRNIVRLQDVVHSEKR--LYLVFEYLDLDLKKFMDSSPEFAKDQRQIKMFLYQILCGIA 448
Query: 221 HCHNRRVLHRDIKGSNLLID-NEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELL 279
+CH+ RVLHRD+K NLLID + LK+ADFGLA F + T VVTLWYR PE+L
Sbjct: 449 YCHSHRVLHRDLKPQNLLIDRSSNALKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEIL 625
Query: 280 LGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW 332
LG+ Y +D+WS GCI E++ +P+ PG +E+++L KI+++ G+P++E W
Sbjct: 626 LGSRHYSTPVDVWSVGCIFAEMINQRPLFPGDSEIDELFKIFRITGTPNEETW 784
Score = 31.6 bits (70), Expect(2) = 7e-55
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +3
Query: 335 SKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
+ LP+ P+ P K + P+ L L+ +L +DP R TA AL E+F
Sbjct: 795 TSLPDFKSAFPKWPAKDLATQV-PNLEPAGLDLLSNMLCLDPTRRITARGALEHEYF 962
>AW573834 similar to PIR|D86320|D86 hypothetical protein AAF27112.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 402
Score = 208 bits (530), Expect = 4e-54
Identities = 96/134 (71%), Positives = 113/134 (83%)
Frame = +1
Query: 178 SCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNL 237
+C +YLVF+YME D+ GL + P I FTESQIKCYM QLLSGLEHCH+R V+HRDIKGSNL
Sbjct: 1 ACIIYLVFEYMELDVTGLWSKPEISFTESQIKCYMKQLLSGLEHCHSRGVMHRDIKGSNL 180
Query: 238 LIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCI 297
L++NEGILK+ADFGLA+F + P+TSRVVTLWYRPPELLLG+TDYG +DLWS GC+
Sbjct: 181 LVNNEGILKVADFGLANFTNSGKKQPLTSRVVTLWYRPPELLLGSTDYGPSVDLWSVGCV 360
Query: 298 LGELLVGKPIMPGR 311
ELLVGKPI+ GR
Sbjct: 361 FAELLVGKPIL*GR 402
>TC81436 similar to PIR|A96571|A96571 hypothetical protein F8L10.9
[imported] - Arabidopsis thaliana, partial (34%)
Length = 1060
Score = 193 bits (490), Expect = 2e-49
Identities = 97/172 (56%), Positives = 130/172 (75%), Gaps = 1/172 (0%)
Frame = +1
Query: 322 KLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERET 381
KLCGSPS++YW+KSKLP+AT+FKP++PY+RC+ ETFK FP A+ LI+ LL+IDP +R T
Sbjct: 1 KLCGSPSEDYWRKSKLPHATIFKPQQPYRRCVAETFKEFPAPAIELIETLLSIDPADRGT 180
Query: 382 ASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAA-SKAQVDGSKKHRT 440
++ AL SEFF+T+P CDPSSLPKYPPSKE DAK RD+E RRQ AA SK Q ++
Sbjct: 181 SASALISEFFSTKPLPCDPSSLPKYPPSKEFDAKVRDEEARRQGAAGSKGQRHDPERRGV 360
Query: 441 RERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPL 492
RE +A+PAP+ANAEL ++ +R+ + ++S+SEKF P +D GFP+
Sbjct: 361 RE--SRAVPAPDANAELVVSMQKRQGQNY--SQSRSEKFNPHPEDAGSGFPI 504
>TC92518 similar to GP|14532508|gb|AAK63982.1 AT3g05050/T12H1_1 {Arabidopsis
thaliana}, partial (8%)
Length = 564
Score = 191 bits (486), Expect = 5e-49
Identities = 94/96 (97%), Positives = 94/96 (97%)
Frame = +3
Query: 1 MGCVIGRQASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSR 60
MGCVIGRQ SSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSR
Sbjct: 267 MGCVIGRQTSSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSR 446
Query: 61 RSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEAL 96
RSKPNPRLSNPPKHLRGEQVAAGWPSWLTA CGEAL
Sbjct: 447 RSKPNPRLSNPPKHLRGEQVAAGWPSWLTADCGEAL 554
>TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2MsF -
alfalfa, complete
Length = 1251
Score = 181 bits (458), Expect = 9e-46
Identities = 111/298 (37%), Positives = 166/298 (55%), Gaps = 11/298 (3%)
Frame = +2
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDH- 163
+ FEK++K+G+GTY VY+A + TGK+VALKK R + RE+ ILR L
Sbjct: 104 EAFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDDEGVPPTTLREVSILRMLSRD 283
Query: 164 PNVIKLQGLVTSRMS---CSLYLVFDYMEHDLAGLAASPVIRFTESQI-----KCYMNQL 215
P+V++L + + LYLVF+YM+ DL S R T I K M QL
Sbjct: 284 PHVVRLLDVKQGQNKEGKTVLYLVFEYMDTDLKKFIRS--FRQTGQNIPPPTIKGLMYQL 457
Query: 216 LSGLEHCHNRRVLHRDIKGSNLLIDNEGI-LKIADFGLASFFDPNYMNPMTSRVVTLWYR 274
G+ CH +LHRD+K NLL+D + + LKIAD GLA F + T ++TLWYR
Sbjct: 458 CKGVAFCHGHGILHRDLKPHNLLMDRKTMMLKIADLGLARAFTVP-LKKYTHEILTLWYR 634
Query: 275 PPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK- 333
PE+LLGAT Y + +D+WS CI EL+ + PG +E++QL I++L G+P+++ W
Sbjct: 635 APEVLLGATHYSMAVDMWSVACIFAELVTKTALFPGDSELQQLLHIFRLLGTPNEDVWPG 814
Query: 334 KSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
SKL N + P + + + G + + L+ ++L +P +R +A A+ +F
Sbjct: 815 VSKLMNWHEYPQWGP--QSLSKAVPGLEETGVDLLSQMLQYEPSKRLSAKKAMEHPYF 982
>TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase cdc2MsC -
alfalfa, partial (38%)
Length = 711
Score = 170 bits (430), Expect = 2e-42
Identities = 93/185 (50%), Positives = 121/185 (65%), Gaps = 15/185 (8%)
Frame = +1
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R D FEK+++IG+GTY VY A + TG++VALKK+ + A +I IL++
Sbjct: 157 RSVDCFEKLEQIGEGTYGMVYMAREIETGEIVALKKIPIG**KEXGFPITAIPQIKILKK 336
Query: 161 LDHPNVIKLQGLVTS--------------RMSCSLYLVFDYMEHDLAGLAASPVIRFTES 206
L H NVIKL+ +VTS + +Y+VF+YM+HDL GLA P +RFT
Sbjct: 337 LHHENVIKLKEIVTSPGPEKDDQGRPDGNKYKGGIYMVFEYMDHDLTGLADRPGMRFTVP 516
Query: 207 QIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTS 266
QIKCYM QLL+GL +CH +VLHRDIKGSNLLIDNEG LK+ADFGLA F + +T+
Sbjct: 517 QIKCYMRQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSFSNEHNANLTN 696
Query: 267 RVVTL 271
RV+TL
Sbjct: 697 RVITL 711
>TC86213 homologue to PIR|S60121|S60121 mitogen-activated protein kinase
MMK2 (EC 2.7.1.-) - alfalfa, complete
Length = 1596
Score = 166 bits (421), Expect = 2e-41
Identities = 105/284 (36%), Positives = 158/284 (54%), Gaps = 7/284 (2%)
Frame = +2
Query: 110 IDKIGQGTYSNVYKAIDSMTGKVVALKKV--RFDNLEPESIKFMAREIIILRRLDHPNVI 167
I +G+G Y V A+++ T + VA+KK+ FDN K REI +LR +DH NV+
Sbjct: 359 IRPVGRGAYGIVCAAVNAETREEVAIKKIGNAFDNRI--DAKRTLREIKLLRHMDHENVM 532
Query: 168 KLQGLVTSRMSCS---LYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHN 224
++ ++ + +Y+V + M+ DL + S T+ + ++ QLL GL++ H+
Sbjct: 533 SIKDIIRPPQKENFNDVYIVSELMDTDLHQIIRSNQ-PMTDDHCRYFVYQLLRGLKYVHS 709
Query: 225 RRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATD 284
VLHRD+K SNLL++ LKI DFGLA + + MT VVT WYR PELLL ++
Sbjct: 710 ANVLHRDLKPSNLLLNANCDLKIGDFGLAR--TTSETDFMTEYVVTRWYRAPELLLNCSE 883
Query: 285 YGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFK 344
Y ID+WS GCILGE++ +P+ PGR V QL + +L GSP D + NA +
Sbjct: 884 YTAAIDIWSVGCILGEIVTRQPLFPGRDYVHQLRLVTELIGSPDDASLGFLRSENARRYV 1063
Query: 345 PREPY--KRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
+ P ++ F P A+ L++K+L DP +R +AL
Sbjct: 1064RQLPQYPQQNFSTRFPSMSPGAVDLLEKMLIFDPSKRIRVDEAL 1195
>TC76803 PIR|T09622|T09622 protein kinase MMK4 (EC 2.7.1.-) cold- and
drought-induced - alfalfa, complete
Length = 1709
Score = 166 bits (420), Expect = 2e-41
Identities = 117/343 (34%), Positives = 177/343 (51%), Gaps = 18/343 (5%)
Frame = +2
Query: 113 IGQGTYSNVYKAIDSMTGKVVALKKVR--FDNLEPESIKFMAREIIILRRLDHPNVIKLQ 170
IG+G Y V +++ T ++VA+KK+ FDN K REI +LR LDH NVI L+
Sbjct: 248 IGRGAYGIVCSLLNTETNELVAVKKIANAFDN--HMDAKRTLREIKLLRHLDHENVIGLR 421
Query: 171 GLVTS---RMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRV 227
++ R +Y+ + M+ DL + S ++ + ++ Q+L GL + H+ +
Sbjct: 422 DVIPPPLRREFNDVYITTELMDTDLHQIIRSNQ-NLSDEHCQYFLYQILRGLRYIHSANI 598
Query: 228 LHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNP-MTSRVVTLWYRPPELLLGATDYG 286
+HRD+K SNLL++ LKI DFGLA P + MT VVT WYR PELLL ++DY
Sbjct: 599 IHRDLKPSNLLLNANCDLKIIDFGLAR---PTMESDFMTEYVVTRWYRAPELLLNSSDYT 769
Query: 287 VGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLF--- 343
ID+WS GCI EL+ KP+ PG+ V Q+ + +L G+P+D K +A +
Sbjct: 770 SAIDVWSVGCIFMELMNKKPLFPGKDHVHQMRLLTELLGTPTDADVGLVKNDDARRYIRQ 949
Query: 344 ---KPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF------TTE 394
PR+P R F P A+ L+DK+L IDP R T +AL + E
Sbjct: 950 LPQYPRQPLNR----VFPHVHPLAIDLVDKMLTIDPTRRITVEEALAHPYLEKLHDVADE 1117
Query: 395 PYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKK 437
P +P S + +D ++ + + R+ A + S+K
Sbjct: 1118PICMEPFSFEF--EQQHLDEEQIKEMIYREALALNPEYA*SRK 1240
>TC81224 homologue to GP|17064846|gb|AAL32577.1 putative protein kinase
{Arabidopsis thaliana}, partial (34%)
Length = 868
Score = 164 bits (415), Expect = 9e-41
Identities = 95/221 (42%), Positives = 133/221 (59%), Gaps = 9/221 (4%)
Frame = +3
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRL 161
R D FE+++KI +GTY VY+A D TG+VVALKKV+ + + REI IL
Sbjct: 138 RSVDEFERLNKIDEGTYGVVYRAKDKKTGEVVALKKVKMEKEKEGFPLTSLREINILLSF 317
Query: 162 DHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEH 221
HP ++ ++ +V S+++V +YMEHDL GL S F++S++KC M QLL G+++
Sbjct: 318 HHPYIVDVKEVVVGSSLDSIFMVMEYMEHDLKGLMESMKQPFSQSEVKCLMIQLLEGVKY 497
Query: 222 CHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWY----RPPE 277
H+ VLHRD+K SNLL++N G LKI DFGLA Y +P + ++T W PE
Sbjct: 498 LHDNWVLHRDLKTSNLLLNNRGELKICDFGLAR----QYGSPFETIILTWWLLFGTGAPE 665
Query: 278 LLLGATDYG-----VGIDLWSAGCILGELLVGKPIMPGRTE 313
LLLGA Y VG+ GCI+ ELL +P+ G+TE
Sbjct: 666 LLLGAKQYSNSY*YVGL----LGCIMAELLSKEPLFNGKTE 776
>TC79500 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabidopsis
thaliana}, partial (55%)
Length = 1381
Score = 157 bits (398), Expect = 8e-39
Identities = 104/297 (35%), Positives = 157/297 (52%), Gaps = 8/297 (2%)
Frame = +2
Query: 103 KADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLD 162
+A ++ + IG+G+Y V AID+ TG+ VA+KK+ + REI +LR L
Sbjct: 419 EASQYQVQEIIGKGSYGVVCSAIDTHTGQKVAIKKINHVFEHVSDATRILREIKLLRLLK 598
Query: 163 HPNVIKLQGLV---TSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGL 219
HP++++++ ++ + R +Y+VF+ ME DL + + T + ++ QLL GL
Sbjct: 599 HPDIVEIRHIMLPPSRREFRDIYVVFELMESDLHQVIKANH-DLTPQHHQFFLYQLLRGL 775
Query: 220 EHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA--SFFDPNYMNPMTSRVVTLWYRPPE 277
+ H V HRD+K N+L + + LKI DFGLA SF D T V T WYR PE
Sbjct: 776 KFIHTANVFHRDLKPKNILANADCKLKICDFGLARVSFNDAPSTIFWTDYVATRWYRAPE 955
Query: 278 LLLGA-TDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSK 336
L + Y GID+WS GCI E+L G+P+ PG+ V QL + L G+P E + +
Sbjct: 956 LCGSFFSKYTPGIDIWSIGCIFAEMLTGRPLFPGKNVVHQLDIMTDLLGTPPPESIARIR 1135
Query: 337 LPNAT--LFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
A L R+ + F P AL ++++LLA DP R +A +AL +F
Sbjct: 1136NEKARRYLNSMRKKQPVPFSQKFPNIDPLALRILERLLAFDPKNRPSAEEALSDPYF 1306
>TC77039 GP|313148|emb|CAA48472.1| protein kinase {Medicago sativa},
complete
Length = 1734
Score = 154 bits (390), Expect = 7e-38
Identities = 110/305 (36%), Positives = 160/305 (52%), Gaps = 14/305 (4%)
Frame = +2
Query: 101 PRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRR 160
P++ ++ +G G++ V++A TG+ VA+KKV D ++ RE+ +R
Sbjct: 434 PKQTISYMAERVVGHGSFGVVFQAKCLETGETVAIKKVLQDK------RYKNRELQTMRL 595
Query: 161 LDHPNVIKLQGL---VTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQ--------IK 209
LDHPNV+ L+ T + L LV +Y+ + S VIR +K
Sbjct: 596 LDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETV-----SRVIRHYNKMNQRMPMIYVK 760
Query: 210 CYMNQLLSGLEHCHNR-RVLHRDIKGSNLLID-NEGILKIADFGLASFFDPNYMNPMTSR 267
Y Q+ L + HN V HRDIK NLL++ + LKI DFG A P S
Sbjct: 761 LYSYQICRALAYIHNSIGVCHRDIKPQNLLVNPHTHQLKICDFGSAKVLVKG--EPNISY 934
Query: 268 VVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSP 327
+ + +YR PEL+ GAT+Y ID+WSAGC+LGELL+G+P+ PG + V+QL +I K+ G+P
Sbjct: 935 ICSRYYRAPELIFGATEYTTAIDIWSAGCVLGELLLGQPLFPGESGVDQLVEIIKVLGTP 1114
Query: 328 SDEYWKKSKLPNATLFK-PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
+ E K PN T FK P+ + K PP A+ L+ +LL P R TA +AL
Sbjct: 1115TREEIKCMN-PNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRSTALEAL 1291
Query: 387 RSEFF 391
F+
Sbjct: 1292VHPFY 1306
>TC89244 similar to GP|8953767|dbj|BAA98122.1 cyclin-dependent protein
kinase-like {Arabidopsis thaliana}, partial (19%)
Length = 809
Score = 153 bits (387), Expect = 2e-37
Identities = 73/106 (68%), Positives = 89/106 (83%), Gaps = 2/106 (1%)
Frame = +2
Query: 388 SEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQ--VDGSKKHRTRERSM 445
SEFFTTEPYAC+PSSLPKYPPSKE+D K RD+E RRQRA S VDG+++ R RERS
Sbjct: 11 SEFFTTEPYACEPSSLPKYPPSKELDVKLRDEEARRQRALSGKSNAVDGARQSRARERSY 190
Query: 446 KAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFP 491
A+PAPEANAE+Q+N+DR R++T+ N KSKSEKFPPPH+DG +G+P
Sbjct: 191 -AIPAPEANAEIQTNLDRLRVVTNGNGKSKSEKFPPPHEDGAVGYP 325
>TC87466 homologue to GP|10185114|emb|CAC08564. wound-induced GSK-3-like
protein {Medicago sativa}, complete
Length = 1934
Score = 149 bits (375), Expect = 4e-36
Identities = 124/434 (28%), Positives = 203/434 (46%), Gaps = 36/434 (8%)
Frame = +3
Query: 10 SSNKGSGAQTNRIKVDEAS-----AATTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKP 64
SS+ G + + R+K+D+ + T + + V+ N+ + + V +++
Sbjct: 75 SSDPGGDSNSKRVKLDQETEKAYEGTNTLDRNDREQRVDSSNEATVGTSNEVSAITKTEK 254
Query: 65 NPRLSNPPKHLRG-------------EQVAAGWPSWLTAVCGEALTGWI------PRKAD 105
PK L + + A S G+ +T I P++
Sbjct: 255 KSGFDELPKELHEMKIRDEKSKNNNEKDIEATTVSGNGTETGQIITTTIAGRDGQPKQTI 434
Query: 106 TFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPN 165
++ +G G++ V++A T + VA+KKV D ++ RE+ ++R +DHPN
Sbjct: 435 SYMAERVVGTGSFGVVFQAKCLETNEAVAIKKVLQDK------RYKNRELQVMRMVDHPN 596
Query: 166 VIKLQGL---VTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQ----IKCYMNQLLSG 218
++KL+ T + L LV +Y+ + ++ + IR + ++ Y Q+L G
Sbjct: 597 IVKLKHCFYSTTEKDELYLNLVLEYVPETVYKVSKN-YIRMHQHMPIIHVQLYTYQILRG 773
Query: 219 LEHCHNR-RVLHRDIKGSNLLIDNEGI-LKIADFGLASFFDPNYMNPMTSRVVTLWYRPP 276
L + H V HRDIK NLL++ + LKI DFG A P P S + + +YR P
Sbjct: 774 LNYLHEVIGVCHRDIKPQNLLVNAQTHQLKICDFGSAKMLVPG--EPNISYICSRYYRAP 947
Query: 277 ELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSK 336
EL+ GAT+Y ID+WS GC+L ELL+G+ + PG + V+QL +I K+ G+P+ E +
Sbjct: 948 ELIFGATEYTTAIDMWSVGCVLAELLLGQAMFPGESGVDQLVEIIKVLGTPTREEIRCMN 1127
Query: 337 LPNATLFK-PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFT--T 393
PN FK P+ + K P A+ L+ +LL P R TA A FF
Sbjct: 1128-PNYNEFKFPQIKAHPWHKLFHKRMPSEAVDLVSRLLQYSPHLRCTALAACAHPFFNDLR 1304
Query: 394 EPYACDPSSLPKYP 407
+P A P+ P P
Sbjct: 1305DPNASLPNGQPLPP 1346
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,552,315
Number of Sequences: 36976
Number of extensions: 253116
Number of successful extensions: 2233
Number of sequences better than 10.0: 517
Number of HSP's better than 10.0 without gapping: 1927
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1959
length of query: 569
length of database: 9,014,727
effective HSP length: 101
effective length of query: 468
effective length of database: 5,280,151
effective search space: 2471110668
effective search space used: 2471110668
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130801.5


