

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
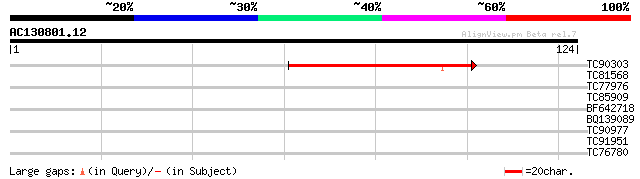
Score E
Sequences producing significant alignments: (bits) Value
TC90303 similar to PIR|T06264|T06264 3-dehydroquinate dehydratas... 45 6e-06
TC81568 similar to GP|13277342|emb|CAC34417. Germin-like protein... 28 0.91
TC77976 similar to GP|7981382|emb|CAB91875.1 putative alcohol de... 28 0.91
TC85909 similar to GP|13277342|emb|CAC34417. Germin-like protein... 27 1.6
BF642718 similar to SP|P51659|DHB4 Estradiol 17 beta-dehydrogena... 26 3.5
BQ139089 homologue to SP|P37114|TCMO Trans-cinnamate 4-monooxyge... 26 3.5
TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {... 26 3.5
TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)... 26 4.5
TC76780 homologue to SP|P37114|TCMO_MEDSA Trans-cinnamate 4-mono... 25 7.7
>TC90303 similar to PIR|T06264|T06264 3-dehydroquinate dehydratase (EC
4.2.1.10) / shikimate 5-dehydrogenase (EC 1.1.1.25) -
tomato, partial (17%)
Length = 632
Score = 45.4 bits (106), Expect = 6e-06
Identities = 23/46 (50%), Positives = 28/46 (60%), Gaps = 5/46 (10%)
Frame = +2
Query: 62 NLVNFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
+L N+ P G ILAN T IGM P D P+++ Y LVFDAVY
Sbjct: 2 DLDNYHPEDGMILANTTSIGMQPKVDETPISKHALKFYSLVFDAVY 139
>TC81568 similar to GP|13277342|emb|CAC34417. Germin-like protein {Pisum
sativum}, partial (96%)
Length = 684
Score = 28.1 bits (61), Expect = 0.91
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +3
Query: 96 LVFDAVYLHFYIKLGGSKAVLFPSF 120
++F LHF I +GGS A+ F SF
Sbjct: 420 MIFPQGLLHFQINVGGSNALTFNSF 494
>TC77976 similar to GP|7981382|emb|CAB91875.1 putative alcohol dehydrogenase
{Lycopersicon esculentum}, partial (48%)
Length = 641
Score = 28.1 bits (61), Expect = 0.91
Identities = 19/56 (33%), Positives = 30/56 (52%), Gaps = 3/56 (5%)
Frame = +3
Query: 8 HPLDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAVFGEVQSY 60
H L+G++ + GGA A AKT GA +VI DI+ + +A ++ + SY
Sbjct: 201 HRLEGKVAIVTGGASGIGAETAKTFVENGAFVVIADINDELGHQVATSIGLDKVSY 368
>TC85909 similar to GP|13277342|emb|CAC34417. Germin-like protein {Pisum
sativum}, partial (95%)
Length = 1551
Score = 27.3 bits (59), Expect = 1.6
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +2
Query: 96 LVFDAVYLHFYIKLGGSKAVLFPSF 120
+VF LHF I GGS A+ F SF
Sbjct: 437 MVFPQGLLHFQINSGGSNALAFVSF 511
>BF642718 similar to SP|P51659|DHB4 Estradiol 17 beta-dehydrogenase 4 (EC
1.1.1.62) (17-beta-HSD 4), partial (24%)
Length = 569
Score = 26.2 bits (56), Expect = 3.5
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +1
Query: 19 GGAGKALAFGAKTRGARIVIFDI 41
GG GKA A +RGA++V+ D+
Sbjct: 55 GGLGKAYALLLASRGAKVVVNDL 123
>BQ139089 homologue to SP|P37114|TCMO Trans-cinnamate 4-monooxygenase (EC
1.14.13.11) (Cinnamic acid 4-hydroxylase) (CA4H) (C4H)
(P450C4H), partial (40%)
Length = 627
Score = 26.2 bits (56), Expect = 3.5
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = +1
Query: 12 GRLFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE 56
G +F+ G + FG++TR V+FDI K + + V+GE
Sbjct: 211 GEMFLHTQG----VEFGSRTRN---VVFDIFTGKGQDMVFTVYGE 324
>TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {Arabidopsis
thaliana}, partial (84%)
Length = 779
Score = 26.2 bits (56), Expect = 3.5
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +3
Query: 80 IGMHPDTDRIPVAEYQLVFDAV 101
+G+HP R V+EYQ +F AV
Sbjct: 315 LGVHPCDKRRSVSEYQFLFPAV 380
>TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)
D22295(C10709) correspond to a region of the predicted
gene.~Similar to, partial (23%)
Length = 526
Score = 25.8 bits (55), Expect = 4.5
Identities = 23/67 (34%), Positives = 29/67 (42%), Gaps = 4/67 (5%)
Frame = -1
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRS----LACAVFGEVQSYKNLVNFQPVKGAI 73
A G A A G GA D D KS+S LACA+ V S + F V +
Sbjct: 187 AAGPPPAGAPGVDLGGA-----DTDRVKSKSFTMLLACAI*SNVPSSSMFLRFGSVSWSR 23
Query: 74 LANATPI 80
+ N TP+
Sbjct: 22 IENLTPV 2
>TC76780 homologue to SP|P37114|TCMO_MEDSA Trans-cinnamate 4-monooxygenase
(EC 1.14.13.11) (Cinnamic acid 4-hydroxylase) (CA4H)
(C4H) (P450C4H), complete
Length = 1871
Score = 25.0 bits (53), Expect = 7.7
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +2
Query: 23 KALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE 56
+ + FG++TR V+FDI K + + V+GE
Sbjct: 431 QGVEFGSRTRN---VVFDIFTGKGQDMVFTVYGE 523
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,522,687
Number of Sequences: 36976
Number of extensions: 36909
Number of successful extensions: 202
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 200
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 201
length of query: 124
length of database: 9,014,727
effective HSP length: 84
effective length of query: 40
effective length of database: 5,908,743
effective search space: 236349720
effective search space used: 236349720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC130801.12


