

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.11 - phase: 0
(275 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
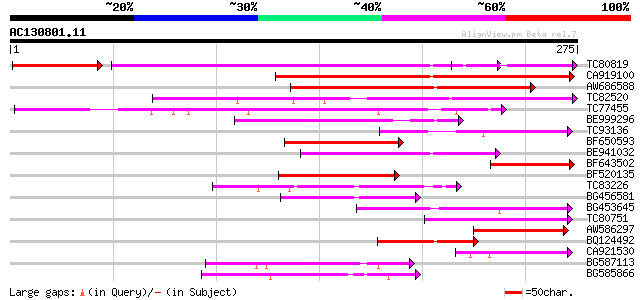
Score E
Sequences producing significant alignments: (bits) Value
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 158 9e-51
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 194 3e-50
AW686588 148 3e-36
TC82520 94 7e-20
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 94 7e-20
BE999296 69 3e-12
TC93136 67 7e-12
BF650593 66 1e-11
BE941032 65 2e-11
BF643502 65 4e-11
BF520135 64 6e-11
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 62 2e-10
BG456581 59 2e-09
BG453645 58 3e-09
TC80751 weakly similar to GP|13161403|dbj|BAB33036. CPRD49 {Vign... 53 1e-07
AW586297 weakly similar to GP|22002156|gb| putative reverse tran... 49 2e-06
BQ124492 weakly similar to GP|9909168|dbj| putative transposable... 49 2e-06
CA921530 49 2e-06
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 47 6e-06
BG585866 45 4e-05
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 158 bits (399), Expect(2) = 9e-51
Identities = 76/190 (40%), Positives = 109/190 (57%)
Frame = +2
Query: 50 ANMLPLRLSRGGEGWSWRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYS 109
A+M L + G W W R L+ WEE + EC LL + L +++D W WL +P+ GYS
Sbjct: 380 ADMGRLGWTVDGRAWEWTRRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNGYS 559
Query: 110 VRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDT 169
V+ Y +T+TG+ S VD +WH +P KVS+F WRL+R+RL T+ NL RG+L
Sbjct: 560 VKVFYRYITSTGHISDRSLVDDVWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATN 739
Query: 170 ALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAK 229
A CV GC E++ HLFL C F LW +R W+G + S + HF+QF + G +
Sbjct: 740 AACVCGCVDSESTTHLFLHCNVFCSLWSLVRNWLG-IPSMSSGELRTHFIQFAKMVGMPR 916
Query: 230 ARRSFLQLIW 239
+ +LI+
Sbjct: 917 VSHLYFRLIF 946
Score = 59.7 bits (143), Expect(2) = 9e-51
Identities = 23/44 (52%), Positives = 31/44 (70%)
Frame = +3
Query: 2 SWFGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDK 45
+WF +++R VGDG D FW+ W G PLR ++PRL DLA+DK
Sbjct: 234 NWFEENIRMVVGDGRDAFFWYDTWAGDVPLRLKYPRLLDLAMDK 365
Score = 45.4 bits (106), Expect = 2e-05
Identities = 27/61 (44%), Positives = 35/61 (57%)
Frame = +1
Query: 215 SNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKLLSL 274
SN F C +G+ SF+ + WVIW ERNNRLF+N VT L+D+IKL S
Sbjct: 871 SNTFYS-VCQNGWHAKSFSFILQV---NFWVIWKERNNRLFQNTVTAPFVLIDRIKLHSF 1038
Query: 275 V 275
+
Sbjct: 1039L 1041
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 194 bits (493), Expect = 3e-50
Identities = 88/145 (60%), Positives = 114/145 (77%)
Frame = -2
Query: 130 DLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSC 189
++IWH QVPLKVS+FAWRL+RDRL T+SNL RG++ + LCV+GCG++E++ HLFLSC
Sbjct: 590 EMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLSC 411
Query: 190 PFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSE 249
+FA LW +R WIG VGVD+N++S+HFVQF +G KA +SFLQLIWLL WV+W+E
Sbjct: 410 SYFASLWSLVRDWIG-FVGVDTNVLSDHFVQFVHSTGGNKASQSFLQLIWLLCAWVLWTE 234
Query: 250 RNNRLFKNVVTEAPRLLDKIKLLSL 274
RNN F + +T PRLLDK+K LSL
Sbjct: 233 RNNMCFNDSITPLPRLLDKVKYLSL 159
>AW686588
Length = 567
Score = 148 bits (373), Expect = 3e-36
Identities = 67/119 (56%), Positives = 87/119 (72%)
Frame = +1
Query: 137 VPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELW 196
VPLKVSI AWRL+RDRL T++NL R L ++ A CV GCG ET++HLFL C F +W
Sbjct: 142 VPLKVSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAVW 321
Query: 197 EQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLF 255
+ +R WIG G D + +S+HF+QF +G +ARRSF+QLIWLL W++W+ERNNRLF
Sbjct: 322 QHIRAWIGVS-GADPHDLSDHFIQFITCTGHTRARRSFMQLIWLLCVWMVWNERNNRLF 495
>TC82520
Length = 833
Score = 93.6 bits (231), Expect = 7e-20
Identities = 60/213 (28%), Positives = 110/213 (51%), Gaps = 7/213 (3%)
Frame = +2
Query: 70 LWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYS---VRGAYEVLTATGNSITD 126
L+AWEE + E LL + L ++ D WL +PI GY+ + ++ + T G +
Sbjct: 2 LFAWEEESVREWYALLHNTVLQENVHDVCRWLLDPI*GYTEGNISLSHYLWTTIG*ESS* 181
Query: 127 SAVDLIWHHQ-VPLKVSIFAWRLMRD---RLSTRSNLASRGILQLDTALCVTGCGSIETS 182
+ + + V ++V+ + + +T S+ S G+ GCG ET+
Sbjct: 182 RCLAKEYSFKGVYVRVASLSQ*TSHEG*FNAATCSSTNSHGLH--------FGCGDSETA 337
Query: 183 DHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLT 242
HLFL C F LW + W+ + + ++I F+QFT ++G + SFLQ++W +
Sbjct: 338 THLFLHCDIFGSLWSHVLRWLHLLLVLPADI-RQFFIQFTSMAGSPRFTHSFLQIMWFAS 514
Query: 243 TWVIWSERNNRLFKNVVTEAPRLLDKIKLLSLV 275
WV+W +RNNR+F+N +++ ++++K+ S +
Sbjct: 515 VWVLWKKRNNRVFQNSLSDPSTFVEQVKMHSFL 613
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 93.6 bits (231), Expect = 7e-20
Identities = 76/261 (29%), Positives = 111/261 (42%), Gaps = 22/261 (8%)
Frame = -2
Query: 3 WFGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGE 62
WF + RKVGDG + FW W PLRE FPR F P RL + G
Sbjct: 727 WFPRELIRKVGDGRSSFFWKDAWDSSVPLRESFPRAF-------------FPYRLLKMGC 587
Query: 63 GWSWR------RWLWAWEEIVL-----EECRTLL--LDVSLFTDISDSWIWLPNPIGGYS 109
G W RW W + L E LL L+ + +D W+W P+ G +S
Sbjct: 586 GDLWDMNAEGVRWRLYWRRLELFEWEKERLLELLGRLEGVVLRYWADIWVWKPDKEGVFS 407
Query: 110 VRGAY------EVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRG 163
V Y +L + + +W + P KV F+W L DR+ T NL R
Sbjct: 406 VNSCYFLLQNLRLLEDRLSYEEEVIFRELWKSKAPAKVLAFSWTLFLDRIPTMVNLGKRR 227
Query: 164 ILQLDTALCVTGCG-SIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIIS--NHFVQ 220
+L+++ + CG ET HLFL C +++ ++ W+ + N+IS N +
Sbjct: 226 LLRVEDSKRCVFCGCQDETVVHLFLHCDVISKV*REVMRWL------NFNLISPPNLLIH 65
Query: 221 FTCLSGFAKARRSFLQLIWLL 241
C S +A++ Q WL+
Sbjct: 64 AICWSREVRAKK-LRQGAWLI 5
>BE999296
Length = 384
Score = 68.6 bits (166), Expect = 3e-12
Identities = 36/111 (32%), Positives = 58/111 (51%)
Frame = -1
Query: 110 VRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDT 169
+ GAY+ L + + +D +WH+ +PLKV F R++R+ L T+ N R ++ +
Sbjct: 306 IHGAYQFLMSADAPLDREYIDSVWHNHIPLKVCFFVLRVLRNCLPTKDNFVRRRVIHEEH 127
Query: 170 ALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQ 220
LC TGC ET+D LF LW + W+ V+S ++ +HF Q
Sbjct: 126 MLCPTGCSFKETTDDLF--------LWPLV*QWLHIS-*VNSCVLRDHFYQ 1
>TC93136
Length = 722
Score = 67.0 bits (162), Expect = 7e-12
Identities = 33/96 (34%), Positives = 52/96 (53%), Gaps = 2/96 (2%)
Frame = +1
Query: 180 ETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFA--KARRSFLQL 237
ETS HLFL CPF + +W ++ W+ ++ +F C+ ++R + L
Sbjct: 262 ETSSHLFLHCPFLSSVWSKILGWL-------------YYSKFVCVFRLLERRSRTKGVWL 402
Query: 238 IWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKLLS 273
IW T WVIW NNR+FKN+ ++++IK+LS
Sbjct: 403 IWHATIWVIWKGINNRIFKNISKAIDEIVEEIKVLS 510
>BF650593
Length = 486
Score = 66.2 bits (160), Expect = 1e-11
Identities = 28/58 (48%), Positives = 36/58 (61%)
Frame = +3
Query: 134 HHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPF 191
H VPLKVS WRL ++ L+TR NL+ RG+L ++ CV CG E+ H F CPF
Sbjct: 297 HKSVPLKVSCLVWRLFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFECPF 470
>BE941032
Length = 435
Score = 65.5 bits (158), Expect = 2e-11
Identities = 34/97 (35%), Positives = 51/97 (52%)
Frame = +2
Query: 142 SIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELWEQLRL 201
SIF W ++ +R T+ NL RG++ CV CG++ + HLFL C FF ++W +
Sbjct: 146 SIFLWCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQIWINVSD 325
Query: 202 WIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLI 238
W+ V V IS+ QF GF+ + +QLI
Sbjct: 326 WLS-FVMVTLLRISDRLAQFGSFGGFSNCKWDTIQLI 433
>BF643502
Length = 631
Score = 64.7 bits (156), Expect = 4e-11
Identities = 29/41 (70%), Positives = 34/41 (82%)
Frame = -1
Query: 234 FLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKLLSL 274
F+QLIWLL WV+W+ERNN +F +VVT P LLDKIKLLSL
Sbjct: 502 FMQLIWLLCVWVVWNERNNVMFNHVVTPIPCLLDKIKLLSL 380
>BF520135
Length = 202
Score = 63.9 bits (154), Expect = 6e-11
Identities = 30/59 (50%), Positives = 38/59 (63%)
Frame = +3
Query: 131 LIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSC 189
L+W + KVSIFAWRL RL T++N+ RGI+ D +CVT C IE+ HLFL C
Sbjct: 12 LLWRKEDLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRCRLIESDVHLFLHC 188
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 62.0 bits (149), Expect = 2e-10
Identities = 43/131 (32%), Positives = 60/131 (44%), Gaps = 10/131 (7%)
Frame = +3
Query: 99 IWLPNPIGGYSVRGAYEVLTA-----TGNSITDSAVDLIWH-----HQVPLKVSIFAWRL 148
+W+ NP G YSV+ Y L N+ T S LIW H +P + + WR+
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIP-RHKVLLWRI 179
Query: 149 MRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVG 208
+ D L RS+L RGI LC ET HLF+SCP +W L C+
Sbjct: 180 LNDSLPVRSSLRKRGIQCY--PLCPRCHSKTETITHLFMSCPLSKRVWFGSNL----CIN 341
Query: 209 VDSNIISNHFV 219
D N+ + +F+
Sbjct: 342 FD-NLPNPNFI 371
>BG456581
Length = 683
Score = 58.9 bits (141), Expect = 2e-09
Identities = 30/68 (44%), Positives = 37/68 (54%)
Frame = +2
Query: 132 IWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPF 191
IW+ +P S WRL DRL T NL+SRG + ++C +ETSDHLFL C F
Sbjct: 80 IWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALV--SMCSFCLEQVETSDHLFLRCKF 253
Query: 192 FAELWEQL 199
LW L
Sbjct: 254 VVTLWSWL 277
>BG453645
Length = 622
Score = 58.2 bits (139), Expect = 3e-09
Identities = 34/107 (31%), Positives = 57/107 (52%), Gaps = 2/107 (1%)
Frame = +3
Query: 169 TALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFA 228
++ CV +ET+ H+FL C + +W ++ W V+ I N FV + C S
Sbjct: 24 SSTCVLCNLKVETTTHIFLHCDVASLVWFRVMKWFD----VNFLIPPNLFVHWECWSEGG 191
Query: 229 KARRSFLQ--LIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKLLS 273
A R L+W T WV+W++RN+ +FK + A ++++IK+LS
Sbjct: 192 SANRVTKGHWLVWHTTIWVLWAKRNDLIFKGLNCVAEDVIEEIKVLS 332
>TC80751 weakly similar to GP|13161403|dbj|BAB33036. CPRD49 {Vigna
unguiculata}, partial (52%)
Length = 1156
Score = 52.8 bits (125), Expect = 1e-07
Identities = 25/72 (34%), Positives = 37/72 (50%)
Frame = +1
Query: 202 WIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTE 261
W D I +H +Q G++++ RS L L+W WVIW ER+ +F
Sbjct: 697 WFLGVSSTDPPCIPDHLIQLVQ*GGYSRSIRSTLLLVWFSYMWVIWKERSKMIFNQKKDH 876
Query: 262 APRLLDKIKLLS 273
+LL+K+KLLS
Sbjct: 877 VHQLLEKVKLLS 912
>AW586297 weakly similar to GP|22002156|gb| putative reverse transcriptase
{Oryza sativa (japonica cultivar-group)}, partial (4%)
Length = 186
Score = 49.3 bits (116), Expect = 2e-06
Identities = 22/46 (47%), Positives = 30/46 (64%)
Frame = +1
Query: 226 GFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKL 271
G AK R S + L W ++WVIW ERN R+F+ + +LL+KIKL
Sbjct: 13 GLAKLRCSIMHLFWFTSSWVIWKERNARIFRAKESTHHQLLEKIKL 150
>BQ124492 weakly similar to GP|9909168|dbj| putative transposable element
Tip100 protein {Oryza sativa (japonica cultivar-group)},
partial (8%)
Length = 694
Score = 48.9 bits (115), Expect = 2e-06
Identities = 22/49 (44%), Positives = 31/49 (62%)
Frame = +2
Query: 179 IETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGF 227
I+T+ HLFL C F+ LW Q+ LW+G V + +HF QFT ++GF
Sbjct: 527 IKTARHLFLDCDIFSSLWSQVWLWLGIS-SVQPGELRHHFPQFTNMAGF 670
>CA921530
Length = 793
Score = 48.9 bits (115), Expect = 2e-06
Identities = 23/62 (37%), Positives = 37/62 (59%), Gaps = 5/62 (8%)
Frame = -1
Query: 217 HFVQFT---CLSGFAKAR--RSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKL 271
HF+ F C +G+ + + R L LIW T W++W NNR+F N EA +++++K+
Sbjct: 670 HFIMFIMSQCWNGWERNKKIRRGLWLIWHATIWILWKAWNNRIFNNQAVEAEDIIEEVKV 491
Query: 272 LS 273
LS
Sbjct: 490 LS 485
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 47.4 bits (111), Expect = 6e-06
Identities = 33/112 (29%), Positives = 49/112 (43%), Gaps = 11/112 (9%)
Frame = -3
Query: 96 DSWIWLPNPIGGYSVRGAYEVLT---ATGNS-------ITDSAVDLIWHHQVPLKVSIFA 145
DS+ W + G YSV+ Y V T A N D +W + KV F
Sbjct: 426 DSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFL 247
Query: 146 WRLMRDRLSTRSNLASRGILQLDTALCVTGCG-SIETSDHLFLSCPFFAELW 196
WR + + L T +N+ SR I + + + CG ET +H+ CP+ +W
Sbjct: 246 WRCISNSLPTAANMRSRHISKDGSC---SRCGMESETVNHILFQCPYARLIW 100
>BG585866
Length = 828
Score = 44.7 bits (104), Expect = 4e-05
Identities = 29/109 (26%), Positives = 52/109 (47%), Gaps = 3/109 (2%)
Frame = +3
Query: 94 ISDSWIWLPNPIGGYSVRGAYEVLTATGNSIT--DSAVDLIWHHQVPLKVSIFAWRLMRD 151
I D++IW N G Y+ + Y + + ++ +S+ IW ++P K F W +
Sbjct: 342 IGDAYIWPHNSNGVYTAKSGYSWILSQTETVNYNNSSWSWIWRLKIPEKYKFFLWLACHN 521
Query: 152 RLSTRSNLASRGILQLDTALCVTGCGSIETS-DHLFLSCPFFAELWEQL 199
+ T S L R + +++A+C + CG E S H C F +W ++
Sbjct: 522 AVPTLSLLNHRNM--VNSAIC-SRCGEHEESFFHCVRDCRFSKIIWHKI 659
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.483
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,023,949
Number of Sequences: 36976
Number of extensions: 210616
Number of successful extensions: 2189
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 2043
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2153
length of query: 275
length of database: 9,014,727
effective HSP length: 95
effective length of query: 180
effective length of database: 5,502,007
effective search space: 990361260
effective search space used: 990361260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC130801.11


