

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.9 + phase: 0
(487 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
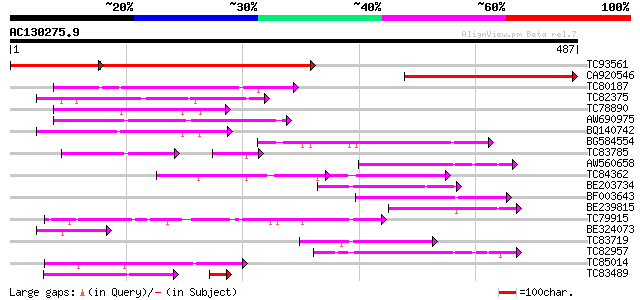
Sequences producing significant alignments: (bits) Value
TC93561 weakly similar to PIR|T45869|T45869 hypothetical protein... 326 e-133
CA920546 301 4e-82
TC80187 similar to PIR|T47858|T47858 hypothetical protein T8B10.... 96 2e-20
TC82375 similar to PIR|T45866|T45866 hypothetical protein F3A4.2... 94 1e-19
TC78890 weakly similar to GP|17978926|gb|AAL47430.1 AT3g47210/F1... 90 2e-18
AW690975 weakly similar to PIR|T04647|T046 hypothetical protein ... 90 2e-18
BQ140742 weakly similar to GP|6957728|gb hypothetical protein {A... 86 3e-17
BG584554 77 1e-14
TC83785 weakly similar to GP|8953391|emb|CAB96664.1 putative pro... 66 1e-14
AW560658 weakly similar to PIR|T45865|T458 hypothetical protein ... 70 2e-12
TC84362 similar to PIR|T45656|T45656 hypothetical protein F13I12... 53 4e-12
BE203734 homologue to GP|19915659|gb cell surface protein {Metha... 60 1e-09
BF003643 weakly similar to PIR|F84686|F846 hypothetical protein ... 59 3e-09
BE239815 58 7e-09
TC79915 similar to GP|17979379|gb|AAL49915.1 unknown protein {Ar... 58 9e-09
BE324073 weakly similar to GP|8953391|emb| putative protein {Ara... 55 6e-08
TC83719 similar to GP|18143763|dbj|BAB79812. hypothetical protei... 53 2e-07
TC82957 weakly similar to GP|6957728|gb|AAF32472.1| hypothetical... 53 3e-07
TC85014 weakly similar to GP|9758661|dbj|BAB09127.1 emb|CAB72473... 51 1e-06
TC83489 weakly similar to GP|17528938|gb|AAL38679.1 unknown prot... 41 2e-06
>TC93561 weakly similar to PIR|T45869|T45869 hypothetical protein F3A4.240 -
Arabidopsis thaliana, partial (9%)
Length = 829
Score = 326 bits (836), Expect(2) = e-133
Identities = 165/184 (89%), Positives = 168/184 (90%)
Frame = +1
Query: 79 QEMEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLV 138
++MEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLV
Sbjct: 277 KKMEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLV 456
Query: 139 DGSFIIQLLRDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSL 198
DGSFIIQLLRDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSL
Sbjct: 457 DGSFIIQLLRDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSL 636
Query: 199 HPQMSFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEEPRGCQ 258
HPQMSFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHV + RGCQ
Sbjct: 637 HPQMSFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHVS*SS*IPT*TETHWRRARGCQ 816
Query: 259 SQMI 262
SQMI
Sbjct: 817 SQMI 828
Score = 168 bits (425), Expect(2) = e-133
Identities = 80/80 (100%), Positives = 80/80 (100%)
Frame = +3
Query: 1 MSFTFQKLIKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNESVCIYRVPPNMLNVEPKA 60
MSFTFQKLIKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNESVCIYRVPPNMLNVEPKA
Sbjct: 42 MSFTFQKLIKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNESVCIYRVPPNMLNVEPKA 221
Query: 61 YIPSNISIGPYHYGSQHLQE 80
YIPSNISIGPYHYGSQHLQE
Sbjct: 222 YIPSNISIGPYHYGSQHLQE 281
>CA920546
Length = 594
Score = 301 bits (771), Expect = 4e-82
Identities = 148/148 (100%), Positives = 148/148 (100%)
Frame = -2
Query: 340 RLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLG 399
RLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLG
Sbjct: 593 RLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLG 414
Query: 400 CWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRS 459
CWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRS
Sbjct: 413 CWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRS 234
Query: 460 LIIIPFRDPPHYTVFDDVSKKIDHGTNN 487
LIIIPFRDPPHYTVFDDVSKKIDHGTNN
Sbjct: 233 LIIIPFRDPPHYTVFDDVSKKIDHGTNN 150
>TC80187 similar to PIR|T47858|T47858 hypothetical protein T8B10.130 -
Arabidopsis thaliana, partial (4%)
Length = 1040
Score = 96.3 bits (238), Expect = 2e-20
Identities = 68/217 (31%), Positives = 116/217 (53%), Gaps = 6/217 (2%)
Frame = +3
Query: 38 EEKNESVCIYRVPPNMLNVEP-KAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPN 96
+ K+ CIY+VPPN+ + +AY P ISIGP+H+ ++ M K ++FH ++
Sbjct: 282 DAKHGICCIYKVPPNLRKLNNGEAYTPYLISIGPFHHTKEN--SMHTQKQRYFHYFWE-R 452
Query: 97 GANGSKLEEACKFLEKEEINARNCYMG-EIKLSSDEFLKMMLVDGSFIIQL-LRDLSDNE 154
N L + FL+ + +N Y + + DEF+ M+++D FI++L LR ++E
Sbjct: 453 MTNKKALVKYKNFLKVKIEVIKNFYFEFDHTIKDDEFVDMIMLDSVFILELFLRKSKESE 632
Query: 155 F-KHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFF- 212
K + W+ I+R++++LENQLP +VL +L++ K+N+ +SF L+F +F
Sbjct: 633 REKDYMFTTSWIYKGIQRDLLLLENQLPFYVLEQLYQKVCKDNN---DLSFLELAFNYFE 803
Query: 213 -YRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPK 248
Y +S+ + E K H DL+R PK
Sbjct: 804 DYNPQRSKKDQNEEMIKYCKKSCRHFTDLVRCFYLPK 914
>TC82375 similar to PIR|T45866|T45866 hypothetical protein F3A4.210 -
Arabidopsis thaliana, partial (37%)
Length = 1103
Score = 94.0 bits (232), Expect = 1e-19
Identities = 65/212 (30%), Positives = 120/212 (55%), Gaps = 12/212 (5%)
Frame = +3
Query: 24 WRGLVEKELQSIKPEEKNES---VCIYRVPPNMLNV--EPKAYIPSNISIGPYHYGSQHL 78
W ++++L+ ++ S + IY++P + + + KA+ P +S+GPYH+G + L
Sbjct: 381 WVISIKEKLEEANQDDVASSWTKLSIYKIPHYLRDSSGDDKAFAPQIVSLGPYHHGKKRL 560
Query: 79 QEMEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLV 138
++ME K + + + + +A K +E++ AR+CY G I LSS+EF++M+++
Sbjct: 561 RQMERHKWRSINHVLKRTKHDIRLYLDAMKEMEEK---ARSCYEGTIGLSSNEFVEMLVL 731
Query: 139 DGSFIIQLLRDLSDNEFKHV------PSLS-RWMLPTIRREMIMLENQLPMFVLTKLFEL 191
DG F+I+L R ++ FK + P + R + +I+R+MIMLENQLP+F+L L +
Sbjct: 732 DGCFVIELFRGATEG-FKELGYSRNDPVFAMRGSMHSIQRDMIMLENQLPLFILDLLLGI 908
Query: 192 TDKNNSLHPQMSFYNLSFKFFYRLLQSESRKT 223
L ++ NL+ +FF L+ ++ T
Sbjct: 909 QIGKPDLKGLVA--NLALRFFDPLMPTDEPLT 998
>TC78890 weakly similar to GP|17978926|gb|AAL47430.1 AT3g47210/F13I12_260
{Arabidopsis thaliana}, partial (6%)
Length = 844
Score = 90.1 bits (222), Expect = 2e-18
Identities = 58/176 (32%), Positives = 95/176 (53%), Gaps = 24/176 (13%)
Frame = +1
Query: 38 EEKNESVC-IYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFD-- 94
EE + + C IY+VP N+ V+ +AY P ISIGP H G++ LQ M+ K ++FH ++
Sbjct: 193 EELSLADCSIYKVPHNLRKVKQEAYTPQLISIGPIHLGNEELQPMQQHKKRYFHFFYERL 372
Query: 95 PNGANGSKLEEACKFLEKEEINARNCYMGEI-KLSSDEFLKMMLVDGSFIIQLL------ 147
S ++ +LE +E R CY + +S ++F++ +L+D FI++LL
Sbjct: 373 SQFNQQSMIKNYKLYLENKEQEIRKCYAEKFPNISKEKFVETILLDAVFIMELLLRNSSW 552
Query: 148 -RDLSDNEFKHVPSLS-------------RWMLPTIRREMIMLENQLPMFVLTKLF 189
D S +E ++ S S W+ +I R++I+LENQ+P FVL L+
Sbjct: 553 KSDTSKHEHEYKQSKSFRWKHSDDYILTQAWLSKSITRDLILLENQIPFFVLKHLY 720
>AW690975 weakly similar to PIR|T04647|T046 hypothetical protein F10N7.210 -
Arabidopsis thaliana, partial (10%)
Length = 657
Score = 90.1 bits (222), Expect = 2e-18
Identities = 63/206 (30%), Positives = 100/206 (47%), Gaps = 1/206 (0%)
Frame = +3
Query: 38 EEKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNG 97
E ++ CIY+VP + AY P+ +SIGP+H+G LQ ME K +F
Sbjct: 42 EPPSDKCCIYKVPYVIRRHNKDAYTPTVVSIGPFHHGHPQLQNMENQKLIYFKDFLQRTK 221
Query: 98 ANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLL-RDLSDNEFK 156
A L + ++E + + CY + S DE +K++L+D +FIIQL R D +
Sbjct: 222 A---CLNDLVCYIESVLSDFKRCYSETLPFSHDELVKLILIDSAFIIQLFWRYYYD---R 383
Query: 157 HVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLL 216
W+ IR ++ +LENQLP FV+ K++ L+ N SF L+ +F+
Sbjct: 384 RCILFKPWLDNGIRYDLWLLENQLPFFVIEKIYSLSLTNVPTTMIHSFLKLTIHYFWLFT 563
Query: 217 QSESRKTPECQTSYKFKIEHVLDLLR 242
+++ + S I H DL+R
Sbjct: 564 INQNWDFDKGDIS----ISHFTDLIR 629
>BQ140742 weakly similar to GP|6957728|gb hypothetical protein {Arabidopsis
thaliana}, partial (10%)
Length = 644
Score = 85.9 bits (211), Expect = 3e-17
Identities = 58/180 (32%), Positives = 87/180 (48%), Gaps = 12/180 (6%)
Frame = +1
Query: 24 WRGLVEKELQSIKPEEKNE-SVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEME 82
W V K L+ EE E S+ I+ VP ++ +P +Y+P ++IGPYHY Q L EM+
Sbjct: 94 WVIQVRKTLEEEFEEENGELSISIFNVPKLLMASDPNSYVPQQVAIGPYHYWRQELFEMQ 273
Query: 83 VLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSF 142
K R K++ L K E R CY + L+ + MM+VD SF
Sbjct: 274 NHKLASTKRF--QKQLQSLKIDNLVDQLTKLEQKIRACYHKFLDLNGETLAWMMIVDASF 447
Query: 143 IIQLL---------RDLSDNEFKHVPSL--SRWMLPTIRREMIMLENQLPMFVLTKLFEL 191
+++ L R + + H + R I R+++MLENQ+P+FVL K+ EL
Sbjct: 448 LLEFLQFYAMQEESRKVISSNMSHFVNYVGRRLSXNAILRDIVMLENQIPLFVLRKMLEL 627
>BG584554
Length = 803
Score = 77.4 bits (189), Expect = 1e-14
Identities = 56/224 (25%), Positives = 96/224 (42%), Gaps = 22/224 (9%)
Frame = +3
Query: 214 RLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLI---GEEPRGC-----QSQMIHSI 265
+L+ + T C++ H DL+RY P+ I G P ++ +
Sbjct: 30 KLISQKLELTQNCKSC-----NHFTDLIRYTYLPREIQVKGVNPSKSFAPFRSEYVLRTA 194
Query: 266 TELKEGGVKIKACESRELMDISFGK----KWGIMV----------KELTIPPLYIGDHRG 311
T+L E G+ + + R D+ F K W + + +L IP L +
Sbjct: 195 TKLSEAGISFEKLQGRSYCDLKFKKTPILSWFLCLGCVPCFKFVESKLQIPHLNVDQATE 374
Query: 312 TVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELI 371
V RN++AFE+CH P + Y+ ++ LI++ DV L +I H LGS +A L+
Sbjct: 375 CVLRNLIAFEQCHYSEQPFICNYVSLIDSLIHTHEDVELLVDTEIISHELGSHAELATLV 554
Query: 372 NNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLT 415
N + K +V + + ++ E NE+ W + L Y +
Sbjct: 555 NGLCKYVV--VTSNCYGKIIKEMNEHYNNWWKHYMGMLRSVYFS 680
>TC83785 weakly similar to GP|8953391|emb|CAB96664.1 putative protein
{Arabidopsis thaliana}, partial (12%)
Length = 692
Score = 66.2 bits (160), Expect(2) = 1e-14
Identities = 35/102 (34%), Positives = 53/102 (51%)
Frame = +1
Query: 45 CIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKLE 104
CIY+VP + + AY P+ +SIGP+H+G LQ ME K F A L
Sbjct: 136 CIYKVPSKIRKLNADAYTPTVVSIGPFHHGHPQLQNMEPQKLIHFKAFLQRTEA---CLN 306
Query: 105 EACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQL 146
+ +++ N + CY + S DE +K++L+D FII+L
Sbjct: 307 DLVCYVDSILSNFKRCYSETLPFSHDELVKLILIDSGFIIEL 432
Score = 31.2 bits (69), Expect(2) = 1e-14
Identities = 17/48 (35%), Positives = 27/48 (55%), Gaps = 4/48 (8%)
Frame = +2
Query: 175 MLENQLPMFVLTKLFELT-DKNNSLHPQM---SFYNLSFKFFYRLLQS 218
+LENQLP FV+ K++ L+ N+ +P F L+ FF+ +S
Sbjct: 506 VLENQLPFFVIEKIYSLSFSSTNASNPNTMIPCFLELTINFFFSFNKS 649
>AW560658 weakly similar to PIR|T45865|T458 hypothetical protein F3A4.200 -
Arabidopsis thaliana, partial (6%)
Length = 589
Score = 69.7 bits (169), Expect = 2e-12
Identities = 48/140 (34%), Positives = 77/140 (54%), Gaps = 3/140 (2%)
Frame = -1
Query: 300 TIPPLYIGDHRGTVFRNIVAFEKCHKRCNP-DMTTYMFFLNRLINSANDVSALHYKGVIH 358
T+P + + D F N +A+E C N + ++ F++ LI++ DV L KG++
Sbjct: 589 TLPEIVVDDTSAATFLNXIAYEMCPDFDNDYGICSFAAFMDSLIDNPEDVKLLRSKGILL 410
Query: 359 HSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRAR-FRASLVHNYLTS- 416
+SLGSDE VAEL N I+ ++VP+ +E+Y + Y C R + A H Y ++
Sbjct: 409 NSLGSDEEVAELFNIISTDLVPN-SETYFEVRAKIHDHY--CNRCNTWIAEGFHTYFSNP 239
Query: 417 WAVGLSTLGALLALYFTFIQ 436
WA+ ++ + A AL TFIQ
Sbjct: 238 WAI-IAFIAAFTALVLTFIQ 182
>TC84362 similar to PIR|T45656|T45656 hypothetical protein F13I12.250 -
Arabidopsis thaliana, partial (5%)
Length = 942
Score = 53.1 bits (126), Expect(2) = 4e-12
Identities = 45/157 (28%), Positives = 73/157 (45%), Gaps = 8/157 (5%)
Frame = +2
Query: 127 LSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHVPS--LSRWMLPTIRREMIMLENQLPMFV 184
L + ++++L+D FII+L +++ L W+ IR ++++LENQLP FV
Sbjct: 140 LLNRSLVQLILIDSGFIIELFWRSYYDDWSQDDGFLLKPWLASNIRLDLLLLENQLPFFV 319
Query: 185 LTKLFELT-DKNNSLHPQM---SFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHVLDL 240
+ +++ L+ N+ P+ SF LSF +F QS + F I H DL
Sbjct: 320 IQEIYNLSFTSTNASVPKTKIPSFLELSFDYFAYYNQSNLG-----FDNGDFSIRHFTDL 484
Query: 241 LRYNIRPKLIGEEPRGCQSQMIH--SITELKEGGVKI 275
+R + P M H S TEL E G ++
Sbjct: 485 IRIFHLQHPLQRRPLRIDEPMKHLQSATELLEAGGEV 595
Score = 35.8 bits (81), Expect(2) = 4e-12
Identities = 29/108 (26%), Positives = 47/108 (42%), Gaps = 1/108 (0%)
Frame = +3
Query: 272 GVKIKACESRE-LMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRCNPD 330
GV+ K E L+D+ F + L IP L + D +FRN+VA E+CH
Sbjct: 585 GVRFKVNTKSECLLDLRFSGR------VLEIPQLKVEDWTEILFRNMVALEQCHYPSESY 746
Query: 331 MTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEI 378
+T Y+ + ++ + LG + VA L N++ K +
Sbjct: 747 ITDYVCCFGFPY*YWQGCGYTCSEEILVNWLGDSDSVANLFNSLWKNV 890
>BE203734 homologue to GP|19915659|gb cell surface protein {Methanosarcina
acetivorans str. C2A} [Methanosarcina acetivorans C2A],
partial (0%)
Length = 571
Score = 60.5 bits (145), Expect = 1e-09
Identities = 37/125 (29%), Positives = 67/125 (53%), Gaps = 1/125 (0%)
Frame = +3
Query: 265 ITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCH 324
I +L+ G+++KA + D+ F W +LTIP +Y+ + + N++A+E C
Sbjct: 159 IEDLRAVGIRLKASATGRPTDVDFTAGW--FTAKLTIPVIYVNNLAASTLLNLIAYEMCP 332
Query: 325 KRCNP-DMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMN 383
N + +Y+ ++ LI+ DV L KG++ ++ SDE VA L N I ++V ++
Sbjct: 333 DFDNDCGICSYVALMDSLIDHPEDVKVLRSKGIL-CTVWSDEEVASLFNIIGTDLVANI- 506
Query: 384 ESYLY 388
+ Y Y
Sbjct: 507 DKYFY 521
>BF003643 weakly similar to PIR|F84686|F846 hypothetical protein At2g28580
[imported] - Arabidopsis thaliana, partial (11%)
Length = 545
Score = 59.3 bits (142), Expect = 3e-09
Identities = 37/134 (27%), Positives = 59/134 (43%)
Frame = +3
Query: 298 ELTIPPLYIGDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVI 357
+L IP + V RN++A E+CH P + Y+ ++ LIN+ DV L K +I
Sbjct: 12 KLQIPQFKVHQTTECVLRNLIALEQCHYSNQPFICNYVSLIDFLINTQEDVELLVDKEII 191
Query: 358 HHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSW 417
H LGS +A +IN + K +V N Y N + Y W+ + W
Sbjct: 192 VHELGSHAELATMINGLCKNVVVTCN-YYGKTSKNLNDHYYNYWKHYMGMLRSVYFRDPW 368
Query: 418 AVGLSTLGALLALY 431
+ +G + L+
Sbjct: 369 RFSSTIVGVFIFLF 410
>BE239815
Length = 586
Score = 58.2 bits (139), Expect = 7e-09
Identities = 36/126 (28%), Positives = 61/126 (47%), Gaps = 12/126 (9%)
Frame = +1
Query: 326 RCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDM--- 382
R N + ++ F++ L+++ +DV L GV + LGSDE +A+L N++ ++ M
Sbjct: 13 RYNYECCSFFTFMDSLVDNGDDVKELRLPGVFQNLLGSDEQLAKLFNDLGDDLPTKMYCN 192
Query: 383 ---------NESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFT 433
+ YL V Y W+ + ++ T WA+ ++ L A+LAL T
Sbjct: 193 NSYTDAVAYSRRYLLIKVQIEKHYTNKWKTWLAQAYNTHFNTPWAM-IAFLAAMLALVLT 369
Query: 434 FIQAIC 439
FIQ C
Sbjct: 370 FIQTWC 387
>TC79915 similar to GP|17979379|gb|AAL49915.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 1090
Score = 57.8 bits (138), Expect = 9e-09
Identities = 78/329 (23%), Positives = 138/329 (41%), Gaps = 36/329 (10%)
Frame = +1
Query: 31 ELQSIKPEEKNESVCIYRVP---PNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNK 87
ELQS K +N I RVP M N+ K Y P +SIGP H+ L+ E K K
Sbjct: 106 ELQS-KQTPQNSRPKIQRVPYYRRRMSNLG-KHYTPKLLSIGPIHHDRSDLKLGEKYKLK 279
Query: 88 FFHRLFDPNGANGSKLEEACKFLEK-EEINARNCYMGEIKLSSDEFLK-----------M 135
+ + N +L + K + +E+ R + ++ + + L+ M
Sbjct: 280 WAAEYIENTALNPEELHQ--KIADNIDELKGR--FSDDVLTLTGKSLEGFHSLEEKLSWM 447
Query: 136 MLVDGSFIIQLLRDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKN 195
+ VDG ++ LL K + L + ++++LENQLP VL L+ KN
Sbjct: 448 LFVDGCSLLYLLE-------KGHMKIKVGQLVLVMMDVLLLENQLPYEVLKLLW----KN 594
Query: 196 NSLHPQMSFYNLSFKFFYRLLQSESRK---------------TPECQT---SYKFKIEHV 237
N ++ +SF + R + ES+ T E Q+ ++ ++H
Sbjct: 595 NE--SELIKCMMSFPNYLRNIPGESQSDNNMEGEGEHSVSIITNESQSETPTHLLDLQHK 768
Query: 238 LDLLRYNIRPKLI---GEEPRGCQSQMIHSITELKEGGVKIKACESRELMDISFGKKWGI 294
+ L + K++ + + I +L+ G+++K +R D+ F W
Sbjct: 769 VILTTSKSKGKILKWTSLKKNEFKPITYRGIEDLRAVGIRLKTSATRRPTDVDFTAGW-- 942
Query: 295 MVKELTIPPLYIGDHRGTVFRNIVAFEKC 323
+LTIP +Y+ + + N++A+E C
Sbjct: 943 FTAKLTIPVIYVNNLAASNLLNLIAYEMC 1029
>BE324073 weakly similar to GP|8953391|emb| putative protein {Arabidopsis
thaliana}, partial (10%)
Length = 272
Score = 55.1 bits (131), Expect = 6e-08
Identities = 26/69 (37%), Positives = 39/69 (55%), Gaps = 5/69 (7%)
Frame = +2
Query: 24 WRGLVEKELQSIKPEEKNESV-----CIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHL 78
W + +E+++ P NE IY+VP + + KAY+P IS GPYHYG +HL
Sbjct: 23 WLDEINEEIENNVPSIANEREQWKKHSIYKVPSQLTELNEKAYVPRAISFGPYHYGKEHL 202
Query: 79 QEMEVLKNK 87
+ ME K++
Sbjct: 203KPMEEHKHR 229
>TC83719 similar to GP|18143763|dbj|BAB79812. hypothetical protein
{Clostridium perfringens str. 13}, partial (7%)
Length = 542
Score = 53.1 bits (126), Expect = 2e-07
Identities = 38/121 (31%), Positives = 60/121 (49%), Gaps = 3/121 (2%)
Frame = -1
Query: 250 IGEEPRGCQSQMIHSITELKEGGVKIKACESREL--MDISFGKKWGIMVKELTIPPLYIG 307
I EE SI +LK G+++ A ++ E +ISF KW EL +P
Sbjct: 359 IDEEEINLNWNTYKSIRDLKTVGIQVVANKTDEWNWSNISFTSKW--FNGELRLPIFLFN 186
Query: 308 DHRGTVFRNIVAFEKCHK-RCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEH 366
D FRN+VA+E C + ++ F++ L+++A+DV L GV + LGSD+
Sbjct: 185 DVTPYFFRNLVAYEMCSDVHHTYECCSFFSFMDSLVDNADDVKELRSAGVFQNLLGSDKE 6
Query: 367 V 367
+
Sbjct: 5 L 3
>TC82957 weakly similar to GP|6957728|gb|AAF32472.1| hypothetical protein
{Arabidopsis thaliana}, partial (11%)
Length = 857
Score = 52.8 bits (125), Expect = 3e-07
Identities = 45/182 (24%), Positives = 84/182 (45%), Gaps = 4/182 (2%)
Frame = +3
Query: 262 IHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFE 321
I S+ EL + GV + ISF K + +P + + + RN+VA+E
Sbjct: 180 IPSVKELVKSGVNFSPTNG-SISSISFDAK----TRTFYLPIIGLDVNTEVFLRNLVAYE 344
Query: 322 KCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPD 381
+T Y +N +I+S +D L KG+I + L SD+ VA++ N ++K +
Sbjct: 345 SSVGSGPLVITRYTELMNGIIDSEDDAKILREKGIILNHLKSDKEVADMWNGMSKSLRLS 524
Query: 382 MNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVG----LSTLGALLALYFTFIQA 437
+L + + N++ + +R + ++ ++ S+ G L+ L A+ L +QA
Sbjct: 525 -RVLFLDKTIEDVNKF---YNSRMKVKML-KFMKSYVFGSWQFLTFLAAIFLLLLMGVQA 689
Query: 438 IC 439
C
Sbjct: 690 FC 695
>TC85014 weakly similar to GP|9758661|dbj|BAB09127.1
emb|CAB72473.1~gene_id:MQJ16.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (6%)
Length = 664
Score = 50.8 bits (120), Expect = 1e-06
Identities = 44/184 (23%), Positives = 83/184 (44%), Gaps = 10/184 (5%)
Frame = +2
Query: 31 ELQSIKPEEKNESVCIYRVPPNMLNVE--PKAYIPSNISIGPYHYGSQHLQEMEVLKNKF 88
ELQ K +N I RV ++ N + K Y P +SIGP H+ + +L E K +
Sbjct: 71 ELQKSKQIPQNSRPKIQRVLKSLRNRKNFEKHYSPKCVSIGPIHHHNTNLILAEKYKLMW 250
Query: 89 FHRLFDPNG--------ANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDG 140
+ + NG ++E + + + + + + ++ M+ VDG
Sbjct: 251 AAKYIENNGYIPEDLHKKIADNIDELKGHFDDDVLTSTRWSLAGFRSLEEKLSWMLFVDG 430
Query: 141 SFIIQLLRDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHP 200
++ +L + NE H+ ++ L + ++++LENQLP VL L+ D++ +
Sbjct: 431 CSLLYILEKANLNEPWHM-NIKVDQLVLVMMDVLLLENQLPYQVLKLLWTDEDESGLIES 607
Query: 201 QMSF 204
M+F
Sbjct: 608 MMNF 619
>TC83489 weakly similar to GP|17528938|gb|AAL38679.1 unknown protein
{Arabidopsis thaliana}, partial (4%)
Length = 1057
Score = 40.8 bits (94), Expect(2) = 2e-06
Identities = 30/116 (25%), Positives = 51/116 (43%)
Frame = +2
Query: 30 KELQSIKPEEKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFF 89
K L +E + I VP ++ AY+P +SIGP + G + L +ME +K +
Sbjct: 248 KALLGAVRKETCQPYSISAVPDDLRKWNESAYMPKVVSIGPLYKGRRELLQMEEIKWRCV 427
Query: 90 HRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQ 145
L + +E + + K + R Y+ EI L + M+ DG F ++
Sbjct: 428 TSLLSRTFGLDA-IERCMEAVIKLDATVRASYIYEITLDRYDLATTMVFDGCFSVR 592
Score = 28.9 bits (63), Expect(2) = 2e-06
Identities = 10/19 (52%), Positives = 17/19 (88%)
Frame = +3
Query: 172 EMIMLENQLPMFVLTKLFE 190
++++LENQ+P+F+L LFE
Sbjct: 693 DLMLLENQIPIFILETLFE 749
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,100,410
Number of Sequences: 36976
Number of extensions: 230208
Number of successful extensions: 1122
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 1098
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1107
length of query: 487
length of database: 9,014,727
effective HSP length: 100
effective length of query: 387
effective length of database: 5,317,127
effective search space: 2057728149
effective search space used: 2057728149
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC130275.9


