

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
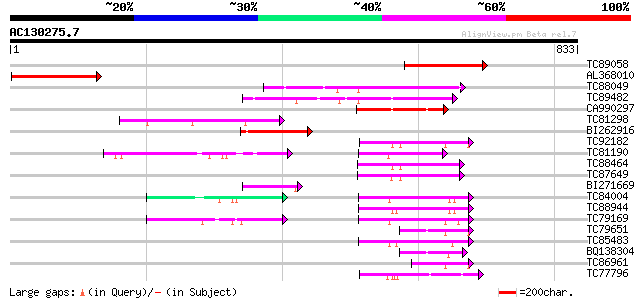
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89058 similar to GP|19071218|gb|AAL84162.1 putative heavy meta... 184 2e-46
AL368010 similar to GP|19071218|g putative heavy metal transport... 143 2e-34
TC88049 similar to GP|13374852|emb|CAC34486. metal-transporting ... 137 2e-32
TC89482 similar to SP|Q9M3H5|AHM1_ARATH Potential cadmium/zinc-t... 132 4e-31
CA990297 similar to GP|15636781|gb copper-transporting P-type AT... 98 1e-20
TC81298 similar to PIR|B96660|B96660 protein F2K11.18 [imported]... 88 2e-17
BI262916 similar to GP|15636781|gb copper-transporting P-type AT... 82 8e-16
TC92182 type IIA calcium ATPase [Medicago truncatula] 80 4e-15
TC81190 type IIB calcium ATPase [Medicago truncatula] 77 2e-14
TC88464 similar to SP|Q9LU41|ACA9_ARATH Potential calcium-transp... 74 2e-13
TC87649 similar to SP|Q9SZR1|ACAA_ARATH Potential calcium-transp... 73 4e-13
BI271669 similar to GP|2668492|dbj metal-transporting P-type ATP... 71 2e-12
TC84004 type IIB calcium ATPase [Medicago truncatula] 71 2e-12
TC88944 type IIB calcium ATPase [Medicago truncatula] 67 2e-11
TC79169 type IIB calcium ATPase [Medicago truncatula] 66 6e-11
TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (E... 65 1e-10
TC85483 type IIB calcium ATPase MCA5 [Medicago truncatula] 62 7e-10
BQ138304 homologue to GP|6688833|emb putative calcium P-type ATP... 61 2e-09
TC86961 similar to GP|1742951|emb|CAA70946.1 Ca2+-ATPase {Arabid... 59 6e-09
TC77796 H+-ATPase 54 3e-07
>TC89058 similar to GP|19071218|gb|AAL84162.1 putative heavy metal
transporter {Arabidopsis thaliana}, partial (10%)
Length = 1618
Score = 184 bits (466), Expect = 2e-46
Identities = 90/123 (73%), Positives = 108/123 (87%)
Frame = +1
Query: 580 IENFKKEGPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAI 639
I FKK+GP AM+GDG+NDAPALA+ADIGISMGISGSALA+ET + ILMSND+RK+PEAI
Sbjct: 1 ITEFKKDGPTAMLGDGLNDAPALASADIGISMGISGSALASETGDIILMSNDLRKIPEAI 180
Query: 640 RLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQEN 699
+LARK RK++EN+++SV K AILALAIAG+P+VW AVL DVGTCLLVILNSML+L
Sbjct: 181 KLARKARRKVIENIVLSVITKVAILALAIAGHPIVWAAVLADVGTCLLVILNSMLLLPRG 360
Query: 700 HKY 702
HK+
Sbjct: 361 HKH 369
>AL368010 similar to GP|19071218|g putative heavy metal transporter
{Arabidopsis thaliana}, partial (11%)
Length = 448
Score = 143 bits (361), Expect = 2e-34
Identities = 72/133 (54%), Positives = 97/133 (72%)
Frame = +3
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ +++S F+V+G+CC++E L+E ILKPL GVK VSVIVP+RTV VVHD LLIS+ +I
Sbjct: 48 KKLQKSYFDVVGLCCSSEVPLIENILKPLQGVKEVSVIVPSRTVIVVHDTLLISQLQIVK 227
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN ARLEA+ R G N++KK P I ++A GLLL LSFLK++Y P ++AL +V G
Sbjct: 228 ALNQARLEANIRIYGNENHKKKWPSIYSVASGLLLLLSFLKFVYTPFKYVALAAVVAGIY 407
Query: 123 KVLFRAIASIRAL 135
+ +AI SIR L
Sbjct: 408 PIFLKAIVSIRNL 446
>TC88049 similar to GP|13374852|emb|CAC34486. metal-transporting ATPase-like
protein {Arabidopsis thaliana}, partial (38%)
Length = 1207
Score = 137 bits (345), Expect = 2e-32
Identities = 93/314 (29%), Positives = 161/314 (50%), Gaps = 18/314 (5%)
Frame = +1
Query: 374 GLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSS 433
GLL++GGD LE L+ + +A DKTGT+TRG+ V +L+ +++E +S
Sbjct: 1 GLLIRGGDVLERLAGVNYIALDKTGTLTRGK-PVVSAIGSIHYGESEILHIAAAVEKTAS 177
Query: 434 HPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIG---------NKRVG 484
HP+A A+++ A + P + + PG G IDGR + +G N R+
Sbjct: 178 HPIAKAIINKAESLELVLPPTKGQIVE--PGFGTLAEIDGRLVAVGSLEWVHERFNTRMN 351
Query: 485 VRAICKRDNCEVQFQRPEISTKKNNC-------EETLVGVFSLVDACRSGALEAMEELKL 537
+ + + S+K + E ++G ++ D R A + LK
Sbjct: 352 PSDLMNLERALMNHSSSTSSSKYSKTVVYVGREGEGIIGAIAISDIVREDAESTVMRLKK 531
Query: 538 LGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-PIAMIGDGI 596
G+++V+L+GD + + + D V A L P +K+ I + K G +AM+GDGI
Sbjct: 532 KGIKTVLLSGDREEAVATIAETVGIENDFVKASLSPQQKSAFISSLKAAGHHVAMVGDGI 711
Query: 597 NDAPALATADIGISM-GISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVII 655
NDAP+LA AD+GI++ + A++ ++ IL+ N I +V +A+ LA+ T K+ +N+
Sbjct: 712 NDAPSLAAADVGIALQNEAQENAASDAASIILLGNKISQVIDALDLAQATMAKVYQNLSW 891
Query: 656 SVGFKCAILALAIA 669
+V + ++A+ IA
Sbjct: 892 AVAYN--VIAIPIA 927
>TC89482 similar to SP|Q9M3H5|AHM1_ARATH Potential cadmium/zinc-transporting
ATPase HMA1 (EC 3.6.3.3) (EC 3.6.3.5). [Mouse-ear
cress], partial (41%)
Length = 1100
Score = 132 bits (333), Expect = 4e-31
Identities = 102/351 (29%), Positives = 176/351 (50%), Gaps = 36/351 (10%)
Frame = +3
Query: 343 ALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITR 402
AL ++++ PCAL ++ P+A A++ A G+LLKGG L+ L+ T+AFDKTGT+T
Sbjct: 15 ALGLMVAASPCALAVA-PLAYATAISSCAKKGILLKGGHVLDALASCHTIAFDKTGTLTT 191
Query: 403 GE--FSVTDFSAGDDINNET--------------LLYWISSIESKSSHPVAGALVDYARL 446
G F + G I N+ L +++E ++HP+ A+V+++
Sbjct: 192 GGLVFKAIEPVYGHHIRNKESNISSCCVPTCEKEALAVAAAMEKGTTHPIGRAVVEHSEG 371
Query: 447 HSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRV---------GVRAICKRDNCEVQ 497
++ V +VENF+ FPG G+ T++ + G + + + C+ ++ E +
Sbjct: 372 KNLPSV--SVENFEYFPGRGLTATVNSIESGAGGANLLKASLGSIDFITSFCQSED-ESK 542
Query: 498 FQRPEISTKKNNCE--------ETLVGVFSLVDACRSGALEAMEELK-LLGVRSVMLTGD 548
+ I+ E V + L D R G + ++EL+ R +MLTGD
Sbjct: 543 KVKEAINASSYGSEFVHAALIINKKVTLIHLEDRPRPGVFDVIQELQDEAKFRVMMLTGD 722
Query: 549 SSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATAD 606
A V S + I H L P +K + +++ ++ G + M+G+GINDAPALA A
Sbjct: 723 HEYSARRVASAV--GIKEFHCNLKPEDKLRHVKDISRDMGGGLIMVGEGINDAPALAAAT 896
Query: 607 IGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISV 657
+GI + SA A ++ +L+ +I VP I +R+TT + +NV +++
Sbjct: 897 VGIVLAHRASATAIAVADVLLLRENISAVPFCIAKSRQTTSLIKQNVALAL 1049
>CA990297 similar to GP|15636781|gb copper-transporting P-type ATPase
{Brassica napus}, partial (19%)
Length = 591
Score = 97.8 bits (242), Expect = 1e-20
Identities = 54/136 (39%), Positives = 90/136 (65%), Gaps = 1/136 (0%)
Frame = +3
Query: 510 CEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHA 569
C L+GV + D+ + A +E L+ +G+ VM+TGD+ + A V ++ I V A
Sbjct: 48 CNGILIGVLGVADSLKREASVVVEGLQKMGITPVMVTGDNWRTARAVAKEV--GIQDVRA 221
Query: 570 ELLPHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNAILM 628
E++P KA ++ +F+K+G I AM+GDGIND+PALA AD+G+++G +G+ +A E +N +LM
Sbjct: 222 EVMPAGKADVVRSFQKDGSIVAMVGDGINDSPALAAADVGMAIG-AGTDIAIEAANYVLM 398
Query: 629 SNDIRKVPEAIRLARK 644
+++ V AI L++K
Sbjct: 399 RDNLEDVITAIDLSKK 446
>TC81298 similar to PIR|B96660|B96660 protein F2K11.18 [imported] -
Arabidopsis thaliana, partial (23%)
Length = 848
Score = 87.8 bits (216), Expect = 2e-17
Identities = 73/261 (27%), Positives = 117/261 (43%), Gaps = 18/261 (6%)
Frame = +1
Query: 162 IIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAET--------GER-VDVNDVKMN 212
+ LF +A + K A++ L ++ P A + GER +D ++ N
Sbjct: 7 LFLLFYLASIWKCWLKGKTSQAIAKLMDLTPDTATLLTLDDDKGNVLGEREIDSRLIQKN 186
Query: 213 TILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----P 267
++ V G + DG VV G+ V+E M+TGE+ PV K +V GT+N N
Sbjct: 187 DVIKVVPGTKVASDGFVVWGQSHVNESMITGEAKPVAKMKGDMVIGGTVNENGVLHVKVT 366
Query: 268 KIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVV 327
+I +T +I + + + K + F VI + + +
Sbjct: 367 RIGSETALSQIVRLVESAQMAKAPVQKYADQISKYFVPIVIVLSLSTWISWFVAGKLHSY 546
Query: 328 PAALSVPDMEPWFHLALV----VLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYL 383
P + +P F LAL V++ CPCAL L+TP A+ A G+L+KGG L
Sbjct: 547 PKSW-IPSSMNSFELALQFGISVMVIACPCALGLATPTAVMVGTGVGATQGVLIKGGQAL 723
Query: 384 ETLSRIKTVAFDKTGTITRGE 404
E+ ++ + FDKTGT+T G+
Sbjct: 724 ESAHKVNCIVFDKTGTLTIGK 786
>BI262916 similar to GP|15636781|gb copper-transporting P-type ATPase
{Brassica napus}, partial (15%)
Length = 527
Score = 82.0 bits (201), Expect = 8e-16
Identities = 43/106 (40%), Positives = 67/106 (62%)
Frame = +3
Query: 340 FHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGT 399
F +++VV+ CPCAL L+TP A+ A A +G+L+KGG+ LE +K V FDKTGT
Sbjct: 165 FSISVVVI--ACPCALGLATPTAVMVATGVGANNGVLIKGGESLERAQMVKYVIFDKTGT 338
Query: 400 ITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYAR 445
+T+G+ +V+ ++ L ++S E+ S HP+A A++ YAR
Sbjct: 339 LTQGKATVSVAKVFAGMDRGEFLKLVASAEAXSEHPLAKAILQYAR 476
>TC92182 type IIA calcium ATPase [Medicago truncatula]
Length = 3148
Score = 79.7 bits (195), Expect = 4e-15
Identities = 62/210 (29%), Positives = 99/210 (46%), Gaps = 43/210 (20%)
Frame = +1
Query: 515 VGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLK------------- 561
VGV L D R +A+E+ K G+R +++TGD+ A + ++K
Sbjct: 1840 VGVVGLRDPPREEVHKAIEDCKQAGIRVMVITGDNKSTAEAICKEIKLFSTDEDLTGQSL 2019
Query: 562 ---NAIDIVHAELL---------------PHEKAKLIENFKKEGPI-AMIGDGINDAPAL 602
+ + H+E + P K +++ K+ G I AM GDG+NDAPAL
Sbjct: 2020 TGKEFMSLSHSEQVKLLLRNGGKVFSRAEPRHKQEIVRLLKEMGEIVAMTGDGVNDAPAL 2199
Query: 603 ATADIGISMGISGSALANETSNAILMSNDIRKVPEAI---RLARKTTRKLVENVIIS-VG 658
ADIGI+MGI+G+ +A E S+ +L ++ + AI R + + +I S VG
Sbjct: 2200 KLADIGIAMGITGTEVAKEASDMVLADDNFSTIVSAIAEGRAIYNNMKAFIRYMISSNVG 2379
Query: 659 FKCAILALAIAGYP-------LVWLAVLTD 681
+I A G P L+W+ ++TD
Sbjct: 2380 EVISIFLTAALGIPECMIPVQLLWVNLVTD 2469
Score = 40.0 bits (92), Expect = 0.004
Identities = 25/105 (23%), Positives = 47/105 (43%)
Frame = +1
Query: 339 WFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTG 398
+F +A+ + ++ P L + K A +++ +ETL + DKTG
Sbjct: 919 YFKIAVALAVAAIPEGLPAVITTCLALGTRKMAQKNAIVRKLPSVETLGCTTVICSDKTG 1098
Query: 399 TITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDY 443
T+T + S T+F + +T + S+E + P G +VD+
Sbjct: 1099 TLTTNQMSATEFFT---LGGKTTACRVISVEGTTYDPKDGGIVDW 1224
>TC81190 type IIB calcium ATPase [Medicago truncatula]
Length = 3277
Score = 77.4 bits (189), Expect = 2e-14
Identities = 49/163 (30%), Positives = 78/163 (47%), Gaps = 32/163 (19%)
Frame = +1
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAM------------------ 554
TL+G+ L D CR +A+E K GV M+TGD+ A
Sbjct: 2011 TLLGIVGLKDPCRPNTKKAVETCKAAGVEIKMITGDNIFTAKAIAIECGILDSNSDHAKA 2190
Query: 555 -----------YVQSQLKNAIDIVH--AELLPHEKAKLIENFKKEGPI-AMIGDGINDAP 600
Y + + +D + A P +K +++ +K+G + A+ GDG NDAP
Sbjct: 2191 GEVVEGVEFRSYTEEERMEKVDNIRVMARSSPMDKLLMVQCLRKKGHVVAVTGDGTNDAP 2370
Query: 601 ALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLAR 643
AL ADIG+SMGI G+ +A E+S+ +++ ++ V +R R
Sbjct: 2371 ALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGR 2499
Score = 51.6 bits (122), Expect = 1e-06
Identities = 70/332 (21%), Positives = 135/332 (40%), Gaps = 54/332 (16%)
Frame = +1
Query: 138 NINILVLLAVCGTAAL----------QDFTDGGVI---IFLFSIAQWLETRATHKALVAM 184
N +++L VC +L + + +GG I +FL + L + +
Sbjct: 532 NDTTIIILLVCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQERQFHKL 711
Query: 185 SSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTG 243
S ++N + +++ + DV + I+++K GD IP DG+ + G +VDE +TG
Sbjct: 712 SKISNNIKVEVVRNGRPQQISIFDVLVGDIVSLKIGDQIPADGVFLSGYSLQVDESSMTG 891
Query: 244 ESFPVTKES--DSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSS---GKNVKT 298
ES V E + +G ++ Y +++ + KN S G+ SS N +T
Sbjct: 892 ESDHVEIEPLRAPFLLSGAKVVDGYAQMLVTSVG------KNTSWGQMMSSISRDTNERT 1053
Query: 299 C*RSF*QKVIST----------------------------KIHR*FR-------KVLYSC 323
++ K+ S+ K ++ FR V+ S
Sbjct: 1054PLQARLDKLTSSIGKVGLAVAFLVLLVLLIRYFTGNSHDEKGNKEFRGSKTDINDVMNSV 1233
Query: 324 IAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYL 383
+++V AA+++ +V + G P A+ L+ + ++ + +++
Sbjct: 1234VSIVAAAVTI---------VVVAIPEGLPLAVTLT----LAYSMKRMMADHAMVRKLSAC 1374
Query: 384 ETLSRIKTVAFDKTGTITRGEFSVTDFSAGDD 415
ET+ + DKTGT+T + VT F G +
Sbjct: 1375ETMGSATVICTDKTGTLTLNQMRVTKFCLGPE 1470
>TC88464 similar to SP|Q9LU41|ACA9_ARATH Potential calcium-transporting
ATPase 9 plasma membrane-type (EC 3.6.3.8) (Ca2+-ATPase
isoform 9)., partial (43%)
Length = 1608
Score = 73.9 bits (180), Expect = 2e-13
Identities = 51/189 (26%), Positives = 90/189 (46%), Gaps = 31/189 (16%)
Frame = +3
Query: 511 EETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQL------KNAI 564
E L+ + + D CR G +A+ GV+ M+TGD+ Q A + + + A+
Sbjct: 249 ELVLLAIVGIKDPCRPGVKDAVRICTDAGVKVRMVTGDNLQTAKAIALECGILASNEEAV 428
Query: 565 D--IVHAELL----------------------PHEKAKLIENFKKEGP-IAMIGDGINDA 599
D I+ ++ P++K L++ +K G +A+ GDG NDA
Sbjct: 429 DPCIIEGKVFRELSEKEREQVAKKITVMGRSSPNDKLLLVQALRKGGDVVAVTGDGTNDA 608
Query: 600 PALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGF 659
PAL ADIG+SMGI G+ +A E+S+ I++ ++ V + +R R + + + +
Sbjct: 609 PALHEADIGLSMGIQGTEVAKESSDIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTV 788
Query: 660 KCAILALAI 668
A L + +
Sbjct: 789 NVAALVINV 815
>TC87649 similar to SP|Q9SZR1|ACAA_ARATH Potential calcium-transporting
ATPase 10 plasma membrane-type (EC 3.6.3.8), partial
(43%)
Length = 2159
Score = 73.2 bits (178), Expect = 4e-13
Identities = 46/189 (24%), Positives = 91/189 (47%), Gaps = 31/189 (16%)
Frame = +3
Query: 511 EETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ---LKNAIDIV 567
E L+ + + D CR G +++ + GV+ M+TGD+ + A + + L + D+
Sbjct: 342 ELVLLAIVGIKDPCRPGVKNSVQLCQKAGVKVKMVTGDNVKTAKAIALECGILSSLADVT 521
Query: 568 HAELL---------------------------PHEKAKLIENFKKEGPI-AMIGDGINDA 599
++ P++K L++ +++G + A+ GDG NDA
Sbjct: 522 ERSVIEGKTFRALSDSEREEIAESISVMGRSSPNDKLLLVQALRRKGHVVAVTGDGTNDA 701
Query: 600 PALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGF 659
PAL ADIG++MGI+G+ +A E+S+ I++ ++ V + +R R + + + +
Sbjct: 702 PALHEADIGLAMGIAGTEVAKESSDIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTV 881
Query: 660 KCAILALAI 668
A L + +
Sbjct: 882 NVAALVINV 908
>BI271669 similar to GP|2668492|dbj metal-transporting P-type ATPase
{Arabidopsis thaliana}, partial (13%)
Length = 509
Score = 70.9 bits (172), Expect = 2e-12
Identities = 41/92 (44%), Positives = 54/92 (58%), Gaps = 4/92 (4%)
Frame = +1
Query: 342 LALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTIT 401
LA VL+ CPCAL L+TP A+ + A GLLL+GG+ LE + + TV FDKTGT+T
Sbjct: 226 LACSVLVIACPCALGLATPTAVLVGTSLGAKRGLLLRGGNILEKFAMVNTVVFDKTGTLT 405
Query: 402 RGEFSVTDFSAGDDINN----ETLLYWISSIE 429
G+ VT G I N +T + +S IE
Sbjct: 406 IGKPVVTKIVTGTCIENANSSQTKINALSDIE 501
>TC84004 type IIB calcium ATPase [Medicago truncatula]
Length = 3655
Score = 70.9 bits (172), Expect = 2e-12
Identities = 54/207 (26%), Positives = 94/207 (45%), Gaps = 38/207 (18%)
Frame = +1
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYV---------------- 556
TL+ + + D R G EA++ G+ M+TGD+ A +
Sbjct: 2176 TLIALVGIKDPVRPGVKEAVQTCIAAGITVRMVTGDNINTAKAIAKECGILTDDGVAIEG 2355
Query: 557 -------QSQLKNAIDIVH--AELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATA 605
Q+K+ I + A LP +K KL+ N + +A+ GDG NDAPAL A
Sbjct: 2356 PSFRELSDEQMKDIIPRIQVMARSLPLDKHKLVTNLRNMFGEVVAVTGDGTNDAPALHEA 2535
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE--------NVI 654
DIG++MGI+G+ +A E ++ I+M ++ + ++ R +K V+ +I
Sbjct: 2536 DIGLAMGIAGTEVAKEKADVIIMDDNFATIVNVVKWGRAVYINIQKFVQFQLTVNVVALI 2715
Query: 655 ISVGFKCAILALAIAGYPLVWLAVLTD 681
I+ C + + L+W+ ++ D
Sbjct: 2716 INFVSACITGSAPLTAVQLLWVNLIMD 2796
Score = 47.0 bits (110), Expect = 3e-05
Identities = 55/232 (23%), Positives = 93/232 (39%), Gaps = 26/232 (11%)
Frame = +1
Query: 202 ERVDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFPV-TKESDSLVWAG 259
+++ + DV + I+ + GD +P DGI + G +DE L+GES PV E + +G
Sbjct: 970 KKISIYDVVVGDIIHLSTGDQVPADGIYISGYSLLIDESSLSGESEPVFITEEHPFLLSG 1149
Query: 260 TINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QK-----------VI 308
T + K++ T VG R GK ++T + I
Sbjct: 1150 TKVQDGQGKMLVTT------------VGMRTEWGKLMETLNEGGEDETPLQVKLNGVATI 1293
Query: 309 STKIHR*FRKVLYSCIAV---VPAAL----------SVPDMEPWFHLALVVLLSGCPCAL 355
KI F V + + V V AL + +F +A+ +++ P L
Sbjct: 1294 IGKIGLFFAIVTFLVLTVRFLVEKALHGEFGNWSSNDATKLLDFFAIAVTIIVVAVPEGL 1473
Query: 356 ILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
L+ +++ A+ K L++ ET+ + DKTGT+T V
Sbjct: 1474 PLAVTLSLAFAMKKLMNDMALVRHLSACETMGSASCICTDKTGTLTTNHMVV 1629
>TC88944 type IIB calcium ATPase [Medicago truncatula]
Length = 1883
Score = 67.4 bits (163), Expect = 2e-11
Identities = 55/206 (26%), Positives = 96/206 (45%), Gaps = 37/206 (17%)
Frame = +3
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQL------------ 560
T VG+ + D R G E++ + G+ M+TGD+ A + +
Sbjct: 711 TCVGIVGIKDPVRPGVRESVAICRSAGITVRMVTGDNINTAKAIARECGILTDGIAIEGP 890
Query: 561 -------KNAIDI-----VHAELLPHEKAKLIENFKK--EGPIAMIGDGINDAPALATAD 606
K +DI V A P +K L+++ + E +A+ GDG NDAPAL AD
Sbjct: 891 EFREMSEKELLDIIPKIQVMARSSPMDKHTLVKHLRTTFEEVVAVTGDGTNDAPALHEAD 1070
Query: 607 IGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE-----NVI-ISV 657
IG++MGI+G+ +A E+++ I++ ++ + + R +K V+ NV+ + V
Sbjct: 1071IGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVALIV 1250
Query: 658 GFKCAILA--LAIAGYPLVWLAVLTD 681
F A L + L+W+ ++ D
Sbjct: 1251NFTSACLTGNAPLTAVQLLWVNMIMD 1328
>TC79169 type IIB calcium ATPase [Medicago truncatula]
Length = 3565
Score = 65.9 bits (159), Expect = 6e-11
Identities = 51/207 (24%), Positives = 93/207 (44%), Gaps = 38/207 (18%)
Frame = +2
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYV---------------- 556
TL+ + + D R G EA+++ G+ M+TGD+ A +
Sbjct: 2102 TLITIVGIKDPVRPGVKEAVQKCLAAGISVRMVTGDNINTAKAIAKECGILTEGGVAIEG 2281
Query: 557 -------QSQLKNAIDIVH--AELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATA 605
+ Q+K+ I + A LP +K L+ + +A+ GDG NDAPAL +
Sbjct: 2282 PEFRNLSEEQMKDIIPRIQVMARSLPLDKHTLVTRLRNMFGEVVAVTGDGTNDAPALHES 2461
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEA--------IRLARKTTRKLVENVIISV 657
DIG++MGI+G+ +A E ++ I+M ++ + + I + + +L NV+ +
Sbjct: 2462 DIGLAMGIAGTEVAKENADVIIMDDNFTTIVKVAKWGRAIYINIQKFVQFQLTVNVVALI 2641
Query: 658 G---FKCAILALAIAGYPLVWLAVLTD 681
C A + L+W+ ++ D
Sbjct: 2642 TNFVSACITGAAPLTAVQLLWVNLIMD 2722
Score = 46.6 bits (109), Expect = 4e-05
Identities = 53/225 (23%), Positives = 95/225 (41%), Gaps = 19/225 (8%)
Frame = +2
Query: 202 ERVDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFPVTKES-DSLVWAG 259
++V + D+ + I+ + GD +P DGI ++G +DE L+GES PV ++ + +G
Sbjct: 902 QKVSIYDLVVGDIVHLSTGDQVPADGIFIQGYSLLIDESSLSGESEPVDIDNRRPFLLSG 1081
Query: 260 TINMNAYPKIIFDTYDFRIYKCK---NYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*F 316
T + K+I T R K S G + VK + + KI F
Sbjct: 1082 TKVQDGQAKMIVTTVGMRTEWGKLMETLSEGGEDETPLQVKLNGVA----TVIGKIGLTF 1249
Query: 317 RKVLYSCIA---VVPAALSVPDMEPW-----------FHLALVVLLSGCPCALILSTPVA 362
+ + + V+ A++ D W F +A+ +++ P L L+ ++
Sbjct: 1250 AVLTFLVLTARFVIEKAIN-GDFTSWSSEDALKLLDYFAIAVTIIVVAIPEGLPLAVTLS 1426
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
+ A+ K L++ ET+ + DKTGT+T V
Sbjct: 1427 LAFAMKKLMNDRALVRHLSACETMGSASCICTDKTGTLTTNHMVV 1561
>TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (EC 3.6.1.38)
ECA3 [imported] - Arabidopsis thaliana, partial (38%)
Length = 1546
Score = 64.7 bits (156), Expect = 1e-10
Identities = 42/121 (34%), Positives = 66/121 (53%), Gaps = 12/121 (9%)
Frame = +3
Query: 573 PHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSND 631
P K L+E + + + AM GDG+NDAPAL ADIGI+MG SG+A+A S+ +L ++
Sbjct: 183 PSHKRMLVEALQHQNEVVAMTGDGVNDAPALKKADIGIAMG-SGTAVAKSASDMVLADDN 359
Query: 632 IRKVPEAI---RLARKTTRKLVENVIIS-VGFKCAILALAIAGYP-------LVWLAVLT 680
+ A+ R T++ + +I S +G I A+ G P L+W+ ++T
Sbjct: 360 FASIVAAVAEGRAIYNNTKQFIRYMISSNIGEVVCIFVAAVLGIPDTLAPVQLLWVNLVT 539
Query: 681 D 681
D
Sbjct: 540 D 542
>TC85483 type IIB calcium ATPase MCA5 [Medicago truncatula]
Length = 2541
Score = 62.4 bits (150), Expect = 7e-10
Identities = 50/207 (24%), Positives = 91/207 (43%), Gaps = 38/207 (18%)
Frame = +2
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYV---------------- 556
T +GV + D R G E++ + G+ M+TGD+ A +
Sbjct: 1004 TCIGVVGIKDPVRPGVKESVALCRSAGITVRMVTGDNINTAKAIARECGILTDDGIAIEG 1183
Query: 557 ----QSQLKNAIDI-----VHAELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATA 605
+ L+ +++ V A P +K L+ + + +A+ GDG NDAPAL A
Sbjct: 1184 PEFREKSLEELLELIPKIQVMARSSPLDKHTLVRHLRTTFGEVVAVTGDGTNDAPALHEA 1363
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL---------VENVIIS 656
DIG++MGI+G+ +A E+++ I++ ++ + + R + V V +
Sbjct: 1364 DIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNIVALI 1543
Query: 657 VGFKCAIL--ALAIAGYPLVWLAVLTD 681
V F A L + L+W+ ++ D
Sbjct: 1544 VNFTSACLTGTAPLTAVQLLWVNMIMD 1624
Score = 35.0 bits (79), Expect = 0.12
Identities = 19/73 (26%), Positives = 37/73 (50%)
Frame = +2
Query: 335 DMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAF 394
+M +F +A+ +++ P L L+ +++ A+ K L++ ET+ T+
Sbjct: 248 EMLEYFAIAVTIVVVAVPEGLPLAVTLSLAFAMKKMMNDKALVRNLAACETMGSATTICS 427
Query: 395 DKTGTITRGEFSV 407
DKTGT+T +V
Sbjct: 428 DKTGTLTTNHMTV 466
>BQ138304 homologue to GP|6688833|emb putative calcium P-type ATPase
{Neurospora crassa}, partial (13%)
Length = 412
Score = 61.2 bits (147), Expect = 2e-09
Identities = 38/105 (36%), Positives = 62/105 (58%), Gaps = 5/105 (4%)
Frame = +3
Query: 573 PHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSND 631
P K+KL++ +K G + AM GDG+NDAPAL ADIG++MG SG+ +A ++ +L+ ++
Sbjct: 57 PTHKSKLVDLLQKAGEVVAMTGDGVNDAPALKKADIGVAMG-SGTDVAKLAADMVLVDDN 233
Query: 632 IRKVPEAIRLAR---KTTRKLVENVIIS-VGFKCAILALAIAGYP 672
+ A+ R T++ + +I S +G +I A AG P
Sbjct: 234 FATIEGAVEEGRSIYNNTQQFIRYLISSNIGEVVSIFLTAAAGMP 368
>TC86961 similar to GP|1742951|emb|CAA70946.1 Ca2+-ATPase {Arabidopsis
thaliana}, partial (77%)
Length = 1488
Score = 59.3 bits (142), Expect = 6e-09
Identities = 36/102 (35%), Positives = 56/102 (54%), Gaps = 11/102 (10%)
Frame = +1
Query: 591 MIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAI---RLARKTTR 647
M GDG+NDAPAL ADIGI+MGI+G+ +A E S+ +L ++ + A+ R +
Sbjct: 55 MTGDGVNDAPALKLADIGIAMGIAGTEVAKEASDMVLADDNFSSIVAAVGEGRSIYNNMK 234
Query: 648 KLVENVIIS-VGFKCAILALAIAGYP-------LVWLAVLTD 681
+ +I S +G +I A G P L+W+ ++TD
Sbjct: 235 AFIRYMISSNIGEVASIFLTAALGIPEGLIPVQLLWVNLVTD 360
>TC77796 H+-ATPase
Length = 3416
Score = 53.5 bits (127), Expect = 3e-07
Identities = 54/211 (25%), Positives = 89/211 (41%), Gaps = 29/211 (13%)
Frame = +1
Query: 515 VGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVA------------MYVQSQL-- 560
VG+ L D R + E + LGV M+TGD +A MY S L
Sbjct: 1537 VGLLPLFDPPRHDSAETITRALNLGVNVKMITGDQLAIAKETGRRLGMGTNMYPSSSLLG 1716
Query: 561 --KNAI-------DIVH-----AELLPHEKAKLIENFKKEGPIA-MIGDGINDAPALATA 605
K+A +++ A + P K ++++ ++ I M GDG+NDAPAL A
Sbjct: 1717 QSKDAAVSALPVDELIEKADGFAGVFPEHKYEIVKKLQERKHICGMTGDGVNDAPALKRA 1896
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILA 665
DIGI++ + A A S+ +L + + A+ +R +++ I +V I+
Sbjct: 1897 DIGIAVADATDA-ARGASDIVLTEPGLSVIISAVLTSRAIFQRMKNYTIYAVSITIRIV- 2070
Query: 666 LAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
L+W ++ ILN I+
Sbjct: 2071 FGFMFIALIWKFDFAPFMVLIIAILNDGTIM 2163
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,889,107
Number of Sequences: 36976
Number of extensions: 311575
Number of successful extensions: 1496
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 1459
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1484
length of query: 833
length of database: 9,014,727
effective HSP length: 104
effective length of query: 729
effective length of database: 5,169,223
effective search space: 3768363567
effective search space used: 3768363567
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC130275.7


