

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.11 - phase: 0
(309 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
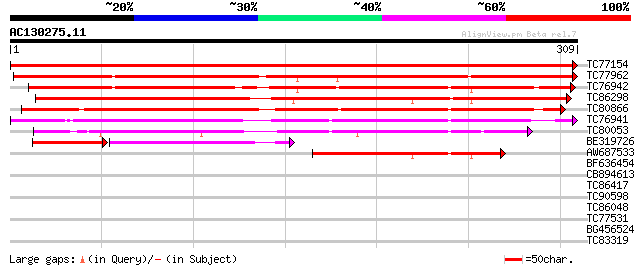
Sequences producing significant alignments: (bits) Value
TC77154 weakly similar to PIR|S49878|S49878 probable narbonin - ... 639 0.0
TC77962 similar to PIR|S49848|S49848 probable narbonin - jack be... 340 4e-94
TC76942 weakly similar to PIR|S44033|S44033 narbonin - Vicia pan... 249 9e-67
TC86298 weakly similar to PIR|S49878|S49878 probable narbonin - ... 248 3e-66
TC80866 weakly similar to PIR|T12156|T12156 nodulin isoform Nvf... 231 3e-61
TC76941 similar to PIR|S44033|S44033 narbonin - Vicia pannonica,... 210 6e-55
TC80053 similar to PIR|JC7335|JC7335 chitinase (EC 3.2.1.14) 1 -... 141 4e-34
BE319726 weakly similar to PIR|S49848|S49 probable narbonin - ja... 83 5e-27
AW687533 weakly similar to PIR|T12157|T121 nodulin - fava bean, ... 107 8e-24
BF636454 similar to PIR|S69190|S691 myb-related protein 1 - toma... 30 1.5
CB894613 similar to GP|16226943|gb AT4g27410/F27G19_10 {Arabidop... 29 2.6
TC86417 similar to GP|16226943|gb|AAL16305.1 AT4g27410/F27G19_10... 29 2.6
TC90598 similar to GP|8953389|emb|CAB96662.1 putative protein {A... 29 2.6
TC86048 weakly similar to PIR|H96561|H96561 probable peptide tra... 28 5.8
TC77531 similar to PIR|T06562|T06562 CP12 protein precursor chl... 28 5.8
BG456524 similar to GP|9279575|dbj emb|CAB36798.1~gene_id:MLN21.... 27 7.6
TC83319 similar to PIR|G86357|G86357 Similar to auxin-independen... 27 10.0
>TC77154 weakly similar to PIR|S49878|S49878 probable narbonin - soybean,
partial (32%)
Length = 1260
Score = 639 bits (1647), Expect = 0.0
Identities = 309/309 (100%), Positives = 309/309 (100%)
Frame = +1
Query: 1 MSHTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGK 60
MSHTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGK
Sbjct: 52 MSHTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGK 231
Query: 61 GKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI 120
GKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI
Sbjct: 232 GKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI 411
Query: 121 KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKS 180
KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKS
Sbjct: 412 KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKS 591
Query: 181 IKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDV 240
IKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDV
Sbjct: 592 IKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDV 771
Query: 241 KLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEE 300
KLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEE
Sbjct: 772 KLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEE 951
Query: 301 VLQDLLTDN 309
VLQDLLTDN
Sbjct: 952 VLQDLLTDN 978
>TC77962 similar to PIR|S49848|S49848 probable narbonin - jack bean, partial
(18%)
Length = 1239
Score = 340 bits (872), Expect = 4e-94
Identities = 171/312 (54%), Positives = 225/312 (71%), Gaps = 5/312 (1%)
Frame = +2
Query: 3 HTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGK 62
H+ I V++P IFREY+GVKD P+ L DFP +I+DD+ +FHFILG E Y DGKG
Sbjct: 77 HSQNIKTVLRPKIFREYVGVKDEPETLDDFPVNIIHDDVNQFHFILGFATEAYK-DGKGT 253
Query: 63 GEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKK 122
G F R WNF+ FSP KV + KK +KN+KV+I+IGG G + PFNPKE + W A SI+
Sbjct: 254 GHFIRDWNFDYFSPEKVFEHKKKYKNMKVMITIGGHGPKYPFNPKEKKVWIFNATSSIRH 433
Query: 123 LINEYQEYSKDSSSTDECHCDDIIDGIDINYEY--SNCNPDEFSSCIGELIRKLKKS--- 177
+I +Y+ Y + +S CHC IIDGIDINYEY S+ +FS+CIGE+I++LKK
Sbjct: 434 IIQDYENYLVNDNS---CHCTSIIDGIDINYEYIDSSVTGADFSNCIGEVIKRLKKDKHV 604
Query: 178 SKSIKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYG 237
SKS++ VSIAPTELL+ HY L+W +K IN+VDYKFYNQT+S+ +E LYN+L+ +YG
Sbjct: 605 SKSMEYVSIAPTELLQAHYRTLFWDHKMNINYVDYKFYNQTISTENEFDELYNQLVTDYG 784
Query: 238 TDVKLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYF 297
++KLL GVSTDP S+T M R VFI+G L+ ++SLPG+FVW+ANDSA P + D PY
Sbjct: 785 GELKLLVGVSTDP-SDTKMKRQVFIEGVTRLINNKSLPGLFVWSANDSANPPSNDIEPYI 961
Query: 298 LEEVLQDLLTDN 309
LEE+L +L T+N
Sbjct: 962 LEEILLELFTNN 997
>TC76942 weakly similar to PIR|S44033|S44033 narbonin - Vicia pannonica,
partial (45%)
Length = 1361
Score = 249 bits (636), Expect = 9e-67
Identities = 140/302 (46%), Positives = 192/302 (63%), Gaps = 4/302 (1%)
Frame = +3
Query: 11 VKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWN 70
V P+IFREYIGVK P +L +FPA++I I EFHFILG E Y DGKG G F +W
Sbjct: 72 VNPVIFREYIGVKSYPDSLNNFPADIIGRHIPEFHFILGFAHETYV-DGKGTGIFNASWK 248
Query: 71 FNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEY 130
F P V +K +H NVKV+ISIGG + PF+P W A +S+KK+ +Q Y
Sbjct: 249 IPFFGPDNVDDIKTNHGNVKVVISIGGRDTKYPFHPAHKLEWCDNAVESLKKI---FQLY 419
Query: 131 SKDSSSTDECHCDDIIDGIDINYEY--SNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAP 188
++ +S C ++IDGIDINYEY + + ++FS CIG++I++LKK I +VSIAP
Sbjct: 420 NRTNS------CYNLIDGIDINYEYIHPDVSEEDFSYCIGDVIKRLKKDV-GIDVVSIAP 578
Query: 189 TELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDVKLLPGVST 248
+ + HY LY A + INWV+Y+FY T+ S DE VNL+ L +EYG+ KLL G ST
Sbjct: 579 SHETQKHYKTLYLARTNDINWVNYQFYIDTLKSKDEFVNLFLNLSDEYGSK-KLLAGAST 755
Query: 249 DP--DSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEEVLQDLL 306
DP ++R+ F++GC L ++SL GIF+WNANDSA SN + P+ LE+ Q++L
Sbjct: 756 DPADAGKGKLSREDFLEGCVDLHSTQSLRGIFIWNANDSA--SNPNGKPFSLEKKAQEIL 929
Query: 307 TD 308
+
Sbjct: 930 NN 935
>TC86298 weakly similar to PIR|S49878|S49878 probable narbonin - soybean,
partial (71%)
Length = 1281
Score = 248 bits (632), Expect = 3e-66
Identities = 138/304 (45%), Positives = 187/304 (61%), Gaps = 12/304 (3%)
Frame = +1
Query: 15 IFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNF 74
IFREYIGVK NL+DFP E+I +I EFHFILG E Y KG G F TWN F
Sbjct: 139 IFREYIGVKPFSTNLRDFPVEIIKTNISEFHFILGFATEEYDAQNKGTGVFKETWNTRAF 318
Query: 75 SPAKVAKLKKDHKNVKVIISIGGFG-AENPFNPKEIESWSTKAKQSIKKLINEYQEYSKD 133
P V LK ++ NVKV+ISIGG + PFNP E W T+A S+K +I +Y++ +
Sbjct: 319 GPEAVRNLKGNNPNVKVVISIGGNDTVKTPFNPVEETIWITRAVSSLKVIIQKYKDQT-- 492
Query: 134 SSSTDECHCDDIIDGIDINY-----EYSNCNPDEFSSCIGELIRKLKKSS-KSIKLVSIA 187
+IIDGIDINY ++ F+ CIGE+I +LK + IK+VSIA
Sbjct: 493 ---------GNIIDGIDINYLNVFHTTNDTGKLRFARCIGEVITQLKNDNYLRIKIVSIA 645
Query: 188 PTELLKPHYHKLYWANKDIINWVDYKFYNQT--VSSADELVNLYNKLLNEYGTDVKLLPG 245
P+E + HY L+W N+ INWV+Y+FYNQ+ VS+ D+ + LY+++ Y + +LPG
Sbjct: 646 PSETNEIHYRNLFWQNEANINWVNYQFYNQSKAVSTLDDFLKLYDQVSRNYKPSI-VLPG 822
Query: 246 VSTDP---DSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEEVL 302
VSTD + M R+ FI GC+ LL+ SLPG+F+WNA+DS +P +N P+ LE++L
Sbjct: 823 VSTDKLHIEPVDKMPREHFIAGCRHLLQIASLPGVFLWNADDSTIPLPNENKPFVLEDIL 1002
Query: 303 QDLL 306
Q LL
Sbjct: 1003QSLL 1014
>TC80866 weakly similar to PIR|T12156|T12156 nodulin isoform Nvf32-A1 -
fava bean, partial (40%)
Length = 970
Score = 231 bits (589), Expect = 3e-61
Identities = 128/297 (43%), Positives = 180/297 (60%)
Frame = +1
Query: 7 INAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFY 66
+ V P IFREYIGVK P +L +FP E+I + FHFILG + Y+ +GKG G F
Sbjct: 52 VTMXVDPFIFREYIGVKPYPASLNNFPYEIII--AKHFHFILGFANDSYNEEGKGTGNFN 225
Query: 67 RTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINE 126
WN + F P V LK+ + +VKV+ISIGG A PF P E W A S+K++I
Sbjct: 226 ANWNSDFFGPQNVMALKRKYPHVKVVISIGGRDANFPFFPAAREEWCGNAVDSLKEIIRS 405
Query: 127 YQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSI 186
Y + S + + +IDGIDI Y+Y N N ++FS+ +G++I +LKK + I +VSI
Sbjct: 406 YNDCSVEDNI--------LIDGIDIFYDYINTNENDFSNYVGDVINRLKKEVR-IDVVSI 558
Query: 187 APTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDVKLLPGV 246
AP+ HY +LY A D INWV+Y+FY Q + S ++ +NL+ L EY ++ KLL G
Sbjct: 559 APSHETHKHYKELYLACTDDINWVNYQFYMQPIPSKNDFLNLFLNLAKEYDSN-KLLVGG 735
Query: 247 STDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEEVLQ 303
S+DP N D F++ C L +++SL GIF+WNANDSA + P++LE+ LQ
Sbjct: 736 SSDPIDADNFNPDEFVEACNDLHKTKSLRGIFIWNANDSA----NNVPPFYLEKKLQ 894
>TC76941 similar to PIR|S44033|S44033 narbonin - Vicia pannonica, partial
(23%)
Length = 1119
Score = 210 bits (534), Expect = 6e-55
Identities = 126/310 (40%), Positives = 175/310 (55%), Gaps = 1/310 (0%)
Frame = +3
Query: 1 MSHTHQINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGK 60
MSH N V +IFREYIGVK PK K FP ++ N I EFH IL E Y+ DGK
Sbjct: 66 MSHPIIDNNAVDRVIFREYIGVKPYPKPFK-FP-DIDNYKIAEFHIILAFAHETYNEDGK 239
Query: 61 GKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI 120
G G F W+ + V +K+ NVK++ SIGG G + PF+P E W A S+
Sbjct: 240 GTGSFSADWDLDICGIQSVKDIKQKCPNVKLVFSIGGRGTKYPFSPIEKNYWCDNAVDSL 419
Query: 121 KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPD-EFSSCIGELIRKLKKSSK 179
K +I +Y +DI GIDINYE+ N N + +FS+ +G++I +L K+
Sbjct: 420 KTIIKQY---------------NDIFAGIDINYEHINTNDENDFSNYVGDVINRL-KNEV 551
Query: 180 SIKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTD 239
I +VSIAP+ +Y LY A+ D INWVDY+FY Q + + +E ++L+ L EY +
Sbjct: 552 GIDVVSIAPSHANDNYYKLLYSAHADDINWVDYQFYMQPIPTENEFLSLFLSLAREYALE 731
Query: 240 VKLLPGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLE 299
KLL G STDP N+ DVF++ C +L++ +SL GIF+WNAND E
Sbjct: 732 -KLLVGASTDPRDGGNVPLDVFVQTCTNLIKHKSLSGIFIWNAND-------------YE 869
Query: 300 EVLQDLLTDN 309
++ D+LT+N
Sbjct: 870 KIALDILTNN 899
>TC80053 similar to PIR|JC7335|JC7335 chitinase (EC 3.2.1.14) 1 - cone shell
(Conus tulipa), partial (81%)
Length = 1081
Score = 141 bits (355), Expect = 4e-34
Identities = 95/280 (33%), Positives = 148/280 (51%), Gaps = 8/280 (2%)
Frame = +2
Query: 14 IIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFIL--GTLREVYSGDGKGKGEFYRTWNF 71
+IFREYIG + D P IN ++E FHFIL G + S G+F W+
Sbjct: 158 LIFREYIGAESNNIKFSDVP---INPNVE-FHFILSFGIDYDTSSSPSPTNGKFNIFWDT 325
Query: 72 NNFSPAKVAKLKKDHKNVKVIISIGGFGAENP---FNPKEIESWSTKAKQSIKKLINEYQ 128
N +P++V+ +K + NVKV +S+GG E F+P +ESW + A S+ K+I EY
Sbjct: 326 KNLNPSQVSSIKSQNPNVKVALSLGGDSVEGGYAYFDPSSVESWLSNAVSSLTKIIKEYN 505
Query: 129 EYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAP 188
+DGIDI+YE+ NP+ F+ CIG LI+ L K++ I SIAP
Sbjct: 506 -----------------LDGIDIDYEHFKGNPNTFAECIGRLIKTL-KANGVITFASIAP 631
Query: 189 --TELLKPHYHKLYWANKDIINWVDYKFY-NQTVSSADELVNLYNKLLNEYGTDVKLLPG 245
+ ++ HY L+ + +I++V+++FY +S + ++ +NK + Y K+L
Sbjct: 632 FDDDQVQSHYLALWKSYGHLIDYVNFQFYAYDKGTSVSQFIDYFNKQSSNYNGG-KVLVS 808
Query: 246 VSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDS 285
+D + + D F K C+ L + L GIFVW+A+DS
Sbjct: 809 FLSDGSGGLSPS-DGFFKACQRLKSQQKLHGIFVWSADDS 925
>BE319726 weakly similar to PIR|S49848|S49 probable narbonin - jack bean,
partial (42%)
Length = 448
Score = 82.8 bits (203), Expect(2) = 5e-27
Identities = 44/101 (43%), Positives = 58/101 (56%)
Frame = +3
Query: 55 YSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWST 114
Y +G G F TWN F P V +LK+++KNVKV+IS GG + PFNP E W+
Sbjct: 168 YDDKKRGTGVFKTTWNVEFFGPEDVKRLKENNKNVKVVISFGGCDEKTPFNPAEDNIWTE 347
Query: 115 KAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEY 155
KA S+K +I +++ S S IIDGIDINYE+
Sbjct: 348 KAVASLKVIILRFKDQSGRS----------IIDGIDINYEH 440
Score = 55.8 bits (133), Expect(2) = 5e-27
Identities = 26/41 (63%), Positives = 30/41 (72%)
Frame = +2
Query: 13 PIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLRE 53
P IFREYIGVK K+L+DFP +IN +I EFHFILG E
Sbjct: 41 PKIFREYIGVKPYSKSLRDFPINIINSNISEFHFILGFASE 163
>AW687533 weakly similar to PIR|T12157|T121 nodulin - fava bean, partial
(30%)
Length = 387
Score = 107 bits (266), Expect = 8e-24
Identities = 57/114 (50%), Positives = 75/114 (65%), Gaps = 9/114 (7%)
Frame = +3
Query: 166 CIGELIRKLKKSSK-SIKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQT--VSSA 222
C+G++I LKK +I +VSIAP+E HY KLYW NKD IN VDYK YNQT V ++
Sbjct: 39 CLGQVITDLKKDRDLNINVVSIAPSEQNDSHYRKLYWENKDNINLVDYKLYNQTKIVQTS 218
Query: 223 DELVNLYNKLLNEYGTDVKLLPGVSTDP------DSNTNMTRDVFIKGCKSLLE 270
+E V LY+K+ N+Y + K LPG+STDP D M R++FI GCK L++
Sbjct: 219 EEFVKLYSKIANDYSPE-KFLPGISTDPGDTEPADKIIKMPREIFIAGCKHLMQ 377
>BF636454 similar to PIR|S69190|S691 myb-related protein 1 - tomato, partial
(14%)
Length = 655
Score = 29.6 bits (65), Expect = 1.5
Identities = 15/54 (27%), Positives = 28/54 (51%)
Frame = +3
Query: 112 WSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSS 165
W++ K+ I+K +S S+ST +++ +NY YSN D+F++
Sbjct: 60 WNSWIKKKIRK-------HSSSSNSTTNNVTQNVVHHSHLNYNYSNIKLDQFAN 200
>CB894613 similar to GP|16226943|gb AT4g27410/F27G19_10 {Arabidopsis
thaliana}, partial (55%)
Length = 657
Score = 28.9 bits (63), Expect = 2.6
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +3
Query: 126 EYQEYSKDSSSTDECHCDDIIDGI 149
EY +YS SSS+ H DD+++ +
Sbjct: 537 EYTQYSNGSSSSSSSHLDDVLESL 608
>TC86417 similar to GP|16226943|gb|AAL16305.1 AT4g27410/F27G19_10
{Arabidopsis thaliana}, partial (67%)
Length = 1628
Score = 28.9 bits (63), Expect = 2.6
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +2
Query: 126 EYQEYSKDSSSTDECHCDDIIDGI 149
EY +YS SSS+ H DD+++ +
Sbjct: 671 EYTQYSNGSSSSSSSHLDDVLESL 742
>TC90598 similar to GP|8953389|emb|CAB96662.1 putative protein {Arabidopsis
thaliana}, partial (18%)
Length = 602
Score = 28.9 bits (63), Expect = 2.6
Identities = 19/49 (38%), Positives = 22/49 (44%)
Frame = +3
Query: 29 LKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPA 77
L DF AE++N DI E E SGDG RTW N + A
Sbjct: 366 LGDFDAELLNTDISETQGDNDV--EKSSGDGNENSLKLRTWQLNMLARA 506
>TC86048 weakly similar to PIR|H96561|H96561 probable peptide transporter
[imported] - Arabidopsis thaliana, partial (47%)
Length = 1364
Score = 27.7 bits (60), Expect = 5.8
Identities = 27/123 (21%), Positives = 52/123 (41%)
Frame = +3
Query: 184 VSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDVKLL 243
V + PT+ KL + NK + K + Q ++S ++N ++ + ++K +
Sbjct: 915 VLVVPTD-------KLRFLNKACV----IKDHEQDIASDGSIINRWSLCTVDQVEELKAI 1061
Query: 244 PGVSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSNEDNTPYFLEEVLQ 303
+ P +T +T + I G LL++ SL + N+N N PY +
Sbjct: 1062 --IKVIPLWSTGITMSINIGGSFGLLQALSLDRHIISNSNFEVPAGIFYNNPYLCNTYMD 1235
Query: 304 DLL 306
+ L
Sbjct: 1236 NYL 1244
>TC77531 similar to PIR|T06562|T06562 CP12 protein precursor chloroplast -
garden pea, partial (52%)
Length = 652
Score = 27.7 bits (60), Expect = 5.8
Identities = 15/39 (38%), Positives = 20/39 (50%), Gaps = 2/39 (5%)
Frame = +2
Query: 108 EIESWSTKAKQSI--KKLINEYQEYSKDSSSTDECHCDD 144
E+E S A + KK + +EY KD+ TDEC D
Sbjct: 329 EVEELSAAASHARDKKKTSDPLEEYCKDNPETDECRTYD 445
>BG456524 similar to GP|9279575|dbj emb|CAB36798.1~gene_id:MLN21.6~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 692
Score = 27.3 bits (59), Expect = 7.6
Identities = 27/101 (26%), Positives = 39/101 (37%), Gaps = 20/101 (19%)
Frame = +2
Query: 215 YNQTVSSADELVNLYNKL------LNEYGTDVKLLPGVSTDPDSNTNMTRDVFIKG---- 264
Y+ TV+S DE+ Y+K L + T V P V DP N + T + +G
Sbjct: 254 YSSTVASGDEIPESYHKKLLSTQPLAKETTVVDNTPVVVDDPSVNDSDTAEKIYQGILAG 433
Query: 265 ----------CKSLLESESLPGIFVWNANDSAMPSNEDNTP 295
L SESL N + + +NE+ P
Sbjct: 434 KSQNGHSQIYANQLSGSESLSPTNAQNHTEKPVITNEEPVP 556
>TC83319 similar to PIR|G86357|G86357 Similar to auxin-independent growth
promoter [imported] - Arabidopsis thaliana, partial
(38%)
Length = 1113
Score = 26.9 bits (58), Expect = 10.0
Identities = 16/44 (36%), Positives = 26/44 (58%)
Frame = -3
Query: 219 VSSADELVNLYNKLLNEYGTDVKLLPGVSTDPDSNTNMTRDVFI 262
+SSA EL+NL++ ++E K+L D S++ +TR FI
Sbjct: 598 ISSARELINLFSSKMSE-----KMLLSCQNDFLSSSGITRVAFI 482
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,485,959
Number of Sequences: 36976
Number of extensions: 151576
Number of successful extensions: 616
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 584
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 585
length of query: 309
length of database: 9,014,727
effective HSP length: 96
effective length of query: 213
effective length of database: 5,465,031
effective search space: 1164051603
effective search space used: 1164051603
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130275.11


