

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.9 - phase: 0
(393 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
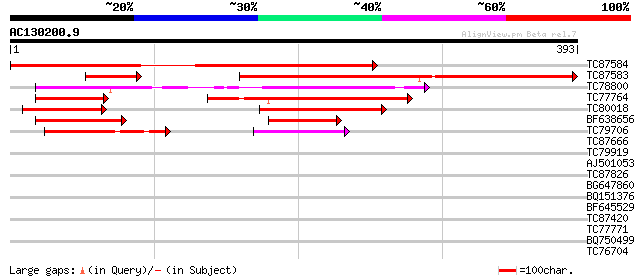
Score E
Sequences producing significant alignments: (bits) Value
TC87584 similar to GP|19715579|gb|AAL91615.1 AT3g24740/K7P8_3 {A... 431 e-121
TC87583 similar to GP|19715579|gb|AAL91615.1 AT3g24740/K7P8_3 {A... 430 e-121
TC78800 similar to GP|15983783|gb|AAL10488.1 AT3g25910/MPE11_6 {... 195 3e-50
TC77764 similar to GP|12324063|gb|AAG51991.1 unknown protein; 39... 136 2e-32
TC80018 similar to GP|18650602|gb|AAL75901.1 AT4g31410/F8F16_230... 98 6e-21
BF638656 weakly similar to GP|21740392|emb OSJNBa0064H22.16 {Ory... 86 3e-17
TC79706 similar to GP|18389230|gb|AAL67058.1 unknown protein {Ar... 77 1e-14
TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger p... 33 0.19
TC79919 similar to GP|3367523|gb|AAC28508.1| ESTs gb|AA728658 an... 32 0.32
AJ501053 similar to GP|17064832|gb putative VP1/ABI3 family regu... 32 0.42
TC87826 similar to SP|Q03211|EXLP_TOBAC Pistil-specific extensin... 30 1.2
BG647860 weakly similar to GP|17065464|gb Unknown protein {Arabi... 30 1.2
BQ151376 28 4.7
BF645529 weakly similar to GP|22475166|gb putative sinapyl alcoh... 28 6.1
TC87420 weakly similar to GP|19347889|gb|AAL86001.1 unknown prot... 28 8.0
TC77771 homologue to GP|6137207|gb|AAF04377.1| P72 DEAD box prot... 28 8.0
BQ750499 similar to OMNI|NT01MC4764 transport protein HasD puta... 28 8.0
TC76704 homologue to SP|Q09134|GRPA_MEDFA Abscisic acid and envi... 28 8.0
>TC87584 similar to GP|19715579|gb|AAL91615.1 AT3g24740/K7P8_3 {Arabidopsis
thaliana}, partial (40%)
Length = 915
Score = 431 bits (1108), Expect = e-121
Identities = 213/255 (83%), Positives = 215/255 (83%)
Frame = +3
Query: 1 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS
Sbjct: 258 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 437
Query: 61 NCLDRFKKLRDNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTL 120
NCLDRFKKLRDNSKENPNLQSSLINTNNSSG
Sbjct: 438 NCLDRFKKLRDNSKENPNLQSSLINTNNSSG----------------------------- 530
Query: 121 LSVGIPQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSL 180
SRQGDAQDPSRHLDQHDEGILETADSENLQD+AVLEEELDVDNSSEDSKSSL
Sbjct: 531 --------SRQGDAQDPSRHLDQHDEGILETADSENLQDKAVLEEELDVDNSSEDSKSSL 686
Query: 181 HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVD 240
HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVD
Sbjct: 687 HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVD 866
Query: 241 PTREQAWQQFERQRE 255
PTREQAW ERQR+
Sbjct: 867 PTREQAWATIERQRD 911
>TC87583 similar to GP|19715579|gb|AAL91615.1 AT3g24740/K7P8_3 {Arabidopsis
thaliana}, partial (41%)
Length = 1220
Score = 430 bits (1105), Expect = e-121
Identities = 217/236 (91%), Positives = 219/236 (91%), Gaps = 2/236 (0%)
Frame = +3
Query: 160 RAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGD 219
RAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGD
Sbjct: 114 RAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGD 293
Query: 220 YLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
YLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVG+
Sbjct: 294 YLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGELCSR 473
Query: 280 NGD--GIGRLPPGGGREGSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHR 337
+G GR GSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHR
Sbjct: 474 KRGWYWVGYHQVVEGR-GSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHR 650
Query: 338 RSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNEDHS 393
RSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNEDHS
Sbjct: 651 RSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNEDHS 818
Score = 85.5 bits (210), Expect = 3e-17
Identities = 39/39 (100%), Positives = 39/39 (100%)
Frame = +1
Query: 53 CDTSYRHSNCLDRFKKLRDNSKENPNLQSSLINTNNSSG 91
CDTSYRHSNCLDRFKKLRDNSKENPNLQSSLINTNNSSG
Sbjct: 1 CDTSYRHSNCLDRFKKLRDNSKENPNLQSSLINTNNSSG 117
>TC78800 similar to GP|15983783|gb|AAL10488.1 AT3g25910/MPE11_6 {Arabidopsis
thaliana}, partial (37%)
Length = 1463
Score = 195 bits (495), Expect = 3e-50
Identities = 104/275 (37%), Positives = 159/275 (57%), Gaps = 2/275 (0%)
Frame = +1
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK--LRDNSKEN 76
+E +E CP+CM+HPHNAVLL+CSSH+KGCR Y+C+TSYRHSNCLD+F K +
Sbjct: 301 KEWEEARCPVCMEHPHNAVLLICSSHEKGCRPYMCNTSYRHSNCLDQFCKSFAEPSPPTV 480
Query: 77 PNLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQD 136
P ++S + N+++ S + N +V E + + T++ V
Sbjct: 481 PQVESEISNSDSPQDQSTEANIV------NVQEETIVDVQEETIVDVQ------------ 606
Query: 137 PSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVE 196
++ EG + T S + +D ++KS L CPLCRG + W+VVE
Sbjct: 607 -----EETSEGFV-TMQSLSCED---------------ETKSKLVCPLCRGHIKEWKVVE 723
Query: 197 EARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREY 256
AR+++N+K RSCS +SC+F G Y +LR+HAR HP RPS VDP R++ W++ ERQR+
Sbjct: 724 GARHFMNDKSRSCSCESCNFTGTYTDLRKHARVEHPLERPSAVDPERQRNWRRLERQRDL 903
Query: 257 GDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGG 291
GD++S +Q++ G+ +++G G+ + GG
Sbjct: 904 GDLLSTLQNSF------GENRVDDGLGLAPIDDGG 990
>TC77764 similar to GP|12324063|gb|AAG51991.1 unknown protein; 3976-4980
{Arabidopsis thaliana}, partial (54%)
Length = 1777
Score = 136 bits (342), Expect = 2e-32
Identities = 70/150 (46%), Positives = 91/150 (60%), Gaps = 8/150 (5%)
Frame = +3
Query: 138 SRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKS--------SLHCPLCRGTV 189
S LDQ+ + + + N Q +E +D DS S L CPLCRG V
Sbjct: 531 SNCLDQYKKAYTKVVSARNGQP---VEGSIDNPFMFHDSNSPHEKNEVTELACPLCRGQV 701
Query: 190 LGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQ 249
GW VVE R++LN KKRSC +D+CSF G+Y EL++H R HP++RP VDP EQ W+
Sbjct: 702 KGWTVVEPVRDFLNEKKRSCMQDNCSFVGNYKELKKHVRAEHPSARPRTVDPDHEQKWRW 881
Query: 250 FERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
E +RE D++S + SAIPGAVV GDYV+E
Sbjct: 882 LEWEREREDVISTVTSAIPGAVVFGDYVIE 971
Score = 89.4 bits (220), Expect = 2e-18
Identities = 35/50 (70%), Positives = 45/50 (90%)
Frame = +3
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK 68
+E ++V+C +CM++PHNAVLLLCSSHDKGCR Y+C TS RHSNCLD++KK
Sbjct: 408 KECEDVTCSVCMEYPHNAVLLLCSSHDKGCRPYMCGTSLRHSNCLDQYKK 557
>TC80018 similar to GP|18650602|gb|AAL75901.1 AT4g31410/F8F16_230
{Arabidopsis thaliana}, partial (44%)
Length = 750
Score = 97.8 bits (242), Expect = 6e-21
Identities = 43/88 (48%), Positives = 59/88 (66%)
Frame = +1
Query: 174 EDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPT 233
ED +S+L CPLCRG V GW VV++AR +L++KKR C C F G YLEL+ HA+ HP
Sbjct: 487 EDGQSNLTCPLCRGQVSGWIVVDKARTHLDDKKRCCDEVKCKFMGSYLELQNHAQIEHPH 666
Query: 234 SRPSDVDPTREQAWQQFERQREYGDIVS 261
+ PS +DP R W+ F++ E D++S
Sbjct: 667 ACPSKIDPARVLDWEDFQQSSEIIDVLS 750
Score = 82.4 bits (202), Expect = 3e-16
Identities = 36/58 (62%), Positives = 43/58 (74%)
Frame = +1
Query: 10 NDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
N+ D L+ D+V CPIC+D PHN VLL CSS+DKGCR+ +CDT HSNCLDRFK
Sbjct: 241 NNMDDIQLNIVWDDVVCPICLDIPHNCVLLKCSSYDKGCRALVCDTDQSHSNCLDRFK 414
>BF638656 weakly similar to GP|21740392|emb OSJNBa0064H22.16 {Oryza sativa},
partial (32%)
Length = 649
Score = 85.5 bits (210), Expect = 3e-17
Identities = 36/63 (57%), Positives = 48/63 (76%)
Frame = +3
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPN 78
+E D+ CPICM+ PHN VLL CSS++KGCR Y+C+TSYRHSNCLD+F K D+ +
Sbjct: 216 KEWDDTRCPICMEIPHNTVLLKCSSYEKGCRPYMCNTSYRHSNCLDQFCKSFDSHLSSAM 395
Query: 79 LQS 81
L++
Sbjct: 396 LEA 404
Score = 64.7 bits (156), Expect = 6e-11
Identities = 29/51 (56%), Positives = 35/51 (67%)
Frame = +3
Query: 180 LHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRV 230
L CPLCRG + G+ V E AR Y+N KKRSCS ++C F G Y EL +HA V
Sbjct: 495 LICPLCRGGIYGYMVSEPARRYMNCKKRSCSSETCEFQGTYXELXKHAXLV 647
>TC79706 similar to GP|18389230|gb|AAL67058.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 676
Score = 77.0 bits (188), Expect = 1e-14
Identities = 41/87 (47%), Positives = 58/87 (66%)
Frame = +3
Query: 25 SCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPNLQSSLI 84
+C +CM+ PHNA+LLLCSS++KGCR Y+C TS R+SNC +++KK + ++QSS
Sbjct: 270 TCSVCMEVPHNAILLLCSSYNKGCRPYMCATSRRYSNCFEQYKKAYTKA---TSVQSSQQ 440
Query: 85 NTNNSSGNSFDINFTMQSDMHDVDELL 111
T+ S+ NS N +SD V ELL
Sbjct: 441 ETDYSNFNS---NSGDRSDNAKVPELL 512
Score = 76.3 bits (186), Expect = 2e-14
Identities = 37/66 (56%), Positives = 39/66 (59%)
Frame = +3
Query: 170 DNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARR 229
D S L CPLCR V GW VVE AR LN KKRSC +D CSFAG Y EL +H R
Sbjct: 477 DRSDNAKVPELLCPLCRRQVKGWTVVEAARKSLNGKKRSCMQDGCSFAGSYKELXKHVRS 656
Query: 230 VHPTSR 235
H SR
Sbjct: 657 KHXCSR 674
>TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger protein
{Oryza sativa}, partial (20%)
Length = 1828
Score = 33.1 bits (74), Expect = 0.19
Identities = 25/99 (25%), Positives = 40/99 (40%), Gaps = 2/99 (2%)
Frame = +2
Query: 1 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRH- 59
M G C + R ++++CP C + N +++ Y C T R+
Sbjct: 635 MEGLSPNSCPRQGLEKKARPQEQINCPRC--NSTNTKFCYYNNYSLTQPRYFCKTCRRYW 808
Query: 60 -SNCLDRFKKLRDNSKENPNLQSSLINTNNSSGNSFDIN 97
R + S++N + SSL N NNSS D+N
Sbjct: 809 TEGGSLRNVPVGGGSRKNKKIPSSLANPNNSSSKIPDLN 925
>TC79919 similar to GP|3367523|gb|AAC28508.1| ESTs gb|AA728658 and gb|N95943
come from this gene. {Arabidopsis thaliana}, partial
(64%)
Length = 1714
Score = 32.3 bits (72), Expect = 0.32
Identities = 40/180 (22%), Positives = 76/180 (42%), Gaps = 27/180 (15%)
Frame = +1
Query: 40 LCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPNLQSS---LINTNNSSGNSFDI 96
LCS + +S + T+ +C F N P L+++ LI ++ N+ +
Sbjct: 310 LCSLNFTNAKSLLSVTAI---DCWGFFAPFVANIMCCPQLEATVTVLIGQSSKHTNALAL 480
Query: 97 NFTMQSD-MHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLDQHDEGI------L 149
N T+ + DV+++L+ Q L + S +A P +H+++ ++ + L
Sbjct: 481 NGTVAKHCLSDVEQILMGQGASGDLRQICSISSSNLTEASCPVKHVNEFNDMVDTSKLLL 660
Query: 150 ETADSENLQD-------RAVLEEE----------LDVDNSSEDSKSSLHCPLCRGTVLGW 192
AD + +++ A+LE LD+D S + + S+ CR VL W
Sbjct: 661 ACADIDPVKECCYQVCHNAILEAATAIASKGSHVLDLDASHDLPEHSIRVNDCRNIVLRW 840
>AJ501053 similar to GP|17064832|gb putative VP1/ABI3 family regulatory
protein {Arabidopsis thaliana}, partial (13%)
Length = 682
Score = 32.0 bits (71), Expect = 0.42
Identities = 24/95 (25%), Positives = 35/95 (36%)
Frame = +3
Query: 15 HALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSK 74
H HR +C +C+ P HD C C+T R L KK
Sbjct: 345 HPRHRP--GCTCIVCIQPPSGQ-----GKHDPTCTCLACETLKRRFKSLTMRKKKNQLES 503
Query: 75 ENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDE 109
E Q++ +N + +G S + + Q H DE
Sbjct: 504 EAVADQNNQVNHRDEAGTS--VGASRQDTSHSTDE 602
>TC87826 similar to SP|Q03211|EXLP_TOBAC Pistil-specific extensin-like
protein precursor (PELP). [Common tobacco] {Nicotiana
tabacum}, partial (6%)
Length = 1222
Score = 30.4 bits (67), Expect = 1.2
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = -3
Query: 263 IQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGN 301
I+ + + G+Y+ G G+ + P GGG EG +G GN
Sbjct: 707 IEEVLRKQIQKGEYLDNGGSGV-KPPGGGGGEGGSGGGN 594
>BG647860 weakly similar to GP|17065464|gb Unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 719
Score = 30.4 bits (67), Expect = 1.2
Identities = 25/106 (23%), Positives = 47/106 (43%), Gaps = 8/106 (7%)
Frame = +3
Query: 83 LINTNNSSGNSFDINFTMQSD-MHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHL 141
L +T +S + F+M SD MHD +++ ++ + + P D D + +
Sbjct: 18 LYSTTFPKIHSIRVFFSMDSDDMHDANDVESLDDDFYSGETEDAPMDDYSDDYDDDNNNN 197
Query: 142 -------DQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSL 180
D D+G ++A S + + + +E D+ + ED SSL
Sbjct: 198 NNNDADDDYFDDGADDSAQSRSTEINFAILKESDIRDRQEDDISSL 335
>BQ151376
Length = 403
Score = 28.5 bits (62), Expect = 4.7
Identities = 14/43 (32%), Positives = 21/43 (48%)
Frame = -1
Query: 324 EPRARSSNGWSRHRRSSDRRRYLWGENLLGLQDNEVEEDLRIF 366
E R + G + +R D+RR WG L D ++ E LR +
Sbjct: 325 ENRKKIKRG*KKKKRDKDKRRIYWGFFLFTYYDIKIVEYLRYY 197
>BF645529 weakly similar to GP|22475166|gb putative sinapyl alcohol
dehydrogenase {Populus tremula x Populus tremuloides},
partial (40%)
Length = 658
Score = 28.1 bits (61), Expect = 6.1
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 4/54 (7%)
Frame = -2
Query: 29 CMDHPHNAVLLLCSSHDKGC---RSYIC-DTSYRHSNCLDRFKKLRDNSKENPN 78
C+ H N SSH + C R Y C T+ HSN LD F + + PN
Sbjct: 480 CVLHICNLRPTFVSSHHR*CYHPRKYHCIQTATSHSNSLDLFCSTHSPNMKQPN 319
>TC87420 weakly similar to GP|19347889|gb|AAL86001.1 unknown protein
{Arabidopsis thaliana}, partial (47%)
Length = 1303
Score = 27.7 bits (60), Expect = 8.0
Identities = 23/77 (29%), Positives = 36/77 (45%), Gaps = 1/77 (1%)
Frame = +2
Query: 106 DVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLDQHDEGILETAD-SENLQDRAVLE 164
++D L + IN +S +P G Q A+ R + + G+ + SE R
Sbjct: 542 NLDSKLNVPSTINMPVSNYMPSGDIQ--ARIVGRTNEVKENGVADNYGYSEQRIQRGPDS 715
Query: 165 EELDVDNSSEDSKSSLH 181
E + DN++EDS SLH
Sbjct: 716 EHIREDNAAEDSNGSLH 766
>TC77771 homologue to GP|6137207|gb|AAF04377.1| P72 DEAD box protein {Pisum
sativum}, partial (69%)
Length = 1736
Score = 27.7 bits (60), Expect = 8.0
Identities = 12/27 (44%), Positives = 14/27 (51%)
Frame = -3
Query: 26 CPICMDHPHNAVLLLCSSHDKGCRSYI 52
C IC P CSSH KGC++ I
Sbjct: 198 CRICWGIPWTHCGSCCSSHSKGCKNTI 118
>BQ750499 similar to OMNI|NT01MC4764 transport protein HasD putative
{Magnetococcus sp. MC-1}, partial (3%)
Length = 800
Score = 27.7 bits (60), Expect = 8.0
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = +3
Query: 34 HNAVLLLCSSHDKGCRSYICD 54
HN VL+LCSS+ GC + CD
Sbjct: 3 HNIVLILCSSYG-GCIKWSCD 62
>TC76704 homologue to SP|Q09134|GRPA_MEDFA Abscisic acid and environmental
stress inducible protein. [Sickle medic] {Medicago
falcata}, partial (81%)
Length = 771
Score = 27.7 bits (60), Expect = 8.0
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = -2
Query: 29 CMDHPHNAVLLLCSSHDKGCRSYICD 54
C HPH LL +H +GC + C+
Sbjct: 275 CCIHPHRGCNLLHCNHHRGCNLHHCN 198
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,973,723
Number of Sequences: 36976
Number of extensions: 203718
Number of successful extensions: 1301
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1276
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1297
length of query: 393
length of database: 9,014,727
effective HSP length: 98
effective length of query: 295
effective length of database: 5,391,079
effective search space: 1590368305
effective search space used: 1590368305
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130200.9


