

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
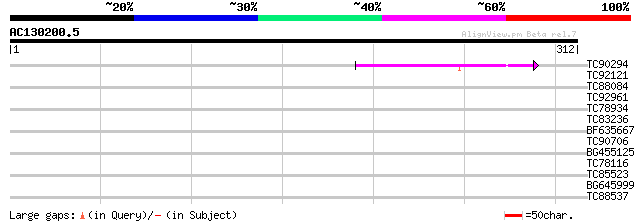
Query= AC130200.5 - phase: 0
(312 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90294 similar to PIR|T45759|T45759 MS5-like protein - Arabidop... 44 1e-04
TC92121 weakly similar to GP|10177302|dbj|BAB10563. gene_id:MDC1... 36 0.017
TC88084 similar to GP|13605619|gb|AAK32803.1 AT3g49260/F2K15_120... 34 0.063
TC92961 similar to GP|22135986|gb|AAM91575.1 unknown protein {Ar... 32 0.41
TC78934 similar to GP|14189890|dbj|BAB55874. response regulator ... 31 0.70
TC83236 homologue to PIR|T06779|T06779 clathrin heavy chain - so... 30 0.92
BF635667 weakly similar to PIR|C85359|C853 hypothetical protein ... 30 1.2
TC90706 similar to PIR|E84825|E84825 probable protein kinase [im... 30 1.6
BG455125 similar to GP|21618178|gb unknown {Arabidopsis thaliana... 28 3.5
TC78116 similar to PIR|F86441|F86441 probable cytochrome P450 [i... 28 4.5
TC85523 similar to GP|10177098|dbj|BAB10432. gene_id:MXC20.6~unk... 28 5.9
BG645999 similar to GP|16930745|gb NADH-ubiquinone oxidoreductas... 28 5.9
TC88537 weakly similar to GP|21536818|gb|AAM61150.1 F-box protei... 27 7.7
>TC90294 similar to PIR|T45759|T45759 MS5-like protein - Arabidopsis
thaliana, partial (31%)
Length = 1124
Score = 43.5 bits (101), Expect = 1e-04
Identities = 30/106 (28%), Positives = 50/106 (46%), Gaps = 5/106 (4%)
Frame = +2
Query: 191 NYMMAEVVFKKAQMIDADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEI---- 246
NY+ AE +++A + D NK NL +CLM+Q R EA L +V + +
Sbjct: 2 NYIEAEEAYRRALCLAPDNNKMCNLGICLMKQGRIAEAKETLHRVKPAVIRWTK*AQILI 181
Query: 247 -KSRNRAEELLVELSANLPQPKFMDDLGLDDDLLKGIDGLLNVWSP 291
K RA+++L +L + + +D L L + G ++W P
Sbjct: 182 *KPSERAQQMLKDLESEMMNRGGVDRLE-QSRLFEAFLGSSSIWQP 316
>TC92121 weakly similar to GP|10177302|dbj|BAB10563.
gene_id:MDC12.18~pir||F69210~similar to unknown protein
{Arabidopsis thaliana}, partial (76%)
Length = 845
Score = 36.2 bits (82), Expect = 0.017
Identities = 22/70 (31%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Frame = +3
Query: 169 VSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKAL-NLALCLMRQSRYEE 227
++ KQE +L N+G Y Q+ Y +A+ +F K+ + A NL L + + EE
Sbjct: 339 LATKQEAHAILSNMGILYRQQKKYELAKAMFTKSLELQPGYAPAFNNLGLVFIAEGLLEE 518
Query: 228 AYLILEQVLQ 237
A E+ LQ
Sbjct: 519 AKHCFEKALQ 548
>TC88084 similar to GP|13605619|gb|AAK32803.1 AT3g49260/F2K15_120
{Arabidopsis thaliana}, partial (33%)
Length = 1706
Score = 34.3 bits (77), Expect = 0.063
Identities = 33/152 (21%), Positives = 67/152 (43%), Gaps = 1/152 (0%)
Frame = +1
Query: 75 AGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEE 134
AG +S ++ A++++ R A A+++ +GL V L + V +
Sbjct: 514 AGHGRNSKVEKAAILIQSYYRGYLARRALRALKGL-----------VRLQALVRGHNVRK 660
Query: 135 QIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMM 194
Q ++ R ++ + + +A R + +HGK + ++Q L GW Y ++++ +
Sbjct: 661 QAQMTMRCMQALVRVQA-RVRARRLQLTHGKHERTVVEQHPTTKLDTNGWDYRRQSSQKI 837
Query: 195 AEVVFKKAQMIDADANKALNLAL-CLMRQSRY 225
+ F+K + KAL A C Q +Y
Sbjct: 838 KDTDFRK-HGTTMNKEKALPYAFNCQQLQKQY 930
>TC92961 similar to GP|22135986|gb|AAM91575.1 unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 842
Score = 31.6 bits (70), Expect = 0.41
Identities = 19/61 (31%), Positives = 34/61 (55%), Gaps = 1/61 (1%)
Frame = +1
Query: 11 GHLCVLEMEGISVLKKSSKGKKEDLYH-VIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYF 69
G+ VL + I +++ + +K+ L+ ++H + +G T +K +H Q V AIVYF
Sbjct: 445 GNQLVLLQQMIQIMEILYRRQKQWLFEELVHALGWGCTFIMKKRHKQAVFCVTLTAIVYF 624
Query: 70 W 70
W
Sbjct: 625 W 627
>TC78934 similar to GP|14189890|dbj|BAB55874. response regulator 9 {Zea
mays}, partial (25%)
Length = 1014
Score = 30.8 bits (68), Expect = 0.70
Identities = 31/127 (24%), Positives = 58/127 (45%), Gaps = 17/127 (13%)
Frame = +2
Query: 187 MQKTNYMMAEVVFKKAQMIDADANKALNLA-----LCLMRQSRYEEAYLILEQVLQG--- 238
M + N M ++V + M D D K L L L ++ S + E L+++ + G
Sbjct: 176 MLQENIDMFDLVIAEVHMPDMDGLKLLELVGLEMDLPVIMLSAHGETELVMKAISHGARD 355
Query: 239 ---KLPGSDEIKS------RNRAEELLVELSANLPQPKFMDDLGLDDDLLKGIDGLLNVW 289
K +E+++ RN+ + + + + + ++ LG+D + K I L+NV
Sbjct: 356 FLLKPVRLEELRNIWQHVIRNKESQFVWSVELHRKFLETVNQLGVDKAVPKKIFDLMNVE 535
Query: 290 SPTRSRR 296
+ TR RR
Sbjct: 536 NITRGRR 556
>TC83236 homologue to PIR|T06779|T06779 clathrin heavy chain - soybean,
partial (12%)
Length = 635
Score = 30.4 bits (67), Expect = 0.92
Identities = 20/68 (29%), Positives = 33/68 (48%), Gaps = 5/68 (7%)
Frame = +2
Query: 79 VDSALKDMAVVMKQLDRAEEAIEAIKSFRGL-----CNKHSQESLDNVLLDLYKKCGRVE 133
V+ AL ++ V + DR E+I+ +F + KH + V +YKK GR +
Sbjct: 233 VNEALNEIYVEEEDYDRLRESIDLHDNFDQIGLAQKIEKHELLEMRRVAAYIYKKAGRWK 412
Query: 134 EQIELLKR 141
+ I L K+
Sbjct: 413 QPIALSKK 436
>BF635667 weakly similar to PIR|C85359|C853 hypothetical protein AT4g30700
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 30.0 bits (66), Expect = 1.2
Identities = 10/20 (50%), Positives = 18/20 (90%)
Frame = +3
Query: 122 LLDLYKKCGRVEEQIELLKR 141
++DL+ +CGR+EE ++LL+R
Sbjct: 114 MVDLFSRCGRLEEALQLLER 173
>TC90706 similar to PIR|E84825|E84825 probable protein kinase [imported] -
Arabidopsis thaliana, partial (38%)
Length = 756
Score = 29.6 bits (65), Expect = 1.6
Identities = 26/113 (23%), Positives = 49/113 (43%), Gaps = 1/113 (0%)
Frame = +3
Query: 18 MEGISVLKKSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAIN-AG 76
++ I +LK +K D +H++ Y + +H +V + + F K +G
Sbjct: 177 LDEIKLLKLVNKHDPGDKHHILRLYDY----FYHQEHLFIVTELLRANLYEFQKFNQESG 344
Query: 77 DKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKC 129
+ LK + ++ +Q +EA++ L H +NVL+ YKKC
Sbjct: 345 GEAYFTLKRLQLITRQ------CLEALQYLHNLGIVHCDLKPENVLIKSYKKC 485
>BG455125 similar to GP|21618178|gb unknown {Arabidopsis thaliana}, partial
(52%)
Length = 638
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/47 (38%), Positives = 25/47 (52%), Gaps = 2/47 (4%)
Frame = +2
Query: 40 HKVPYGDTPYVKAKHAQLVDKDPEVAIVYFW--KAINAGDKVDSALK 84
++ P GD P+V A + QL KD A+ YF + I G +D LK
Sbjct: 338 NEFPDGDRPFVWAFNLQLPTKDNYSAVAYFTTKEPIAEGSVMDRFLK 478
>TC78116 similar to PIR|F86441|F86441 probable cytochrome P450 [imported] -
Arabidopsis thaliana, partial (80%)
Length = 1547
Score = 28.1 bits (61), Expect = 4.5
Identities = 18/72 (25%), Positives = 34/72 (47%), Gaps = 6/72 (8%)
Frame = +3
Query: 232 LEQVLQGKLPGSDEIKSRNRAEELLVELSANLPQPKFMDDLGLDDDLL------KGIDGL 285
++ VL + P +++K ++ E PQP + ++DD+L +G D
Sbjct: 1050 VDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIF 1229
Query: 286 LNVWSPTRSRRL 297
++VW+ RS L
Sbjct: 1230 ISVWNLHRSPTL 1265
>TC85523 similar to GP|10177098|dbj|BAB10432. gene_id:MXC20.6~unknown
protein {Arabidopsis thaliana}, partial (89%)
Length = 871
Score = 27.7 bits (60), Expect = 5.9
Identities = 19/57 (33%), Positives = 31/57 (54%)
Frame = +3
Query: 90 MKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
+K++ E +A++SF K QE D ++ CG+VEE IE + +L+LI
Sbjct: 207 IKKVPEDEGYRKAVESFTSHRLKVCQEEEDWEKIENRLGCGQVEELIEEAQDELKLI 377
>BG645999 similar to GP|16930745|gb NADH-ubiquinone oxidoreductase {Retama
raetam}, partial (84%)
Length = 742
Score = 27.7 bits (60), Expect = 5.9
Identities = 19/57 (33%), Positives = 31/57 (54%)
Frame = +3
Query: 90 MKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
+K++ E +A++SF K QE D ++ CG+VEE IE + +L+LI
Sbjct: 81 IKKVPEDEGYRKAVESFTSHRLKVCQEEEDWEKIENRLGCGQVEELIEEAQDELKLI 251
>TC88537 weakly similar to GP|21536818|gb|AAM61150.1 F-box protein family
AtFBW2 {Arabidopsis thaliana}, partial (52%)
Length = 1395
Score = 27.3 bits (59), Expect = 7.7
Identities = 17/60 (28%), Positives = 26/60 (43%)
Frame = +1
Query: 102 AIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTAR 161
A F G C +S + +LLDL + G + EL+ L +I+ + R T R
Sbjct: 217 ACAGFFGSCFHNSPPGTNTLLLDLMGEAGELRSWDELIPDALGVIFTNLSLKERVTVIPR 396
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,968,779
Number of Sequences: 36976
Number of extensions: 87837
Number of successful extensions: 418
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 418
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 418
length of query: 312
length of database: 9,014,727
effective HSP length: 96
effective length of query: 216
effective length of database: 5,465,031
effective search space: 1180446696
effective search space used: 1180446696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130200.5


