

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
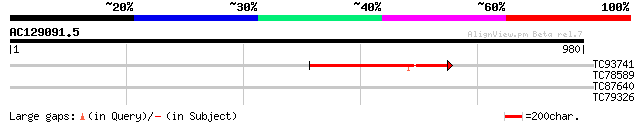
Query= AC129091.5 - phase: 0 /pseudo
(980 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93741 similar to GP|22165058|gb|AAM93675.1 putative glycine-ri... 308 5e-84
TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago trun... 32 1.2
TC87640 similar to PIR|C96618|C96618 probable squalene monooxyge... 31 2.7
TC79326 homologue to GP|14190381|gb|AAK55671.1 AT5g11200/F2I11_9... 29 7.8
>TC93741 similar to GP|22165058|gb|AAM93675.1 putative glycine-rich protein
{Oryza sativa (japonica cultivar-group)}, partial (20%)
Length = 877
Score = 308 bits (790), Expect = 5e-84
Identities = 173/293 (59%), Positives = 193/293 (65%), Gaps = 48/293 (16%)
Frame = +2
Query: 513 SLLVGVDWWDVVGCTQRAAEEGIVSVNGVIAVLDADFHSLPTAQHRQQYCLGLDMIKCRL 572
SLLVGVDWWD VGCTQ AAE+GIVS+N VIAVLDADFHSLP+ QHRQQY LD IKCRL
Sbjct: 2 SLLVGVDWWDAVGCTQSAAEDGIVSLNSVIAVLDADFHSLPSVQHRQQYGPSLDRIKCRL 181
Query: 573 LVGSNAQEVRATVLDMQARVLLDMLGKGIESALINPSALLPDPWQASEEILSNFDMEEMA 632
L GSNAQEVRA VLDMQAR+LLDMLG+GIESALI+P+ +P+ WQAS E LSN D E MA
Sbjct: 182 LEGSNAQEVRAMVLDMQARLLLDMLGRGIESALIDPTQFVPELWQASGETLSNIDSETMA 361
Query: 633 VEPELTPCIQAYVDSVIDLASHL-ITRLRHYAKIFRTLANQAVTVASGS----------- 680
VEP L PC+QAYV + L ITRLR YA RTLA+ AVT +GS
Sbjct: 362 VEPALVPCVQAYV*CNS*FSFTLCITRLRRYASFCRTLASHAVTAGTGSNRNMVASPTQS 541
Query: 681 ------------------------------------TSGGVPNLTLNPSISGPSSLMLIS 704
T+ G N T NP ISGPSS M IS
Sbjct: 542 SATPATSQGGQNGSTSSMGTVQLQTWVQGAIAKISNTTEGGSNPTPNP-ISGPSSFMPIS 718
Query: 705 INTGTFPGTPAVRLIGDCHFLHRLCQLLFFCFFFKRSQLARYKSGLRRTAETS 757
INTGTFPGTPAVRLIGDC FLHRLCQLL FCFFF+R+Q+ RY RT +++
Sbjct: 719 INTGTFPGTPAVRLIGDCQFLHRLCQLLLFCFFFRRTQIPRYMGAANRTNDSN 877
>TC78589 1-deoxy-D-xylulose 5-phosphate synthase 2 [Medicago truncatula]
Length = 2760
Score = 32.0 bits (71), Expect = 1.2
Identities = 22/78 (28%), Positives = 31/78 (39%), Gaps = 5/78 (6%)
Frame = -3
Query: 453 SISVPAFSATSCCSASVWHDTSKGEAMLKIIRVLPPPFPIGQEKATSSTWEHAISERFWF 512
S + PA + T+CCSA G K R +PPP S I+E +
Sbjct: 1552 SAANPAANVTACCSAMPTSKHRSGNLFWK*FRPVPPPIAA*IATILLSISASFINESAKY 1373
Query: 513 SLLV-----GVDWWDVVG 525
+ G++WW V G
Sbjct: 1372 CVYASVEGFGLNWWPVFG 1319
>TC87640 similar to PIR|C96618|C96618 probable squalene monooxygenase
F9K23.3 [imported] - Arabidopsis thaliana, partial (55%)
Length = 1288
Score = 30.8 bits (68), Expect = 2.7
Identities = 25/98 (25%), Positives = 44/98 (44%), Gaps = 3/98 (3%)
Frame = +1
Query: 3 PMKESEAEAEAAATVNDPMMEEEEVEAGTVFDISLEQSPSTSL--HKITVHESMRSNFSA 60
P+++ +++ + N ++ +A T+ + LEQ TSL T+ N S
Sbjct: 637 PLEKFDSDVSGRSFHNGRFIQRMREKASTIPTVKLEQGTVTSLLEENGTIKGVNYKNKSG 816
Query: 61 VAWCAKLNV-IACATETCVNGIPRSSVNPWFWIPIHIV 97
+ AK + I C + C + + RS NP +P H V
Sbjct: 817 QEFTAKAPLTIVC--DGCFSNLRRSLCNPKVEVPSHFV 924
>TC79326 homologue to GP|14190381|gb|AAK55671.1 AT5g11200/F2I11_90
{Arabidopsis thaliana}, partial (94%)
Length = 1883
Score = 29.3 bits (64), Expect = 7.8
Identities = 19/57 (33%), Positives = 23/57 (40%), Gaps = 10/57 (17%)
Frame = -3
Query: 72 CATETCVN----------GIPRSSVNPWFWIPIHIVIPERPTEIAAFNVVADSSLDS 118
C +E C N G R S+NP WIP P+ P +A A SS S
Sbjct: 318 CTSEGCSNPESTIARRSSGFRRKSLNPELWIPT*PFFPDSPLTLAPTESGAFSSSSS 148
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,924,483
Number of Sequences: 36976
Number of extensions: 511520
Number of successful extensions: 3355
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2480
Number of HSP's successfully gapped in prelim test: 100
Number of HSP's that attempted gapping in prelim test: 815
Number of HSP's gapped (non-prelim): 2672
length of query: 980
length of database: 9,014,727
effective HSP length: 105
effective length of query: 875
effective length of database: 5,132,247
effective search space: 4490716125
effective search space used: 4490716125
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC129091.5


