

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.3 + phase: 0
(170 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
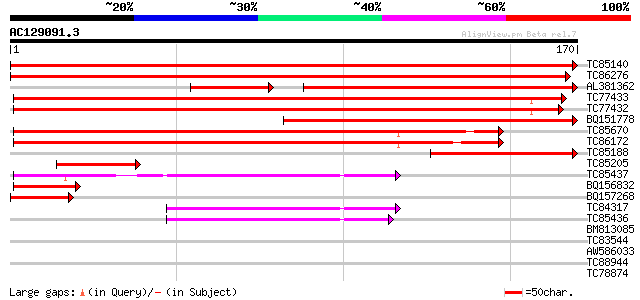
Score E
Sequences producing significant alignments: (bits) Value
TC85140 similar to PIR|T08937|T08937 hypothetical protein F27G19... 354 8e-99
TC86276 similar to GP|9857931|gb|AAG00940.1| unknown {Glycine ma... 261 7e-71
AL381362 weakly similar to GP|643469|gb|AA unknown {Lycopersicon... 175 6e-56
TC77433 similar to GP|20340249|gb|AAM19711.1 aluminum-induced pr... 193 2e-50
TC77432 similar to GP|18252901|gb|AAL62377.1 unknown protein {Ar... 190 3e-49
BQ151778 weakly similar to GP|643469|gb|AA unknown {Lycopersicon... 187 2e-48
TC85670 homologue to GP|2970051|dbj|BAA25187.1 ARG10 {Vigna radi... 141 1e-34
TC86172 similar to GP|2970051|dbj|BAA25187.1 ARG10 {Vigna radiat... 134 2e-32
TC85188 similar to GP|643469|gb|AAA61967.1|| unknown {Lycopersic... 93 5e-20
TC85205 similar to GP|4559292|gb|AAD22970.1| trehalase 1 GMTRE1 ... 55 1e-08
TC85437 homologue to PIR|S11443|AJPMN2 asparagine synthase (glut... 48 2e-06
BQ156832 homologue to PIR|F86214|F862 protein T6D22.2 [imported]... 46 8e-06
BQ157268 homologue to PIR|F86214|F862 protein T6D22.2 [imported]... 43 5e-05
TC84317 similar to GP|3859536|gb|AAC72837.1| asparagine syntheta... 43 5e-05
TC85436 GP|2522320|gb|AAB81011.1| asparagine synthetase {Medicag... 43 5e-05
BM813085 weakly similar to GP|15795110|dbj gene_id:MJK13.11~unkn... 34 0.033
TC83544 weakly similar to GP|15795110|dbj|BAB02374. gene_id:MJK1... 30 0.47
AW586033 similar to GP|14030723|gb| AT4g27450/F27G19_50 {Arabido... 30 0.61
TC88944 type IIB calcium ATPase [Medicago truncatula] 27 3.0
TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.... 27 5.2
>TC85140 similar to PIR|T08937|T08937 hypothetical protein F27G19.50 -
Arabidopsis thaliana, partial (63%)
Length = 1022
Score = 354 bits (909), Expect = 8e-99
Identities = 170/170 (100%), Positives = 170/170 (100%)
Frame = +2
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK
Sbjct: 326 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 505
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN
Sbjct: 506 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 685
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS 170
FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS
Sbjct: 686 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS 835
>TC86276 similar to GP|9857931|gb|AAG00940.1| unknown {Glycine max}, partial
(97%)
Length = 1336
Score = 261 bits (668), Expect = 7e-71
Identities = 123/168 (73%), Positives = 143/168 (84%)
Frame = +3
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+L+NLS LNKQYGLSKG +EAMF+IEAYRTLRDRGPYP DQV+K L+GSFAFV+YD
Sbjct: 387 LGSLNNLSLLNKQYGLSKGTDEAMFLIEAYRTLRDRGPYPPDQVVKELDGSFAFVVYDST 566
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
GTVFAA GSDG + LYWGIAADGSVVIS++L +++ CAKSFAPFP GC+FHSE GL++
Sbjct: 567 FGTVFAALGSDGGLKLYWGIAADGSVVISDDLNVIQEGCAKSFAPFPAGCMFHSEGGLMS 746
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
FEHP K+KAMPRIDSEGVMCGANF VD SR +PRVGSE+NW W
Sbjct: 747 FEHPLNKLKAMPRIDSEGVMCGANFKVDKFSRVNSIPRVGSESNWMEW 890
>AL381362 weakly similar to GP|643469|gb|AA unknown {Lycopersicon
esculentum}, partial (33%)
Length = 518
Score = 175 bits (443), Expect(2) = 6e-56
Identities = 81/82 (98%), Positives = 82/82 (99%)
Frame = +3
Query: 89 SENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVD 148
+ENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVD
Sbjct: 108 TENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVD 287
Query: 149 SQSRNQMMPRVGSEANWATWGS 170
SQSRNQMMPRVGSEANWATWGS
Sbjct: 288 SQSRNQMMPRVGSEANWATWGS 353
Score = 58.9 bits (141), Expect(2) = 6e-56
Identities = 25/25 (100%), Positives = 25/25 (100%)
Frame = +1
Query: 55 VIYDHKDGTVFAASGSDGHIGLYWG 79
VIYDHKDGTVFAASGSDGHIGLYWG
Sbjct: 1 VIYDHKDGTVFAASGSDGHIGLYWG 75
>TC77433 similar to GP|20340249|gb|AAM19711.1 aluminum-induced protein-like
protein {Thellungiella halophila}, partial (96%)
Length = 938
Score = 193 bits (491), Expect = 2e-50
Identities = 87/168 (51%), Positives = 121/168 (71%), Gaps = 2/168 (1%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G++ N++ L +QYGL+K NE IIEAYRTLRDRGPYPADQV++ +G FAF+++D
Sbjct: 257 GHIDNVANLKQQYGLNKTANEVTIIIEAYRTLRDRGPYPADQVVRDFQGKFAFILFDSSS 436
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
T F AS DG + +WGI AD ++V+S++ E+V SC KS+APFP GC F + GL +F
Sbjct: 437 QTAFVASDVDGSVPFFWGIDADENLVLSDDTEIVSKSCGKSYAPFPKGCFFTTSGGLRSF 616
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQM--MPRVGSEANWAT 167
EHP ++K +PRIDS G +CG+ F VD+ ++ + MPRVGS ANW++
Sbjct: 617 EHPLHELKPVPRIDSSGEVCGSTFKVDADTKKEATGMPRVGSAANWSS 760
>TC77432 similar to GP|18252901|gb|AAL62377.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 966
Score = 190 bits (482), Expect = 3e-49
Identities = 82/167 (49%), Positives = 121/167 (72%), Gaps = 2/167 (1%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+L N++ L +QYGL+K NE + +IEAYRTLRDRGPYPA QV++ +G FAF+++D
Sbjct: 293 GHLENVANLKQQYGLNKTANEVIIVIEAYRTLRDRGPYPASQVVRDFQGKFAFILFDSGS 472
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
F ++ +DG++ +WG ADG++V+S+ ++V SC KS+APFP GC F + GL +F
Sbjct: 473 KNAFISADADGNVPFFWGTDADGNLVLSDETDIVTKSCGKSYAPFPKGCFFTTSGGLSSF 652
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQM--MPRVGSEANWA 166
EHP ++K +PR+DS G +CGA F VD++++ + MPRVGS ANW+
Sbjct: 653 EHPLNELKPVPRVDSSGQVCGATFKVDAEAKKESTGMPRVGSAANWS 793
>BQ151778 weakly similar to GP|643469|gb|AA unknown {Lycopersicon
esculentum}, partial (29%)
Length = 491
Score = 187 bits (474), Expect = 2e-48
Identities = 88/88 (100%), Positives = 88/88 (100%)
Frame = +2
Query: 83 DGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCG 142
DGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCG
Sbjct: 32 DGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCG 211
Query: 143 ANFNVDSQSRNQMMPRVGSEANWATWGS 170
ANFNVDSQSRNQMMPRVGSEANWATWGS
Sbjct: 212 ANFNVDSQSRNQMMPRVGSEANWATWGS 295
>TC85670 homologue to GP|2970051|dbj|BAA25187.1 ARG10 {Vigna radiata},
complete
Length = 1144
Score = 141 bits (356), Expect = 1e-34
Identities = 69/148 (46%), Positives = 96/148 (64%), Gaps = 1/148 (0%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+L NL L +QYGLSK NE + +IEAY+ LRDR PYPA+ V+ L GSFAF+++D
Sbjct: 315 GSLDNLGSLRQQYGLSKSANEVVLMIEAYKALRDRAPYPANHVVGHLSGSFAFIVFDKST 494
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI ADG V +++ +L+K +C KS A FP GC + + GL
Sbjct: 495 STLFVASDQSGKVPLYWGITADGYVAFADDADLLKGACGKSLASFPQGCFYSTAVGGLRC 674
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVD 148
+E+P K+ A+P + E + GA F V+
Sbjct: 675 YENPKNKITAVPANEEE--IWGATFKVE 752
>TC86172 similar to GP|2970051|dbj|BAA25187.1 ARG10 {Vigna radiata}, partial
(98%)
Length = 1015
Score = 134 bits (337), Expect = 2e-32
Identities = 65/148 (43%), Positives = 95/148 (63%), Gaps = 1/148 (0%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL +L +QYGL+K NE + +IEAY+ LRDR PYP + V+ L G+FAF+++D
Sbjct: 317 GALDNLGRLRQQYGLAKSANEVVLVIEAYKALRDRAPYPPNHVVGHLSGTFAFILFDKST 496
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + L+WGI ADG +++ EL+K++C KS A FP GC + + GL+
Sbjct: 497 STLFVASDQFGKVPLFWGITADGYAAFADDAELLKSACGKSLASFPQGCFYSTAVGGLMC 676
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVD 148
+E+P K+ A+P +E GA F V+
Sbjct: 677 YENPKNKITAVPA--NEEDFWGATFKVE 754
>TC85188 similar to GP|643469|gb|AAA61967.1|| unknown {Lycopersicon
esculentum}, partial (13%)
Length = 351
Score = 93.2 bits (230), Expect = 5e-20
Identities = 42/44 (95%), Positives = 44/44 (99%)
Frame = +2
Query: 127 KMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS 170
++KAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS
Sbjct: 29 EVKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS 160
>TC85205 similar to GP|4559292|gb|AAD22970.1| trehalase 1 GMTRE1 {Glycine
max}, partial (34%)
Length = 814
Score = 55.1 bits (131), Expect = 1e-08
Identities = 25/25 (100%), Positives = 25/25 (100%)
Frame = +1
Query: 15 GLSKGGNEAMFIIEAYRTLRDRGPY 39
GLSKGGNEAMFIIEAYRTLRDRGPY
Sbjct: 1 GLSKGGNEAMFIIEAYRTLRDRGPY 75
>TC85437 homologue to PIR|S11443|AJPMN2 asparagine synthase
(glutamine-hydrolyzing) (EC 6.3.5.4) 2 - garden pea,
complete
Length = 2248
Score = 48.1 bits (113), Expect = 2e-06
Identities = 38/118 (32%), Positives = 55/118 (46%), Gaps = 2/118 (1%)
Frame = +3
Query: 2 GNLHNLSQLNKQYG--LSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N L KQ + G++ I Y Y D V K L+G F+FV+ D
Sbjct: 402 GEIYNHEALRKQLSNHTFRTGSDCDVIAHLYEE------YGEDFVDK-LDGIFSFVLLDT 560
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHG 117
+D + A + G LY G DGSV IS ++ + C + F FP G L+ S+ G
Sbjct: 561 RDNSYIVARDAIGVTSLYIGWGLDGSVWISSEMKALNDDC-EHFECFPPGHLYSSKDG 731
>BQ156832 homologue to PIR|F86214|F862 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 662
Score = 45.8 bits (107), Expect = 8e-06
Identities = 20/20 (100%), Positives = 20/20 (100%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGN 21
GNLHNLSQLNKQYGLSKGGN
Sbjct: 602 GNLHNLSQLNKQYGLSKGGN 661
>BQ157268 homologue to PIR|F86214|F862 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 656
Score = 43.1 bits (100), Expect = 5e-05
Identities = 19/19 (100%), Positives = 19/19 (100%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKG 19
MGNLHNLSQLNKQYGLSKG
Sbjct: 598 MGNLHNLSQLNKQYGLSKG 654
>TC84317 similar to GP|3859536|gb|AAC72837.1| asparagine synthetase
{Arabidopsis thaliana}, partial (44%)
Length = 802
Score = 43.1 bits (100), Expect = 5e-05
Identities = 23/70 (32%), Positives = 38/70 (53%)
Frame = +3
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + AA + G LY G DGS+ + ++ + C + F FP
Sbjct: 381 LDGMFSFVLLDTRDKSFIAARDAIGITPLYLGWGHDGSIWFASEMKALIDDC-EQFISFP 557
Query: 108 TGCLFHSEHG 117
G ++ S+ G
Sbjct: 558 PGHIYSSKQG 587
>TC85436 GP|2522320|gb|AAB81011.1| asparagine synthetase {Medicago sativa},
complete
Length = 2147
Score = 43.1 bits (100), Expect = 5e-05
Identities = 25/68 (36%), Positives = 37/68 (53%)
Frame = +1
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + A + G LY G DGSV I+ L+ + C + F FP
Sbjct: 442 LDGIFSFVLLDTRDNSFIVARDAIGVTSLYIGWGLDGSVWIASELKGLNDEC-EHFEVFP 618
Query: 108 TGCLFHSE 115
G L+ S+
Sbjct: 619 PGHLYSSK 642
>BM813085 weakly similar to GP|15795110|dbj gene_id:MJK13.11~unknown protein
{Arabidopsis thaliana}, partial (8%)
Length = 774
Score = 33.9 bits (76), Expect = 0.033
Identities = 17/37 (45%), Positives = 22/37 (58%)
Frame = +2
Query: 80 IAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEH 116
I DGSVVIS++L +++ CAK FP G S H
Sbjct: 206 ITGDGSVVISDDLNVIQEGCAK----FPAGTNIFSHH 304
>TC83544 weakly similar to GP|15795110|dbj|BAB02374.
gene_id:MJK13.11~unknown protein {Arabidopsis thaliana},
partial (8%)
Length = 687
Score = 30.0 bits (66), Expect = 0.47
Identities = 18/52 (34%), Positives = 27/52 (51%), Gaps = 7/52 (13%)
Frame = +3
Query: 80 IAADGSVVISENLELVKASCAKSFAPFPTGCLFH-------SEHGLLNFEHP 124
I DGSVVIS++L +++ CAK FP + + S +G +N P
Sbjct: 282 ITGDGSVVISDDLNVIQEGCAK----FPAATMVNNVRFTLCSNYGYMNSWFP 425
>AW586033 similar to GP|14030723|gb| AT4g27450/F27G19_50 {Arabidopsis
thaliana}, partial (7%)
Length = 450
Score = 29.6 bits (65), Expect = 0.61
Identities = 12/22 (54%), Positives = 17/22 (76%)
Frame = +2
Query: 80 IAADGSVVISENLELVKASCAK 101
I DGSVVIS++L +++ CAK
Sbjct: 230 ITGDGSVVISDDLNVIQEGCAK 295
>TC88944 type IIB calcium ATPase [Medicago truncatula]
Length = 1883
Score = 27.3 bits (59), Expect = 3.0
Identities = 21/74 (28%), Positives = 35/74 (46%)
Frame = +3
Query: 22 EAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIA 81
E + II + + P ++K L +F V+ DGT A + + IGL GIA
Sbjct: 915 ELLDIIPKIQVMARSSPMDKHTLVKHLRTTFEEVVAVTGDGTNDAPALHEADIGLAMGIA 1094
Query: 82 ADGSVVISENLELV 95
G+ V E+ +++
Sbjct: 1095--GTEVAKESADVI 1130
>TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.300 -
Arabidopsis thaliana, partial (74%)
Length = 1630
Score = 26.6 bits (57), Expect = 5.2
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +2
Query: 61 DGTVFAASGSDGHI 74
DG+ FAA GSDGH+
Sbjct: 551 DGSKFAAGGSDGHL 592
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,985,383
Number of Sequences: 36976
Number of extensions: 58900
Number of successful extensions: 269
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 267
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 267
length of query: 170
length of database: 9,014,727
effective HSP length: 89
effective length of query: 81
effective length of database: 5,723,863
effective search space: 463632903
effective search space used: 463632903
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC129091.3


