

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
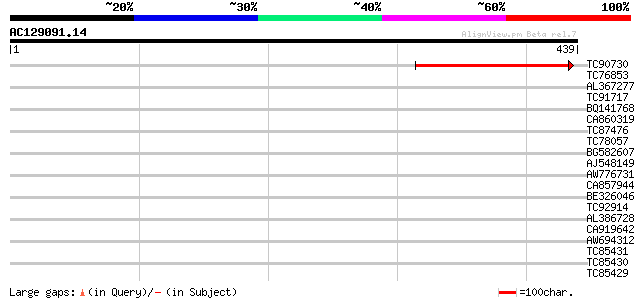
Query= AC129091.14 + phase: 0
(439 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90730 similar to PIR|T01247|T01247 hypothetical protein At2g38... 157 6e-39
TC76853 similar to PIR|T09581|T09581 probable high mobility grou... 34 0.13
AL367277 homologue to PIR|S14974|S149 extensin class I (clone uG... 33 0.22
TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein N... 32 0.63
BQ141768 weakly similar to GP|11036868|gb| PxORF73 peptide {Plut... 32 0.63
CA860319 weakly similar to PIR|T38663|T386 probable translation ... 31 0.83
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 31 0.83
TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice... 31 0.83
BG582607 31 1.1
AJ548149 similar to GP|14583094|gb chromomethylase CMT2 {Arabido... 30 1.4
AW776731 GP|19683016|g hypothetical protein {Dictyostelium disco... 30 1.4
CA857944 30 1.8
BE326046 similar to PIR|A84810|A848 probable guanylate binding p... 30 1.8
TC92914 similar to PIR|F85023|F85023 probable potassium channel ... 30 1.8
AL386728 30 1.8
CA919642 similar to PIR|E85433|E854 SCARECROW-like protein [impo... 30 2.4
AW694312 similar to PIR|A84810|A84 probable guanylate binding pr... 30 2.4
TC85431 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transf... 30 2.4
TC85430 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transf... 30 2.4
TC85429 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transf... 30 2.4
>TC90730 similar to PIR|T01247|T01247 hypothetical protein At2g38150
[imported] - Arabidopsis thaliana, partial (8%)
Length = 630
Score = 157 bits (398), Expect = 6e-39
Identities = 67/122 (54%), Positives = 89/122 (72%)
Frame = +3
Query: 315 FDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNFSVVPPSAFYPVDW 374
FDK HPL+L+FI EFALTFDGNKWGHNGPYL+SRVV R+ R+G+NF+++PP AFYPV W
Sbjct: 3 FDKNHPLVLRFINEFALTFDGNKWGHNGPYLVSRVVERLKKRQGFNFTILPPMAFYPVSW 182
Query: 375 RGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISSCCI 434
I FR P KW+ K+ Q+ E++ VHLWN+QS L + +GS++ ++S+ CI
Sbjct: 183 TKIGGFFRKPKTRSEEKWVEAKLKQLSGETFGVHLWNKQSSGLVIEEGSVMARLVSNHCI 362
Query: 435 FC 436
C
Sbjct: 363 IC 368
>TC76853 similar to PIR|T09581|T09581 probable high mobility group protein
HMG1 - sword bean, partial (97%)
Length = 907
Score = 33.9 bits (76), Expect = 0.13
Identities = 31/92 (33%), Positives = 39/92 (41%), Gaps = 19/92 (20%)
Frame = -2
Query: 51 LFLLAYNAFTVFCIHIPH--HYSPHK-----------PLLLPPPS------HTKTKLSSS 91
L L AYN FT H PH H PH P L PP S T LSSS
Sbjct: 579 LVLQAYNNFTHHLHHHPHPPHLHPHH*LQILTCQNHLPRLFPPVSCCRLV*QPHTLLSSS 400
Query: 92 VFFAHKEDNSPIINKTHFPLLHKISIHKFKRV 123
+ HKE P + +T +LH+ + +R+
Sbjct: 399 MP*QHKE---PCLLQTSISMLHRKLFRQRQRI 313
>AL367277 homologue to PIR|S14974|S149 extensin class I (clone uG-18) -
tomato (fragment), partial (46%)
Length = 254
Score = 33.1 bits (74), Expect = 0.22
Identities = 29/90 (32%), Positives = 37/90 (40%), Gaps = 7/90 (7%)
Frame = +2
Query: 4 HTKHHHNPFLYFPLLLLHKLHHFLHNFKKKI-------LSILFKLPTSFFALLVLFLLAY 56
H HHH+ L+ ++ LH HH L + I +L TS L L L Y
Sbjct: 14 HHPHHHHHLLHHTIISLHLHHHHLPHHLIIISHHPHHPRPLLHHTTTS----LHLHLKKY 181
Query: 57 NAFTVFCIHIPHHYSPHKPLLLPPPSHTKT 86
+ + H PHH H PL P HT T
Sbjct: 182HILHITISHHPHH---HHPL----PLHTIT 250
>TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein NFH2
{Nicotiana tabacum}, partial (6%)
Length = 477
Score = 31.6 bits (70), Expect = 0.63
Identities = 27/94 (28%), Positives = 37/94 (38%)
Frame = +3
Query: 19 LLHKLHHFLHNFKKKILSILFKLPTSFFALLVLFLLAYNAFTVFCIHIPHHYSPHKPLLL 78
L H LHH LH+ ++ + F L L L F +H P PH L L
Sbjct: 192 LHHLLHHHLHH----------QIQNTHFLLHPLTLHLLLPHPPFSLHTPQ---PHHHLHL 332
Query: 79 PPPSHTKTKLSSSVFFAHKEDNSPIINKTHFPLL 112
P H++ S+ K N + N + PLL
Sbjct: 333 LPSLHSQLTFHLSLSHKPKNPNRHLPNSSPSPLL 434
>BQ141768 weakly similar to GP|11036868|gb| PxORF73 peptide {Plutella
xylostella granulovirus}, partial (29%)
Length = 958
Score = 31.6 bits (70), Expect = 0.63
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +1
Query: 65 HIPHHYSPHKPLLLPPPSHTK 85
+ P HY PH ++ PPPSH K
Sbjct: 274 YTPPHYIPHSTIIHPPPSHFK 336
>CA860319 weakly similar to PIR|T38663|T386 probable translation initiation
factor eif-2b beta subunit - fission yeast, partial
(22%)
Length = 767
Score = 31.2 bits (69), Expect = 0.83
Identities = 17/47 (36%), Positives = 28/47 (59%)
Frame = -1
Query: 7 HHHNPFLYFPLLLLHKLHHFLHNFKKKILSILFKLPTSFFALLVLFL 53
HHH+P L+ PLL+ ++ LH L ++ ++PTSF L ++ L
Sbjct: 377 HHHHPLLHLPLLVKMIAYYQLHQLLHL*LWVV-QVPTSFQHL*LILL 240
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 31.2 bits (69), Expect = 0.83
Identities = 26/76 (34%), Positives = 32/76 (41%), Gaps = 4/76 (5%)
Frame = +1
Query: 2 EDHTKHHHNPFLYFPLLLLHKLHHFLHNFKK----KILSILFKLPTSFFALLVLFLLAYN 57
+ H HHH PLLLL LH LH F +L L + S LL L L +
Sbjct: 367 DHHPLHHHLHMSISPLLLL-LLHPLLHMFTNLLPHHLLHPLHPMSISHLHLLQLHHLLHT 543
Query: 58 AFTVFCIHIPHHYSPH 73
T +H+ HH H
Sbjct: 544 T-TNLHLHLLHHLLHH 588
Score = 27.7 bits (60), Expect = 9.1
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = +3
Query: 4 HTKHHHNPFLYFPLLLLHKLHHFLH 28
H H H+ + LLL H LHH LH
Sbjct: 273 HRLHPHHLHTFINLLLHHLLHHRLH 347
>TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice, partial
(50%)
Length = 878
Score = 31.2 bits (69), Expect = 0.83
Identities = 29/101 (28%), Positives = 42/101 (40%), Gaps = 16/101 (15%)
Frame = +3
Query: 4 HTKHHHN-----PFLYF------PLLLLHKLHHFLHNFKKKI-LSILFKLPTSFFALLV- 50
HT HHHN P L P++ + + HN KK+ S L P + +
Sbjct: 165 HTHHHHNNNHINPKLKSSTPGTCPVMTISTITLNNHNLTKKLKCSTLDTCPMKTSTVTLN 344
Query: 51 ---LFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKL 88
L L+ N T H+ HH P + PP H++T+L
Sbjct: 345 HHHLNLIINNPATALRTHLHHH-----PYRITPPPHSQTQL 452
>BG582607
Length = 754
Score = 30.8 bits (68), Expect = 1.1
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 8/75 (10%)
Frame = -2
Query: 7 HHHNPFLYFPLLLLHKLHHFLHNFKKKILSILFKLPTSF--------FALLVLFLLAYNA 58
HHH+ L F LLH LH ++ + +LF++ F + L L L+ N
Sbjct: 726 HHHHW*LQFLFRLLHVLH---SEYQFPLHLVLFRMNNIFLQQYLKPHYLQLFLLLMMPNI 556
Query: 59 FTVFCIHIPHHYSPH 73
FC H+ + P+
Sbjct: 555 LHQFCYHVMTQHFPN 511
>AJ548149 similar to GP|14583094|gb chromomethylase CMT2 {Arabidopsis
thaliana}, partial (6%)
Length = 474
Score = 30.4 bits (67), Expect = 1.4
Identities = 27/88 (30%), Positives = 40/88 (44%), Gaps = 5/88 (5%)
Frame = +2
Query: 10 NPFLYFPLLLLHKLHHFLHNF----KKKILSILFKLPTSF-FALLVLFLLAYNAFTVFCI 64
N L+F L++LH F H+F K S L + F F LL+ +LL A V +
Sbjct: 143 NGVLFFVSPTLYRLHSFSHSFNSCRKSSADSFLTCVSGWFSFKLLLAYLLLSTAHLVTRL 322
Query: 65 HIPHHYSPHKPLLLPPPSHTKTKLSSSV 92
H P+ +PP ++ +SSV
Sbjct: 323 TDDILAPKHSPVDMPPQXE*RSSNASSV 406
>AW776731 GP|19683016|g hypothetical protein {Dictyostelium discoideum},
partial (1%)
Length = 413
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/50 (26%), Positives = 25/50 (50%)
Frame = +1
Query: 64 IHIPHHYSPHKPLLLPPPSHTKTKLSSSVFFAHKEDNSPIINKTHFPLLH 113
+++ H+ H P PP H +++++ F + + SP +K PL H
Sbjct: 67 LYLITHFLLHPPYYYPPSKHHPLQITTTTLFKNPKTLSPKFSKPLTPLPH 216
>CA857944
Length = 796
Score = 30.0 bits (66), Expect = 1.8
Identities = 26/95 (27%), Positives = 43/95 (44%), Gaps = 5/95 (5%)
Frame = +1
Query: 31 KKKILSILFKLPTSFFALLVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSS 90
K I+S+ PT +A+ L L Y+ F V Y PL+LP P+H + +++
Sbjct: 10 KLLIISLTVSFPTDPYAMYAL--LPYSPFKVSV------Y*TCDPLILPNPNHLRAEMNK 165
Query: 91 S-VFFAHKEDNSPIINKTHFPLLHKISI----HKF 120
S F+ + + I + F +H I H+F
Sbjct: 166 SPEFYKSYQT*AEIYKSSAFMKIHNPQIIAGKHRF 270
>BE326046 similar to PIR|A84810|A848 probable guanylate binding protein
[imported] - Arabidopsis thaliana, partial (31%)
Length = 527
Score = 30.0 bits (66), Expect = 1.8
Identities = 20/56 (35%), Positives = 30/56 (52%)
Frame = +2
Query: 48 LLVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSSSVFFAHKEDNSPI 103
L +LFLL + + TV I + + P P++ P P HTK +LS A + +PI
Sbjct: 203 LSLLFLLFFLSPTVSSETIINFHQPF-PIVEPDPGHTKLRLSQDGLEAIERITNPI 367
>TC92914 similar to PIR|F85023|F85023 probable potassium channel [imported]
- Arabidopsis thaliana, partial (24%)
Length = 672
Score = 30.0 bits (66), Expect = 1.8
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +2
Query: 3 DHTKHHHNPFLYFPLLLLHKLHHFLHNFKK 32
++ HHHN Y P LH +HFL KK
Sbjct: 110 NNNNHHHNHLRYNPSTSLHNNNHFLITNKK 199
>AL386728
Length = 452
Score = 30.0 bits (66), Expect = 1.8
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 1/43 (2%)
Frame = -1
Query: 37 ILFKLPTSFFALLVLFLLAYNAFTV-FCIHIPHHYSPHKPLLL 78
I ++PT +F + L L Y FTV F I P+ SP+ +L
Sbjct: 320 IRIRIPTLYFIIFTLIFLIYIIFTVIFIIQPPNEVSPNSDRVL 192
>CA919642 similar to PIR|E85433|E854 SCARECROW-like protein [imported] -
Arabidopsis thaliana, partial (12%)
Length = 661
Score = 29.6 bits (65), Expect = 2.4
Identities = 14/36 (38%), Positives = 21/36 (57%), Gaps = 3/36 (8%)
Frame = +3
Query: 72 PHKPLLLPPPSHTKTKLSSSV---FFAHKEDNSPII 104
P PLL PPP+H T SSS + H ++ +P++
Sbjct: 408 PTLPLLPPPPTHPMTPASSSKTPNYSQHHDEKTPLL 515
>AW694312 similar to PIR|A84810|A84 probable guanylate binding protein
[imported] - Arabidopsis thaliana, partial (59%)
Length = 570
Score = 29.6 bits (65), Expect = 2.4
Identities = 17/56 (30%), Positives = 29/56 (51%)
Frame = +3
Query: 48 LLVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSSSVFFAHKEDNSPI 103
LL L +++ ++ F IH ++ P++ P P HTK +LS A + +PI
Sbjct: 9 LLFLLIISLSSIPSFSIH---NFHQPFPIVEPDPGHTKLRLSREGLEAIERITNPI 167
>TC85431 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transfer protein
precursor (LTP). [Chickpea Garbanzo] {Cicer arietinum},
complete
Length = 616
Score = 29.6 bits (65), Expect = 2.4
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +1
Query: 49 LVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSSS 91
LVL + A + V +H+ H + L +PPP H +L ++
Sbjct: 190 LVLLVHALDILKVVLVHLQHAVVESRDLTVPPPPHLTVRLHAT 318
>TC85430 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transfer protein
precursor (LTP). [Chickpea Garbanzo] {Cicer arietinum},
complete
Length = 646
Score = 29.6 bits (65), Expect = 2.4
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +3
Query: 49 LVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSSS 91
LVL + A + V +H+ H + L +PPP H +L ++
Sbjct: 141 LVLLVHALDILKVVLVHLQHAVVESRDLTVPPPPHLTVRLHAT 269
>TC85429 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transfer protein
precursor (LTP). [Chickpea Garbanzo] {Cicer arietinum},
complete
Length = 698
Score = 29.6 bits (65), Expect = 2.4
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +1
Query: 49 LVLFLLAYNAFTVFCIHIPHHYSPHKPLLLPPPSHTKTKLSSS 91
LVL + A + V +H+ H + L +PPP H +L ++
Sbjct: 163 LVLLVHALDILKVVLVHLQHAVVESRDLTVPPPPHLTVRLHAT 291
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,576,700
Number of Sequences: 36976
Number of extensions: 347379
Number of successful extensions: 3977
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 3809
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3954
length of query: 439
length of database: 9,014,727
effective HSP length: 99
effective length of query: 340
effective length of database: 5,354,103
effective search space: 1820395020
effective search space used: 1820395020
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC129091.14


