

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127429.3 - phase: 0 /pseudo
(423 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
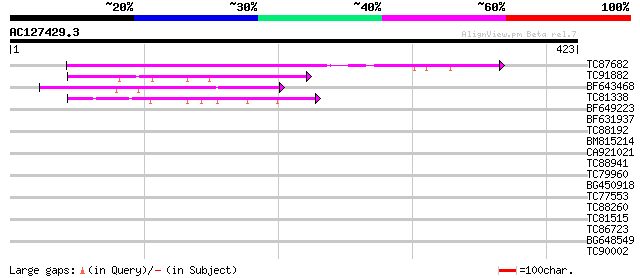
TC87682 weakly similar to PIR|T04527|T04527 hypothetical protein... 210 9e-55
TC91882 similar to GP|10176714|dbj|BAB09944. dimethylaniline mon... 69 3e-12
BF643468 weakly similar to PIR|F96682|F96 hypothetical protein F... 61 9e-10
TC81338 weakly similar to PIR|B86326|B86326 hypothetical protein... 53 2e-07
BF649223 weakly similar to GP|21740857|emb OSJNBb0072N21.6 {Oryz... 37 0.019
BF631937 similar to GP|10177621|db phytoene dehydrogenase-like {... 32 0.35
TC88192 weakly similar to PIR|T05682|T05682 monooxygenase (EC 1.... 32 0.35
BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis ... 32 0.46
CA921021 32 0.60
TC88941 similar to PIR|T52419|T52419 CTF2A protein [imported] - ... 32 0.60
TC79960 similar to PIR|T52419|T52419 CTF2A protein [imported] - ... 32 0.60
BG450918 similar to PIR|T00665|T006 hypothetical protein F3I6.28... 31 1.0
TC77553 homologue to GP|1066499|gb|AAB41904.1| NADH-dependent gl... 31 1.0
TC88260 homologue to GP|15215696|gb|AAK91394.1 AT3g16950/K14A17_... 30 2.3
TC81515 TRE1 protein 29 3.0
TC86723 similar to PIR|B96562|B96562 unknown protein [imported] ... 28 5.1
BG648549 similar to PIR|E85359|E853 hypothetical protein AT4g307... 28 5.1
TC90002 weakly similar to GP|9757850|dbj|BAB08484.1 dimethylanil... 28 6.7
>TC87682 weakly similar to PIR|T04527|T04527 hypothetical protein F16A16.170
- Arabidopsis thaliana, partial (38%)
Length = 1040
Score = 210 bits (534), Expect = 9e-55
Identities = 135/349 (38%), Positives = 191/349 (54%), Gaps = 22/349 (6%)
Frame = +1
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEF 102
+I+GAG SGLA AA L ++ +P +ILER NC ASLW+ TYDR+ LHL KQ+CELP F
Sbjct: 46 IIIGAGTSGLATAACLTKQSIPFIILERENCFASLWQNYTYDRVHLHLRKQLCELPHFPF 225
Query: 103 PSGFPTYPTKQQFIEYLESYSKNFDIRPWFNETVMHAEF-DATLGFWRVRSEGK-AGMVT 160
P +P Y K+QFIEYL +Y NF+I P +N V AE+ D WRV++E K +G V
Sbjct: 226 PPSYPHYVPKKQFIEYLGNYVNNFNINPIYNRAVELAEYVDDDEKKWRVKAENKSSGEVE 405
Query: 161 EFVCRWLIVATGENAEAVVPEIEGVDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGME 220
E+ R+L+VA+GE AE VP +EG++ F G + H++ YK+G+EF+ + VLVVG GNSGME
Sbjct: 406 EYSARFLVVASGETAEPRVPVVEGLENFKGKVIHSTRYKNGKEFKDEHVLVVGSGNSGME 585
Query: 221 VCLDLCNHDAAPSIVVRDSPTKRYAGKINFWVVHVVAQVATRATGGSHPSHCVMVNARQH 280
+ LDL N A PSI+VR SP VH++++ GG V ++ +H
Sbjct: 586 IALDLANFGAKPSIIVR-SP------------VHILSRDMMYYGGG-----FVELSVTKH 711
Query: 281 RTVRFGSASVGSP*TQKTVW-----------KDTGPRCGC----PCQD*K-RRHQGIKRL 324
G + + +W +D + C CQ+ + R G+
Sbjct: 712 SGETSGDSKQDCVWRSEQIWYTFS**RPIHHEDEVWQVSCN*RWNCQENQIRGDSGVASR 891
Query: 325 KRY----TVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYP 369
R + FD+II TG++ + WL +F EDG+P
Sbjct: 892 NREHKW*PSVVP*WKSYPFDSIIFCTGFQRSTQKWL*GWVIF*NEDGFP 1038
>TC91882 similar to GP|10176714|dbj|BAB09944. dimethylaniline
monooxygenase-like protein {Arabidopsis thaliana},
partial (47%)
Length = 800
Score = 69.3 bits (168), Expect = 3e-12
Identities = 55/218 (25%), Positives = 99/218 (45%), Gaps = 36/218 (16%)
Frame = +2
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKL---------------------KT 82
++GAGP+GL A ++++G ++LE+++ I W
Sbjct: 68 VIGAGPAGLVAAREIRKEGHKVIVLEQNHDIGGQWLYDDKNIEGEDPLGRNPFLKVHSSI 247
Query: 83 YDRLRLHLPKQVCELPLMEFPSG------FPTYPTKQQFIEYLESYSKNFDIRPW--FNE 134
Y+ LR P++V + +FP +P + + YL+ + + F +R FN
Sbjct: 248 YNSLRTQSPREV--MGFTDFPFSTKKGRDMRRFPGHVEILMYLKDFCEWFGLREMIRFNT 421
Query: 135 TVMHAEFDATLGF------WRVRSEGK-AGMVTEFVCRWLIVATGENAEAVVPEIEGVDE 187
V + G W VRS+ K + V E V ++VATG + +P I+G+D
Sbjct: 422 RVNYVGMLDDYGVCGNNLKWIVRSKEKNSDKVVEEVFDAVVVATGHYCQPKLPSIKGMDT 601
Query: 188 FVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDL 225
+ H+ +Y++ E F + V+VVG SG ++ ++L
Sbjct: 602 WKRKQMHSHIYRTPEPFHNEVVVVVGNSFSGQDIAIEL 715
>BF643468 weakly similar to PIR|F96682|F96 hypothetical protein F12P19.2
[imported] - Arabidopsis thaliana, partial (33%)
Length = 655
Score = 60.8 bits (146), Expect = 9e-10
Identities = 48/207 (23%), Positives = 90/207 (43%), Gaps = 24/207 (11%)
Frame = +2
Query: 23 INNNKNSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLW---- 78
+N ++NS + + + + ++GAG SGL A L+Q+G ++ E++N + W
Sbjct: 32 LNFDQNSQRFACQLITMVKVAVIGAGVSGLVAAKELQQEGHNVVVFEKNNRVGGTWIYTP 211
Query: 79 ----------------KLKTYDRLRLHLPKQV---CELPLMEFPSGFP-TYPTKQQFIEY 118
Y LR +LP+Q+ + PL + SG P T+P ++ + +
Sbjct: 212 KTDSDPLSMDPARETVHSSVYHSLRTNLPRQIMGFLDYPLSKRESGDPRTFPGHEEVLRF 391
Query: 119 LESYSKNFDIRPWFNETVMHAEFDATLGFWRVRSEGKAGMVTEFVCRWLIVATGENAEAV 178
LE ++ F I + + W V S G V++ V ++V +G E
Sbjct: 392 LEEFAGEFGIHELTRFETEVVKVERKENEWIVESRG-GDSVSQEVFEAVVVCSGHFVEPR 568
Query: 179 VPEIEGVDEFVGSIRHTSLYKSGEEFR 205
+ + G++ F G H+ +S F+
Sbjct: 569 LAVVPGIETFPGFQMHSHKLQSSSFFQ 649
>TC81338 weakly similar to PIR|B86326|B86326 hypothetical protein AAF82235.1
[imported] - Arabidopsis thaliana, partial (32%)
Length = 784
Score = 53.1 bits (126), Expect = 2e-07
Identities = 52/224 (23%), Positives = 95/224 (42%), Gaps = 35/224 (15%)
Frame = +2
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFP 103
I+GAG SGLAVA L ++ E ++ I +W+ +Y +L Q +FP
Sbjct: 77 IIGAGVSGLAVAKQLSHYN--PIVFEATDSIGGVWRHCSYRCTKLQ--SQTWNYEFSDFP 244
Query: 104 ---SGFPTYPTKQQFIEYLESYSKNFDIRPW--FNETVMHAEF---DATLGFWRVRSE-- 153
YP+ + +EYL Y+ FD+ + FN V+ +F F R+ +
Sbjct: 245 WPERESKDYPSHAEILEYLHLYAVYFDLFKYVKFNTKVVEIKFVRDKEGFDFGRLPGDHG 424
Query: 154 ---------------GKAGMVTEFVCRWLIVATGENAE----AVVPEIEGVDEFVGSIRH 194
++ + ++ +++V TG+ + P +G + F G + H
Sbjct: 425 NPLPERPVWELSVQTNESDAIQKYCFEFVVVCTGKYGDIPLMPTFPCNKGPEVFKGKVLH 604
Query: 195 TSLY------KSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAP 232
T Y + + +GKKV+VVG S +++ ++ + P
Sbjct: 605 TIDYCKLDKEATSDLVKGKKVVVVGYKKSAIDLTMECVQANQRP 736
>BF649223 weakly similar to GP|21740857|emb OSJNBb0072N21.6 {Oryza sativa},
partial (11%)
Length = 278
Score = 36.6 bits (83), Expect = 0.019
Identities = 20/61 (32%), Positives = 30/61 (48%)
Frame = +2
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFP 103
IVGAG SGL Y+ Q G ++ E + + LW+ T + +L PKQ + +
Sbjct: 83 IVGAGISGLLACKYVTQIGFNPIVFEADDGVGGLWR-HTIESTKLQNPKQAYQFSXFPWD 259
Query: 104 S 104
S
Sbjct: 260 S 262
>BF631937 similar to GP|10177621|db phytoene dehydrogenase-like {Arabidopsis
thaliana}, partial (26%)
Length = 688
Score = 32.3 bits (72), Expect = 0.35
Identities = 15/32 (46%), Positives = 21/32 (64%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
LI+G G +GL AAYL + G+ +LER + I
Sbjct: 120 LIIGGGHNGLTAAAYLARGGLSVAVLERRHII 215
>TC88192 weakly similar to PIR|T05682|T05682 monooxygenase (EC 1.-.-.-) 2 -
Arabidopsis thaliana, partial (61%)
Length = 1557
Score = 32.3 bits (72), Expect = 0.35
Identities = 17/32 (53%), Positives = 23/32 (71%)
Frame = +2
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
+IVGAG +GLA A L + GV SL+LE S+ +
Sbjct: 128 VIVGAGIAGLATALGLHRLGVRSLVLESSDSL 223
>BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 596
Score = 32.0 bits (71), Expect = 0.46
Identities = 17/38 (44%), Positives = 24/38 (62%)
Frame = +1
Query: 329 VEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKED 366
V F DG + DAII TGYK ++P+ L+ G+ + ED
Sbjct: 238 VAFEDGFSTYADAIIHCTGYKYHIPF-LETNGIVTIED 348
>CA921021
Length = 261
Score = 31.6 bits (70), Expect = 0.60
Identities = 13/21 (61%), Positives = 16/21 (75%)
Frame = -1
Query: 387 GFTKRGLLGASMDAKNIAEDI 407
G T+RG GA MDA+N+A DI
Sbjct: 261 GSTRRGFYGAKMDAQNVANDI 199
>TC88941 similar to PIR|T52419|T52419 CTF2A protein [imported] - Arabidopsis
thaliana, partial (39%)
Length = 659
Score = 31.6 bits (70), Expect = 0.60
Identities = 16/32 (50%), Positives = 22/32 (68%)
Frame = +1
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
+IVG G +GLA A L + GV SL+LE+S +
Sbjct: 154 VIVGGGIAGLATALSLHRLGVRSLVLEQSESL 249
Score = 28.1 bits (61), Expect = 6.7
Identities = 17/58 (29%), Positives = 30/58 (51%)
Frame = +1
Query: 189 VGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEVCLDLCNHDAAPSIVVRDSPTKRYAG 246
+GS+R + L K + + V++VG G +G+ L L + S+V+ S + R G
Sbjct: 91 IGSLRSSKLIKVQSSVQKEHVVIVGGGIAGLATALSL-HRLGVRSLVLEQSESLRTGG 261
>TC79960 similar to PIR|T52419|T52419 CTF2A protein [imported] - Arabidopsis
thaliana, partial (53%)
Length = 907
Score = 31.6 bits (70), Expect = 0.60
Identities = 16/32 (50%), Positives = 22/32 (68%)
Frame = +1
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCI 74
+IVG G +GLA A L + GV SL+LE+S +
Sbjct: 202 VIVGGGIAGLATALSLHRLGVRSLVLEQSESL 297
>BG450918 similar to PIR|T00665|T006 hypothetical protein F3I6.28 -
Arabidopsis thaliana, partial (6%)
Length = 646
Score = 30.8 bits (68), Expect = 1.0
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCIA 75
LI+GAGP GL ++ L + G+ +LER+ +
Sbjct: 366 LIIGAGPVGLVLSILLTKLGINCTVLERNKAFS 464
>TC77553 homologue to GP|1066499|gb|AAB41904.1| NADH-dependent glutamate
synthase {Medicago sativa}, partial (37%)
Length = 3067
Score = 30.8 bits (68), Expect = 1.0
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = +1
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASL 77
IVG+GPSGLA A L + G + ER++ I L
Sbjct: 1387 IVGSGPSGLAAADQLNKMGHIVTVFERADRIGGL 1488
>TC88260 homologue to GP|15215696|gb|AAK91394.1 AT3g16950/K14A17_7
{Arabidopsis thaliana}, partial (80%)
Length = 2274
Score = 29.6 bits (65), Expect = 2.3
Identities = 52/216 (24%), Positives = 82/216 (37%), Gaps = 14/216 (6%)
Frame = +3
Query: 19 IEKMINNNKNSTSSSDRCLWIPGPLIVGAGPSGLAVAAYLKQKGVPSLILE--------- 69
I I+NN S D L LI+GAG G A + +KG+ + I+E
Sbjct: 255 ISAAISNNGTPPKSFDYDL-----LIIGAGVGGHGAALHAVEKGLKTAIVEGDVVGGTCV 419
Query: 70 RSNCIASLWKLKTYDRLR-LHLPKQVCELPLMEFPSGFPTYPTKQQFIEYLESYSKNFDI 128
C+ S L R+R L + L + +G+ +Q ++ + + I
Sbjct: 420 NRGCVPSKALLAVSGRMRDLRNDHHLKSLGIQVSNAGY----DRQAVADHANNLASK--I 581
Query: 129 RPWFNETVMHAEFDATLGFWRV---RSEGKAGMVTEFV-CRWLIVATGENAEAVVPEIEG 184
R ++ D GFW + + K G V + +I+ATG VP +
Sbjct: 582 RGNLTNSMKALGVDILTGFWELFWGPQKVKIGSSNNVVTAKDIIIATGS-----VPFVPK 746
Query: 185 VDEFVGSIRHTSLYKSGEEFRGKKVLVVGCGNSGME 220
E G TS + E + +VG G G+E
Sbjct: 747 GIEVDGKTVITSDHALKLETVPDWIAIVGSGYIGLE 854
>TC81515 TRE1 protein
Length = 1516
Score = 29.3 bits (64), Expect = 3.0
Identities = 22/69 (31%), Positives = 30/69 (42%), Gaps = 7/69 (10%)
Frame = +2
Query: 350 SNVPYWLKDKGMF-------SKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKN 402
SNV LK G+ S D + FPNGW + G K GL +A++
Sbjct: 980 SNVLKSLKTSGLLRAAGVATSLSDSGQQWDFPNGWAPLQHMLVEGLIKSGL----EEARS 1147
Query: 403 IAEDIERCW 411
+AE+I W
Sbjct: 1148 LAEEIAIRW 1174
>TC86723 similar to PIR|B96562|B96562 unknown protein [imported] -
Arabidopsis thaliana, partial (66%)
Length = 745
Score = 28.5 bits (62), Expect = 5.1
Identities = 24/69 (34%), Positives = 33/69 (47%), Gaps = 5/69 (7%)
Frame = +1
Query: 334 GSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPR--KPFPNGW-KGEN--GLYAVGF 388
G+ +NF L G +SNVP+ +K G S+ K N W K E+ GL+ +GF
Sbjct: 148 GNFQNFSVSSLPPGRRSNVPFVVKASGESSESSTSLTVFKSVQNVWDKPEDRLGLFGLGF 327
Query: 389 TKRGLLGAS 397
L AS
Sbjct: 328 AAVVALWAS 354
>BG648549 similar to PIR|E85359|E853 hypothetical protein AT4g30720
[imported] - Arabidopsis thaliana, partial (7%)
Length = 736
Score = 28.5 bits (62), Expect = 5.1
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = +2
Query: 40 PGPLIVGAGPSGLAVAAYLKQKGVPSLILER 70
P IVG+GPSGL A L + G ++ER
Sbjct: 632 PKIAIVGSGPSGLFAALVLAELGADVTLIER 724
>TC90002 weakly similar to GP|9757850|dbj|BAB08484.1 dimethylaniline
monooxygenase (N-oxide-forming)-like protein
{Arabidopsis thaliana}, partial (26%)
Length = 778
Score = 28.1 bits (61), Expect = 6.7
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = +3
Query: 329 VEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKED 366
V DG + DAII TGYK ++P+ L+ G + ED
Sbjct: 87 VPLEDGFSIYADAIIHCTGYKYHIPF-LETNGTVTIED 197
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,859,960
Number of Sequences: 36976
Number of extensions: 214669
Number of successful extensions: 916
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 902
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 912
length of query: 423
length of database: 9,014,727
effective HSP length: 99
effective length of query: 324
effective length of database: 5,354,103
effective search space: 1734729372
effective search space used: 1734729372
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC127429.3


