

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.6 + phase: 0 /pseudo
(408 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
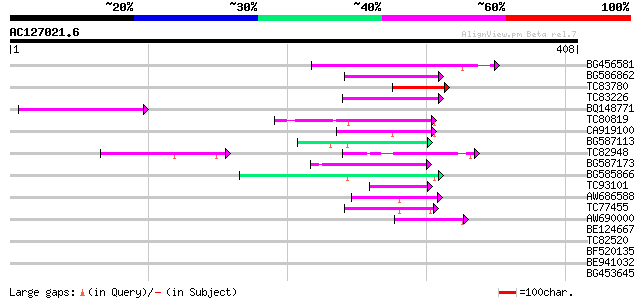
Score E
Sequences producing significant alignments: (bits) Value
BG456581 122 3e-28
BG586862 60 2e-09
TC83780 59 3e-09
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 59 3e-09
BQ148771 53 2e-07
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 51 7e-07
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 50 1e-06
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 48 6e-06
TC82948 47 1e-05
BG587173 weakly similar to GP|4512630|gb| Contains reverse trans... 46 2e-05
BG585866 45 5e-05
TC93101 43 3e-04
AW686588 43 3e-04
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 42 4e-04
AW690000 41 7e-04
BE124667 41 0.001
TC82520 35 0.001
BF520135 39 0.004
BE941032 39 0.005
BG453645 36 0.031
>BG456581
Length = 683
Score = 122 bits (305), Expect = 3e-28
Identities = 61/141 (43%), Positives = 76/141 (53%), Gaps = 6/141 (4%)
Frame = +2
Query: 218 GHLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIV 277
G LT KL Y F+ PW +WN+ IPP+ FICWR H+RLPTD+NL RGC +V
Sbjct: 8 GKLTIKLVYSFLTSHTSCAPWASTIWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALV 187
Query: 278 SICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLSSCQNLF------VKQNWRR 331
S+C FC +Q+ETSDH+FLRC +W WL + LD SS + L R
Sbjct: 188 SMCSFCLEQVETSDHLFLRCKFVVTLWSWLCSQLRVGLDFSSFKALLSSLPRHCSSQVRD 367
Query: 332 QYVGAANSDLNHQKCHMVHMV 352
YV A HMVH +
Sbjct: 368 LYVAAV--------VHMVHSI 406
>BG586862
Length = 804
Score = 60.1 bits (144), Expect = 2e-09
Identities = 27/71 (38%), Positives = 35/71 (49%)
Frame = -1
Query: 242 VWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTS 301
VW P WR +HN LP + L KRG +C C+ ++ET H+FL C VT
Sbjct: 654 VWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQ 475
Query: 302 QIWDWLNQGIN 312
+ W GIN
Sbjct: 474 KEWFGSQLGIN 442
>TC83780
Length = 400
Score = 59.3 bits (142), Expect = 3e-09
Identities = 25/41 (60%), Positives = 29/41 (69%)
Frame = +3
Query: 276 IVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLD 316
IVSICCFC Q +S HIFL CPVT + DWL+ G QQ+D
Sbjct: 3 IVSICCFCHNQAVSSHHIFLSCPVTMVLCDWLSSGTGQQID 125
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 59.3 bits (142), Expect = 3e-09
Identities = 24/73 (32%), Positives = 40/73 (53%)
Frame = +3
Query: 240 KLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPV 299
K +W+ + P + WR +++ LP +LRKRG +C C + ET H+F+ CP+
Sbjct: 123 KKIWSLHTIPRHKVLLWRILNDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPL 302
Query: 300 TSQIWDWLNQGIN 312
+ ++W N IN
Sbjct: 303 SKRVWFGSNLCIN 341
>BQ148771
Length = 680
Score = 52.8 bits (125), Expect = 2e-07
Identities = 30/94 (31%), Positives = 44/94 (45%)
Frame = -3
Query: 7 LSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSGDCNQQK 66
L+ W LS+ R L VI + +Y PK + EI K+ R FVW ++
Sbjct: 573 LANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRR 394
Query: 67 ICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHL 100
V W TM +PK GL +R ++N A ++ L
Sbjct: 393 YHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKL 292
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 51.2 bits (121), Expect = 7e-07
Identities = 38/126 (30%), Positives = 57/126 (45%), Gaps = 9/126 (7%)
Frame = +2
Query: 191 VLTNKILQTTLPIKPVPDALQWLSTTDGHLTNKLTYLFVRGTCIKVPWHKLV---WNTYI 247
+L N +LQ V D +WL + K+ Y ++ T + LV W+ +I
Sbjct: 482 LLNNFVLQDN-----VNDKWRWLLDPVNGYSVKVFYRYITSTG-HISDRSLVDDVWHKHI 643
Query: 248 PPTRVFICWRFIHNRLPTDENLRKRGCTI-VSICCFCK-KQLETSDHIFLRCPVTSQIW- 304
P WR + NRLPT +NL RG + + C C E++ H+FL C V +W
Sbjct: 644 PSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNVFCSLWS 823
Query: 305 ---DWL 307
+WL
Sbjct: 824 LVRNWL 841
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 50.4 bits (119), Expect = 1e-06
Identities = 24/78 (30%), Positives = 40/78 (50%), Gaps = 6/78 (7%)
Frame = -2
Query: 236 VPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGC--TIVSICCFCKKQLETSDHI 293
+P+ +++W+ +P WR + +RLPT NL RG T +C LE++ H+
Sbjct: 602 LPYAEMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHL 423
Query: 294 FLRCPVTSQIW----DWL 307
FL C + +W DW+
Sbjct: 422 FLSCSYFASLWSLVRDWI 369
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 48.1 bits (113), Expect = 6e-06
Identities = 33/109 (30%), Positives = 42/109 (38%), Gaps = 12/109 (11%)
Frame = -3
Query: 208 DALQWLSTTDGHLTNKLTYLFV---------RGTCIKVPWHKL---VWNTYIPPTRVFIC 255
D+ W + GH + K Y RGT + L VW P
Sbjct: 426 DSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFL 247
Query: 256 WRFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIW 304
WR I N LPT N+R R + C C + ET +HI +CP IW
Sbjct: 246 WRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIW 100
>TC82948
Length = 705
Score = 47.4 bits (111), Expect = 1e-05
Identities = 32/101 (31%), Positives = 46/101 (44%), Gaps = 2/101 (1%)
Frame = +3
Query: 240 KLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPV 299
KL W+ I + + R HNR I +C C K E++ H+F C V
Sbjct: 354 KLCWDMMISKEQWYAFMR--HNRALLGH--------IDFMCSLCFKSFESTHHLFFECSV 503
Query: 300 TSQIWDWLNQGINQQLDLSSCQNLFVKQNWR--RQYVGAAN 338
+ QIWDW + IN +++ +L WR + VGAAN
Sbjct: 504 SVQIWDWFSNLINLNFHINNTDDL-----WRICDREVGAAN 611
Score = 42.7 bits (99), Expect = 3e-04
Identities = 29/111 (26%), Positives = 46/111 (41%), Gaps = 17/111 (15%)
Frame = +3
Query: 66 KICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQWAQLCRH-------- 117
K+ V+W +CRP EG L IR +N A L L + S++QW RH
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKEQWYAFMRHNRALLGHI 434
Query: 118 -----ILLSNAKPKNH-YIHSSIWIGLKSWVDVVIQ---HSNLEHW*WKIC 159
+ + + +H + S+ + + W +I H N W+IC
Sbjct: 435 DFMCSLCFKSFESTHHLFFECSVSVQIWDWFSNLINLNFHINNTDDLWRIC 587
>BG587173 weakly similar to GP|4512630|gb| Contains reverse transcriptase
domain (rvt) PF|00078. {Arabidopsis thaliana}, partial
(5%)
Length = 292
Score = 46.2 bits (108), Expect = 2e-05
Identities = 29/87 (33%), Positives = 39/87 (44%)
Frame = +1
Query: 217 DGHLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTI 276
D T+K T+ VR V W +VW + P F W ++RLPT +I
Sbjct: 28 DSFSTSK-TWESVRPRGT*VDWESVVWFSGHIPKHAFNMWVANYDRLPTRCRFAAWDTSI 204
Query: 277 VSICCFCKKQLETSDHIFLRCPVTSQI 303
V C C E+ DH+FLRC + I
Sbjct: 205 VKTCLLCGHYDESRDHLFLRCSFSEHI 285
>BG585866
Length = 828
Score = 45.1 bits (105), Expect = 5e-05
Identities = 39/169 (23%), Positives = 60/169 (35%), Gaps = 22/169 (13%)
Frame = +3
Query: 166 LSMSVSDYIVDGEWNLPTYFHVKDDVLTNKILQTTLPIKP-VPDALQWLSTTDGHLTNKL 224
L ++V D G + + + + + I T L + DA W ++G T K
Sbjct: 219 LHLTVKDVFTTGGQHTQSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKS 398
Query: 225 TYLFVRGTCIKVPWHKL----VWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSIC 280
Y ++ V ++ +W IP F W HN +PT L R +IC
Sbjct: 399 GYSWILSQTETVNYNNSSWSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAIC 578
Query: 281 CFCKKQLETSDHIFLRCPVTSQIW-----------------DWLNQGIN 312
C + E+ H C + IW DWL GI+
Sbjct: 579 SRCGEHEESFFHCVRDCRFSKIIWHKIGFSSPDFFSSSSVQDWLKDGIS 725
>TC93101
Length = 675
Score = 42.7 bits (99), Expect = 3e-04
Identities = 18/45 (40%), Positives = 24/45 (53%)
Frame = -1
Query: 260 HNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIW 304
H LP L KRG +C C LET++HIF+ C T ++W
Sbjct: 672 HTTLPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVW 538
>AW686588
Length = 567
Score = 42.7 bits (99), Expect = 3e-04
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 2/67 (2%)
Frame = +1
Query: 247 IPPTRVFICWRFIHNRLPTDENLRKRGCTIVSI--CCFCKKQLETSDHIFLRCPVTSQIW 304
+P + WR I +RLPT NL +R C V C ET++H+FL C +W
Sbjct: 142 VPLKVSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAVW 321
Query: 305 DWLNQGI 311
+ I
Sbjct: 322 QHIRAWI 342
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 42.0 bits (97), Expect = 4e-04
Identities = 26/74 (35%), Positives = 34/74 (45%), Gaps = 7/74 (9%)
Frame = -2
Query: 242 VWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSI---CCFCKKQLETSDHIFLRCP 298
+W + P + W +R+PT NL KR V C FC Q ET H+FL C
Sbjct: 322 LWKSKAPAKVLAFSWTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHCD 143
Query: 299 VTS----QIWDWLN 308
V S ++ WLN
Sbjct: 142 VISKV*REVMRWLN 101
>AW690000
Length = 652
Score = 41.2 bits (95), Expect = 7e-04
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 2/55 (3%)
Frame = +2
Query: 278 SICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLSSCQNLF--VKQNWR 330
S+C C KQ E+S H+F C +W W +N+ L S ++++ ++WR
Sbjct: 8 SMCSLCCKQAESSLHLFFECSYAVNLWCWFASVLNKTLHFQSLEDMWSLCNRSWR 172
>BE124667
Length = 540
Score = 40.8 bits (94), Expect = 0.001
Identities = 17/57 (29%), Positives = 31/57 (53%), Gaps = 2/57 (3%)
Frame = -1
Query: 278 SICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLSSCQNLF--VKQNWRRQ 332
S+C C+ ET+ H+F C +++W WL +N L S+ +++ ++ NW Q
Sbjct: 390 SMCSSCQASSETTFHLFFECNFATKMWSWLASLLNTPLQFSTSADMWSPLQLNWTPQ 220
>TC82520
Length = 833
Score = 34.7 bits (78), Expect(2) = 0.001
Identities = 15/31 (48%), Positives = 16/31 (51%)
Frame = +3
Query: 242 VWNTYIPPTRVFICWRFIHNRLPTDENLRKR 272
VW IP WR HNRLPT NL +R
Sbjct: 186 VWQKNIPSKVSMFVWRLFHNRLPTKVNLMQR 278
Score = 25.0 bits (53), Expect(2) = 0.001
Identities = 9/25 (36%), Positives = 14/25 (56%), Gaps = 4/25 (16%)
Frame = +2
Query: 288 ETSDHIFLRCPVTSQIWD----WLN 308
ET+ H+FL C + +W WL+
Sbjct: 329 ETATHLFLHCDIFGSLWSHVLRWLH 403
>BF520135
Length = 202
Score = 38.9 bits (89), Expect = 0.004
Identities = 21/64 (32%), Positives = 28/64 (42%), Gaps = 2/64 (3%)
Frame = +3
Query: 238 WHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCT--IVSICCFCKKQLETSDHIFL 295
W L+W WR H RLPT N+ KRG +C + +E+ H+FL
Sbjct: 3 WCLLLWRKEDLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRCRLIESDVHLFL 182
Query: 296 RCPV 299
C V
Sbjct: 183 HCDV 194
>BE941032
Length = 435
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/59 (38%), Positives = 28/59 (46%), Gaps = 6/59 (10%)
Frame = +2
Query: 256 WRFIHNRLPTDENLRKRGC--TIVSICCFCKKQLETSDHIFLRCPVTSQIW----DWLN 308
W + NR PT +NL KRG I C L + H+FL C QIW DWL+
Sbjct: 158 WCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQIWINVSDWLS 334
>BG453645
Length = 622
Score = 35.8 bits (81), Expect = 0.031
Identities = 13/29 (44%), Positives = 18/29 (61%)
Frame = +3
Query: 276 IVSICCFCKKQLETSDHIFLRCPVTSQIW 304
+ S C C ++ET+ HIFL C V S +W
Sbjct: 21 VSSTCVLCNLKVETTTHIFLHCDVASLVW 107
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.142 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,959,683
Number of Sequences: 36976
Number of extensions: 337743
Number of successful extensions: 2646
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 2614
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2643
length of query: 408
length of database: 9,014,727
effective HSP length: 98
effective length of query: 310
effective length of database: 5,391,079
effective search space: 1671234490
effective search space used: 1671234490
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC127021.6


