

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.2 - phase: 0
(377 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
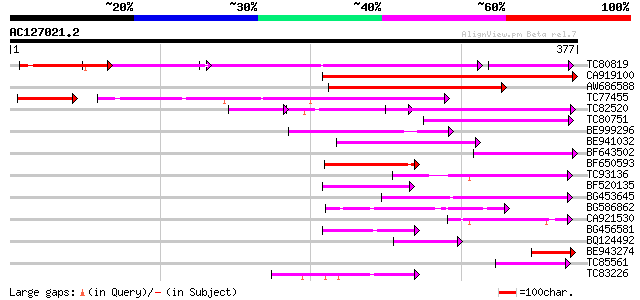
Score E
Sequences producing significant alignments: (bits) Value
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 159 5e-68
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 184 4e-47
AW686588 141 5e-34
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 112 1e-33
TC82520 117 9e-27
TC80751 weakly similar to GP|13161403|dbj|BAB33036. CPRD49 {Vign... 78 6e-15
BE999296 77 1e-14
BE941032 72 5e-13
BF643502 71 6e-13
BF650593 66 3e-11
TC93136 62 5e-10
BF520135 60 1e-09
BG453645 55 4e-08
BG586862 47 9e-06
CA921530 44 8e-05
BG456581 44 8e-05
BQ124492 weakly similar to GP|9909168|dbj| putative transposable... 44 1e-04
BE943274 44 1e-04
TC85561 weakly similar to GP|4530126|gb|AAD21872.1| receptor-lik... 44 1e-04
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 44 1e-04
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 159 bits (401), Expect(3) = 5e-68
Identities = 78/189 (41%), Positives = 106/189 (55%), Gaps = 1/189 (0%)
Frame = +2
Query: 127 EMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPNIGDGYTV 186
+M LGW DG AW W RRL WEEE V E LL N LQD D W W + +GY+V
Sbjct: 383 DMGRLGWTVDGRAWEWTRRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNGYSV 562
Query: 187 RGVYQMLMRQ-EVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDT 245
+ Y+ + + + +V D WHK +P KVS+ WRLLRNR PTKDNLV RGV+
Sbjct: 563 KVFYRYITSTGHISDRSLVDDV-WHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATN 739
Query: 246 QLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNP 305
CV GC +E+ HL +HC +F LW ++ W+G+ S+ ++ H+ QF G
Sbjct: 740 AACVCGCVDSESTTHLFLHCNVFCSLWSLVRNWLGIPSMSSGELRTHFIQFAKMVGMPRV 919
Query: 306 RRSFLQLIW 314
+ +LI+
Sbjct: 920 SHLYFRLIF 946
Score = 79.0 bits (193), Expect(3) = 5e-68
Identities = 37/86 (43%), Positives = 51/86 (59%)
Frame = +3
Query: 49 GRLREGGVTGSAWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFM 108
G L EGG S WW+ I K++ G+G G+WFEE+I +G+G + FFW D W G V
Sbjct: 147 GWLCEGGRQSSMWWKTICKVREGVGEGVGNWFEENIRMVVGDGRDAFFWYDTWAGDVPLR 326
Query: 109 ERFRRLYDLSIHKDLSVGEMHALGWG 134
++ RL DL++ K+ VG GWG
Sbjct: 327 LKYPRLLDLAMDKECKVGLTWG-GWG 401
Score = 64.3 bits (155), Expect = 7e-11
Identities = 32/65 (49%), Positives = 43/65 (65%), Gaps = 3/65 (4%)
Frame = +1
Query: 7 GLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRG---RLREGGVTGSAWWR 63
GLGV FN++LLGKWCWR LVD+EGLW +VL +RYGE+ G R+ + + G +
Sbjct: 31 GLGVGA---FNLSLLGKWCWRLLVDKEGLWHRVLKARYGEEGGGFVRVVDSLLCGGRLYA 201
Query: 64 EIVKI 68
VK+
Sbjct: 202RCVKV 216
Score = 59.3 bits (142), Expect(3) = 5e-68
Identities = 24/57 (42%), Positives = 33/57 (57%)
Frame = +1
Query: 319 WVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLG 375
WV+ ERN RLF NT L++ +K+ S WLK+K V F + Y WW+ C+G
Sbjct: 946 WVIWKERNNRLFQNTVTAPFVLIDRIKLHSFLWLKSKQVGFAYSYLDWWKNSLLCMG 1116
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 184 bits (468), Expect = 4e-47
Identities = 86/169 (50%), Positives = 105/169 (61%)
Frame = -2
Query: 209 WHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIF 268
WH+ VPLKVS+ AWRLLR+R PTK NL+ RGVI + LCV GCG E+ HL + C F
Sbjct: 581 WHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLSCSYF 402
Query: 269 GDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQR 328
LW ++ WIG VD + DH+ QFV+S+G +SFLQLIWL WVL ERN
Sbjct: 401 ASLWSLVRDWIGFVGVDTNVLSDHFVQFVHSTGGNKASQSFLQLIWLLCAWVLWTERNNM 222
Query: 329 LFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLGIG 377
F ++ +LL+ VK SL WLKA+N F FG WW P CLGIG
Sbjct: 221 CFNDSITPLPRLLDKVKYLSLGWLKARNASFLFGTFSWWSNPLQCLGIG 75
>AW686588
Length = 567
Score = 141 bits (355), Expect = 5e-34
Identities = 65/118 (55%), Positives = 81/118 (68%)
Frame = +1
Query: 213 VPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLW 272
VPLKVSI AWRL+R+R PTK NLVRR + + CV GCG ET +HL +HC FG +W
Sbjct: 142 VPLKVSILAWRLIRDRLPTKANLVRRRCLAVEAAGCVVGCGIAETANHLFLHCATFGAVW 321
Query: 273 QRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLF 330
Q I+ WIGV DP + DH+ QF+ +G RRSF+QLIWL +W++ NERN RLF
Sbjct: 322 QHIRAWIGVSGADPHDLSDHFIQFITCTGHTRARRSFMQLIWLLCVWMVWNERNNRLF 495
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 112 bits (279), Expect(2) = 1e-33
Identities = 66/244 (27%), Positives = 119/244 (48%), Gaps = 10/244 (4%)
Frame = -2
Query: 59 SAWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLS 118
S WW++++ ++ V WF + +++G+G ++FFW D W +V E F R +
Sbjct: 778 SRWWKDLMSLEE---VGRVRWFPRELIRKVGDGRSSFFWKDAWDSSVPLRESFPRAF--F 614
Query: 119 IHKDLSVGEMHALGWGEDGEAWR--WRR-RLLAWEEELVVEIRNLLTNVTLQDTQPDVWL 175
++ L +G +G WR WRR L WE+E ++E+ L V L+ D+W+
Sbjct: 613 PYRLLKMGCGDLWDMNAEGVRWRLYWRRLELFEWEKERLLELLGRLEGVVLR-YWADIWV 437
Query: 176 WQPNIGDGYTVRGVYQMLMRQEV------HNHDVVSDAPWHKSVPLKVSICAWRLLRNRW 229
W+P+ ++V Y +L + + +V+ W P KV +W L +R
Sbjct: 436 WKPDKEGVFSVNSCYFLLQNLRLLEDRLSYEEEVIFRELWKSKAPAKVLAFSWTLFLDRI 257
Query: 230 PTKDNLVRRGVI-TNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQ 288
PT NL +R ++ D++ CV ++ET+ HL +HC + + + + W+ + P
Sbjct: 256 PTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHCDVISKV*REVMRWLNFNLISPPN 77
Query: 289 VLDH 292
+L H
Sbjct: 76 LLIH 65
Score = 48.9 bits (115), Expect(2) = 1e-33
Identities = 21/40 (52%), Positives = 27/40 (67%)
Frame = -3
Query: 6 GGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYG 45
GGLGVR ++ N++LL KW WR L D+ LW +VL YG
Sbjct: 951 GGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYG 832
>TC82520
Length = 833
Score = 117 bits (292), Expect = 9e-27
Identities = 49/126 (38%), Positives = 69/126 (53%)
Frame = +2
Query: 251 GCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFL 310
GCG +ET HL +HC IFG LW + W+ + V P + + QF +G SFL
Sbjct: 314 GCGDSETATHLFLHCDIFGSLWSHVLRWLHLLLVLPADIRQFFIQFTSMAGSPRFTHSFL 493
Query: 311 QLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRP 370
Q++W +WVL +RN R+F N+ +E VK+ S WLK + F F YH WW+ P
Sbjct: 494 QIMWFASVWVLWKKRNNRVFQNSLSDPSTFVEQVKMHSFLWLKFQQATFSFSYHDWWKHP 673
Query: 371 FACLGI 376
C+G+
Sbjct: 674 LLCMGV 691
Score = 52.4 bits (124), Expect(2) = 9e-11
Identities = 31/86 (36%), Positives = 45/86 (52%), Gaps = 2/86 (2%)
Frame = +3
Query: 185 TVRGVYQMLM--RQEVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVIT 242
T RG Y+ L Q + + V D W K++P KVS+ WRL NR PTK NL++R V+
Sbjct: 117 TRRGTYRFLTISGQPLDRNQV--DDVWQKNIPSKVSMFVWRLFHNRLPTKVNLMQRHVLQ 290
Query: 243 NDTQLCVRGCGKNETIDHLIIHCLIF 268
C+ G + H+ + +IF
Sbjct: 291 QTHTACISGVAIRKR-QHICFYIVIF 365
Score = 31.6 bits (70), Expect(2) = 9e-11
Identities = 19/40 (47%), Positives = 21/40 (52%)
Frame = +2
Query: 146 LLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPNIGDGYT 185
L AWEEE V E LL N LQ+ DV W + GYT
Sbjct: 2 LFAWEEESVREWYALLHNTVLQENVHDVCRWLLDPI*GYT 121
>TC80751 weakly similar to GP|13161403|dbj|BAB33036. CPRD49 {Vigna
unguiculata}, partial (52%)
Length = 1156
Score = 77.8 bits (190), Expect = 6e-15
Identities = 41/100 (41%), Positives = 52/100 (52%)
Frame = +1
Query: 276 KTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTAR 335
K ++GV S DP + DH Q V G RS L L+W +WV+ ER++ +F
Sbjct: 694 KWFLGVSSTDPPCIPDHLIQLVQ*GGYSRSIRSTLLLVWFSYMWVIWKERSKMIFNQKKD 873
Query: 336 TTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLG 375
QLLE VK+ S WLKA N FPF YH P C+G
Sbjct: 874 HVHQLLEKVKLLSF*WLKATNSTFPFDYHSRRLNPLLCMG 993
>BE999296
Length = 384
Score = 76.6 bits (187), Expect = 1e-14
Identities = 42/110 (38%), Positives = 54/110 (48%)
Frame = -1
Query: 186 VRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDT 245
+ G YQ LM + D+ WH +PLKV R+LRN PTKDN VRR VI +
Sbjct: 306 IHGAYQFLMSADAPLDREYIDSVWHNHIPLKVCFFVLRVLRNCLPTKDNFVRRRVIHEEH 127
Query: 246 QLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQ 295
LC GC ET D L LW + W+ + V+ + DH+YQ
Sbjct: 126 MLCPTGCSFKETTDDLF--------LWPLV*QWLHIS*VNSCVLRDHFYQ 1
>BE941032
Length = 435
Score = 71.6 bits (174), Expect = 5e-13
Identities = 37/96 (38%), Positives = 50/96 (51%)
Frame = +2
Query: 218 SICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKT 277
SI W +L NR+PTKDNL++RGVI+ Q CV CG HL +HC F +W +
Sbjct: 146 SIFLWCVLLNRFPTKDNLLKRGVISAIYQSCVGECGNLYDATHLFLHCNFFRQIWINVSD 325
Query: 278 WIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLI 313
W+ V ++ D QF G N + +QLI
Sbjct: 326 WLSFVMVTLLRISDRLAQFGSFGGFSNCKWDTIQLI 433
>BF643502
Length = 631
Score = 71.2 bits (173), Expect = 6e-13
Identities = 33/69 (47%), Positives = 41/69 (58%)
Frame = -1
Query: 309 FLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQ 368
F+QLIWL +WV+ NERN +F + LL+ +K+ SL WLKAKN F FG
Sbjct: 502 FMQLIWLLCVWVVWNERNNVMFNHVVTPIPCLLDKIKLLSLGWLKAKNATFVFGIQQGRS 323
Query: 369 RPFACLGIG 377
P CLGIG
Sbjct: 322 NPLVCLGIG 296
>BF650593
Length = 486
Score = 65.9 bits (159), Expect = 3e-11
Identities = 29/63 (46%), Positives = 39/63 (61%)
Frame = +3
Query: 210 HKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFG 269
HKSVPLKVS WRL +N T+DNL +RGV+ ++ CV CG+ E++ H C F
Sbjct: 297 HKSVPLKVSCLVWRLFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFEC-PFS 473
Query: 270 DLW 272
+W
Sbjct: 474 XVW 482
Score = 36.2 bits (82), Expect = 0.021
Identities = 18/42 (42%), Positives = 24/42 (56%)
Frame = +1
Query: 34 GLWFKVLSSRYGEQRGRLREGGVTGSAWWREIVKIQNGIGVE 75
GLWF+ L ++YG RG + S WW++I I G GVE
Sbjct: 85 GLWFRALVNKYGLNRGSITIENRGVSLWWKDICYIDFG-GVE 207
>TC93136
Length = 722
Score = 61.6 bits (148), Expect = 5e-10
Identities = 34/122 (27%), Positives = 54/122 (43%), Gaps = 2/122 (1%)
Frame = +1
Query: 255 NETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYN--PRRSFLQL 312
NET HL +HC +W +I W+ +Y +FV R + L
Sbjct: 259 NETSSHLFLHCPFLSSVWSKILGWL------------YYSKFVCVFRLLERRSRTKGVWL 402
Query: 313 IWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFA 372
IW IWV+ N R+F N ++ +++E +K+ S +W + C P Y+ W P
Sbjct: 403 IWHATIWVIWKGINNRIFKNISKAIDEIVEEIKVLSWRWSLTRLKCPPCFYYEWCWDPGD 582
Query: 373 CL 374
C+
Sbjct: 583 CV 588
>BF520135
Length = 202
Score = 60.5 bits (145), Expect = 1e-09
Identities = 27/61 (44%), Positives = 36/61 (58%)
Frame = +3
Query: 209 WHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIF 268
W K KVSI AWRL R PTK N+ +RG++ +D +CV C E+ HL +HC +
Sbjct: 18 WRKEDLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRCRLIESDVHLFLHCDVL 197
Query: 269 G 269
G
Sbjct: 198 G 200
>BG453645
Length = 622
Score = 55.1 bits (131), Expect = 4e-08
Identities = 33/127 (25%), Positives = 55/127 (42%)
Frame = +3
Query: 248 CVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRR 307
CV K ET H+ +HC + +W R+ W V + P + H+ + G N
Sbjct: 33 CVLCNLKVETTTHIFLHCDVASLVWFRVMKWFDVNFLIPPNLFVHW-ECWSEGGSANRVT 209
Query: 308 SFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWW 367
L+W IWVL +RN +F ++E +K+ S +W+ ++ ++ W
Sbjct: 210 KGHWLVWHTTIWVLWAKRNDLIFKGLNCVAEDVIEEIKVLS*RWMLERSSTPSCFFYEWS 389
Query: 368 QRPFACL 374
P CL
Sbjct: 390 WNPRLCL 410
>BG586862
Length = 804
Score = 47.4 bits (111), Expect = 9e-06
Identities = 37/122 (30%), Positives = 60/122 (48%)
Frame = -1
Query: 211 KSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGD 270
K++P S WRLL N P KD L +RG+ + LC R K ET+ HL ++C +
Sbjct: 642 KTIPRHKSFL-WRLLHNALPVKDELHKRGI--RCSLLCPRCESKIETVQHLFLNCEVTQK 472
Query: 271 LWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLF 330
W + I S VL H++ ++ + N + + L L ++ + + RNQ++F
Sbjct: 471 EWFGSQLGINFHS---SGVL-HFHDWITNFILKNDEETIIALTAL--LYSIWHARNQKVF 310
Query: 331 TN 332
N
Sbjct: 309 EN 304
>CA921530
Length = 793
Score = 44.3 bits (103), Expect = 8e-05
Identities = 30/92 (32%), Positives = 41/92 (43%), Gaps = 9/92 (9%)
Frame = -1
Query: 292 HYYQFVYSSGCYN------PRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVK 345
H+ F+ S C+N R L LIW IW+L N R+F N A ++E VK
Sbjct: 670 HFIMFIMSQ-CWNGWERNKKIRRGLWLIWHATIWILWKAWNNRIFNNQAVEAEDIIEEVK 494
Query: 346 ITSLKWLKAK---NVCFPFGYHMWWQRPFACL 374
+ S +W + VC Y+ W P CL
Sbjct: 493 VLSWRWTFNRINIPVCM---YYEWCWNPKYCL 407
>BG456581
Length = 683
Score = 44.3 bits (103), Expect = 8e-05
Identities = 23/64 (35%), Positives = 29/64 (44%)
Frame = +2
Query: 209 WHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIF 268
W+ +P S WRL +R PT DNL RG +C + ET DHL + C
Sbjct: 83 WNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCAL--VSMCSFCLEQVETSDHLFLRCKFV 256
Query: 269 GDLW 272
LW
Sbjct: 257 VTLW 268
>BQ124492 weakly similar to GP|9909168|dbj| putative transposable element
Tip100 protein {Oryza sativa (japonica cultivar-group)},
partial (8%)
Length = 694
Score = 43.9 bits (102), Expect = 1e-04
Identities = 17/46 (36%), Positives = 27/46 (57%)
Frame = +2
Query: 256 ETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSG 301
+T HL + C IF LW ++ W+G+ SV P ++ H+ QF +G
Sbjct: 530 KTARHLFLDCDIFSSLWSQVWLWLGISSVQPGELRHHFPQFTNMAG 667
>BE943274
Length = 597
Score = 43.5 bits (101), Expect = 1e-04
Identities = 17/29 (58%), Positives = 19/29 (64%)
Frame = -3
Query: 348 SLKWLKAKNVCFPFGYHMWWQRPFACLGI 376
SL WLK K+V F FG +WW P CLGI
Sbjct: 331 SLWWLKVKHVTFVFGIQIWWSNPLMCLGI 245
>TC85561 weakly similar to GP|4530126|gb|AAD21872.1| receptor-like protein
kinase homolog RK20-1 {Phaseolus vulgaris}, partial (18%)
Length = 1549
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/50 (38%), Positives = 26/50 (52%)
Frame = +2
Query: 324 ERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFAC 373
ERN R+F + + QLL ++K S WLK + F F Y+ WW C
Sbjct: 1265 ERNSRIFNDNMSSKDQLLASIKFHS*WWLKMHKLGFAFDYYYWWSMASLC 1414
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/107 (28%), Positives = 47/107 (42%), Gaps = 9/107 (8%)
Frame = +3
Query: 175 LWQPNIGDGYTVRGVYQML---MRQEVHNHDVVSDAP--WHKSVPLKV----SICAWRLL 225
+W N Y+V+ Y L Q+++N SD W K L + WR+L
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 226 RNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLW 272
+ P + +L +RG+ LC R K ETI HL + C + +W
Sbjct: 183 NDSLPVRSSLRKRGI--QCYPLCPRCHSKTETITHLFMSCPLSKRVW 317
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.141 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,158,208
Number of Sequences: 36976
Number of extensions: 280981
Number of successful extensions: 2237
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 2193
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2227
length of query: 377
length of database: 9,014,727
effective HSP length: 98
effective length of query: 279
effective length of database: 5,391,079
effective search space: 1504111041
effective search space used: 1504111041
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC127021.2


