

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127020.7 + phase: 0 /pseudo
(1459 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
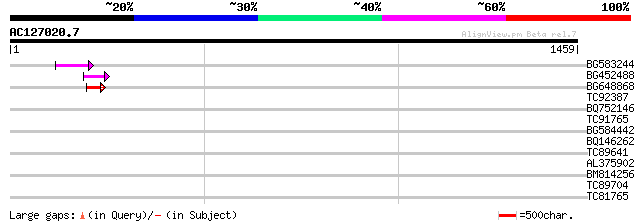
Score E
Sequences producing significant alignments: (bits) Value
BG583244 74 4e-13
BG452488 68 2e-11
BG648868 53 1e-06
TC92387 40 0.009
BQ752146 similar to GP|7958612|gb|A PalH {Emericella nidulans}, ... 35 0.16
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 34 0.36
BG584442 33 0.62
BQ146262 33 0.81
TC89641 similar to GP|9988432|dbj|BAB12698.1 hypothetical protei... 31 4.0
AL375902 similar to GP|9759134|dbj mutator-like transposase-like... 30 6.9
BM814256 similar to PIR|D96745|D96 hypothetical protein T9N14.9 ... 30 9.0
TC89704 weakly similar to GP|15010754|gb|AAK74036.1 AT3g08600/F1... 30 9.0
TC81765 similar to GP|12323731|gb|AAG51827.1 hypothetical protei... 30 9.0
>BG583244
Length = 633
Score = 73.9 bits (180), Expect = 4e-13
Identities = 36/100 (36%), Positives = 57/100 (57%), Gaps = 2/100 (2%)
Frame = -1
Query: 119 VPLNHIEFWVQVHGLPFGFVQPKVGQGIGSFLGTLKAYDVRNTI--HSSYMRIKVAIDVT 176
+P+N I+ WVQ H LPFGF+ +G +GS + + +D N Y++I+V I++
Sbjct: 300 IPMNTIDIWVQDHQLPFGFMDTTIGALVGSHIC*MVIFDEENNYGPWRKYVKIRVEIEIE 121
Query: 177 VPLKKEWRVRASNGNFVTVNFKYEKLGVFCHRCGVLGHTD 216
PLK++ + + + FKYEKL FC CG +GH +
Sbjct: 120 QPLKQDLILEGDMCKNIWLVFKYEKLDKFCFVCGSIGHRE 1
>BG452488
Length = 643
Score = 68.2 bits (165), Expect = 2e-11
Identities = 31/69 (44%), Positives = 41/69 (58%), Gaps = 1/69 (1%)
Frame = +1
Query: 189 NGNFVTVNFKYEKLGVFCHRCGVLGHTDKVCPELFELDSDDGVRNWGPYLK-PVTQRVGT 247
+G + VNF+YEK G FC+ CG++GHTD CP+ FE +G + WGPYL+ G
Sbjct: 19 DGAVIRVNFRYEKPGNFCYECGLIGHTDGSCPKRFEKGFVEGQQQWGPYLRSDYVGPEGG 198
Query: 248 AATNRWLQD 256
N WL D
Sbjct: 199 MIENPWLHD 225
>BG648868
Length = 801
Score = 52.8 bits (125), Expect = 1e-06
Identities = 28/50 (56%), Positives = 32/50 (64%), Gaps = 2/50 (4%)
Frame = -2
Query: 198 KYEKLGVFCHRCGVLGHTDK--VCPELFELDSDDGVRNWGPYLKPVTQRV 245
K+EK+G F CG+LGHTDK LFEL SDDG R WGP VT R+
Sbjct: 797 KHEKIGDFYFLCGLLGHTDKSFSLARLFELASDDGARQWGP--DRVTSRI 654
>TC92387
Length = 713
Score = 39.7 bits (91), Expect = 0.009
Identities = 23/68 (33%), Positives = 35/68 (50%)
Frame = +2
Query: 40 LIGGVLADREIQFAYFSERMSRAWKPGKRVTITKSVADRYLFQFHHKVDDARVLEEGPWL 99
L+G +L +R +++ W K V I K + Y F FHH +D +L++ PW
Sbjct: 203 LVGRLLTNRPN*SYVMIKKIEAFWSLVKGVKIQKI*SGLYTF*FHHHLDLQSILKKCPWN 382
Query: 100 YDNFHIVL 107
+DN IVL
Sbjct: 383 FDNGIIVL 406
>BQ752146 similar to GP|7958612|gb|A PalH {Emericella nidulans}, partial (5%)
Length = 641
Score = 35.4 bits (80), Expect = 0.16
Identities = 40/153 (26%), Positives = 64/153 (41%), Gaps = 2/153 (1%)
Frame = +3
Query: 226 DSDDGVRNWGPYLKPVTQRVGTAATNRWLQDPIPAVTPSHNSDVAAASVGRKTQTAGTGV 285
++ +G R+ G L+P W P PAVTP +D A+A A T
Sbjct: 240 NAQNGQRSMGGLLRPPM----------WPSRPAPAVTPVSRTDTASA--------ASTVY 365
Query: 286 VTNFH--DRMTAFQTQLTAMRHDVLAAQNAVLVKKGKGAHSACQVHLLPSSSTGTSASNL 343
+H T+ ++ R D + Q V + + + S+ SSS+ S S+
Sbjct: 366 AVRYHLPSEATSRTPEVANQRGDQPSPQRHVAIPMSRSSSSS------SSSSSSRSGSST 527
Query: 344 LPDRPIVLGLPAAPL*SENTDDALEEIGSELKK 376
D P + AAP+ + T A+ EI +E +K
Sbjct: 528 ETDEPSL----AAPVPRQATHKAV-EIDAEAQK 611
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 34.3 bits (77), Expect = 0.36
Identities = 33/104 (31%), Positives = 44/104 (41%)
Frame = +1
Query: 1125 SFLK*MRTKLLS*KISCLFMKQRPDRPLVYQRVRFIVAKMSRILSNRTLLLFLAYKWCWA 1184
S LK + KL *K S ++ + LV+ R+I A M +F + CW
Sbjct: 310 SELKSIMRKL--*KTSLFCTRKTLVKLLVFANHRYIAAGMFLTF*RLPSHIFWGFNLCWV 483
Query: 1185 HVSTLVYLL*LAEIVIQLLHTSKTVCGIKLILGVESACLKQGAR 1228
HVS + * AEI QL + CG K I CL + R
Sbjct: 484 HVSIWDFRP*SAEIEQQLFLLLRVECG-KKINSWSDICLSKACR 612
>BG584442
Length = 775
Score = 33.5 bits (75), Expect = 0.62
Identities = 33/96 (34%), Positives = 49/96 (50%)
Frame = +2
Query: 1213 KLILGVESACLKQGARL**NLFCRQFLLML*VFSSCQRL*SIQLRR**IPFGGVMDEQRS 1272
KLI + C K R * ++ + +L M *VF S L ++L+R *I F G M E+ +
Sbjct: 383 KLIFRETNVCQKLCER**LSMHSKAYLPM**VFFSF*ILK*MKLKRL*ILFHGFMLEKIA 562
Query: 1273 EV*TG*TGKSYPHQRYMEVWVSKISPLLI*LC*VNR 1308
+ G ++ + + ME WV + S I C VNR
Sbjct: 563 KECIGCLRRNCLYIKIMEAWVLQTSRRSIFQCLVNR 670
>BQ146262
Length = 687
Score = 33.1 bits (74), Expect = 0.81
Identities = 21/86 (24%), Positives = 40/86 (46%)
Frame = -3
Query: 158 VRNTIHSSYMRIKVAIDVTVPLKKEWRVRASNGNFVTVNFKYEKLGVFCHRCGVLGHTDK 217
+R+ + Y+ I + +D++ L V G V +YE+ FC+ C +LGH+ +
Sbjct: 598 IRSXLFGHYVGILMDVDMSSKLFDSVVVERE-GFVFPVAVEYERKPSFCYNCKLLGHSIQ 422
Query: 218 VCPELFELDSDDGVRNWGPYLKPVTQ 243
+C + + +N KP+ Q
Sbjct: 421 LCNRINNIQHHAAPKN--QQKKPIQQ 350
>TC89641 similar to GP|9988432|dbj|BAB12698.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (44%)
Length = 1080
Score = 30.8 bits (68), Expect = 4.0
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +3
Query: 200 EKLGVFCHRCGVLGHTDKVCPELFELDSDDGVRNWGPYL 238
+K+G C CG +GH + C L S GVR +G L
Sbjct: 78 QKMGRKCSHCGNIGHNSRTCNSLRGSGSFVGVRLFGVQL 194
>AL375902 similar to GP|9759134|dbj mutator-like transposase-like protein
{Arabidopsis thaliana}, partial (20%)
Length = 542
Score = 30.0 bits (66), Expect = 6.9
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = +2
Query: 196 NFKYEKLGVFCHRCGVLGHTDKVC 219
NFK K V C RC +LGH+ K C
Sbjct: 302 NFKRPKRVVQCGRCHMLGHSQKKC 373
>BM814256 similar to PIR|D96745|D96 hypothetical protein T9N14.9 [imported] -
Arabidopsis thaliana, partial (37%)
Length = 599
Score = 29.6 bits (65), Expect = 9.0
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +3
Query: 191 NFVTVNFKYEKLGVFCHRCGVLGH 214
N +T N + +GVFC+ C + GH
Sbjct: 210 NHITNNLSHGSIGVFCNACNMRGH 281
>TC89704 weakly similar to GP|15010754|gb|AAK74036.1 AT3g08600/F17O14_7
{Arabidopsis thaliana}, partial (39%)
Length = 1315
Score = 29.6 bits (65), Expect = 9.0
Identities = 19/59 (32%), Positives = 33/59 (55%)
Frame = +3
Query: 1092 SLGTPSLGVFSRALKFVVKLPLSLTFYLLTIVFSFLK*MRTKLLS*KISCLFMKQRPDR 1150
SL T SLG + R K V+ L L ++F T++FSF + ++ +C F+++ +R
Sbjct: 60 SLFTLSLGEYQRKSKRVMDLLLRISF*STTLIFSFFRFFCSQ------NCCFIERERER 218
>TC81765 similar to GP|12323731|gb|AAG51827.1 hypothetical protein;
55509-55015 {Arabidopsis thaliana}, partial (25%)
Length = 710
Score = 29.6 bits (65), Expect = 9.0
Identities = 21/76 (27%), Positives = 39/76 (50%), Gaps = 1/76 (1%)
Frame = +2
Query: 634 IVHL-LIICGFRIFPLLMLKLWWLRLLITTQFTSILLLHLGPMLLSVIFAMKMRGILNRA 692
++HL L++ ++FP+L L WL +I Q I + M++ V + ++
Sbjct: 401 LIHLRLLLLQSQLFPILSL*FHWLLTMIVKQ---IQKKKVALMMMVVKMTAILMTVIMET 571
Query: 693 SRISLLILGRNILLIR 708
ISL L +N+LL++
Sbjct: 572 MAISLCPLIKNLLLVK 619
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.351 0.158 0.526
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,726,138
Number of Sequences: 36976
Number of extensions: 712205
Number of successful extensions: 6268
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2628
Number of HSP's successfully gapped in prelim test: 358
Number of HSP's that attempted gapping in prelim test: 3498
Number of HSP's gapped (non-prelim): 3264
length of query: 1459
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1351
effective length of database: 5,021,319
effective search space: 6783801969
effective search space used: 6783801969
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC127020.7


