

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127019.14 + phase: 0 /pseudo
(532 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
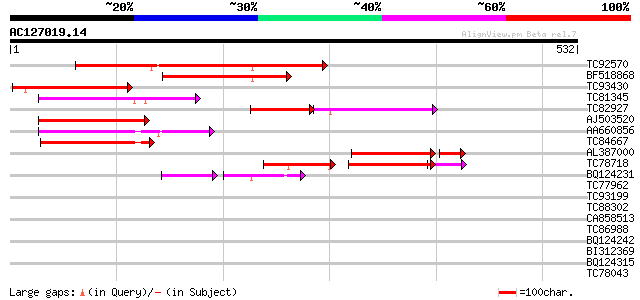
Score E
Sequences producing significant alignments: (bits) Value
TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane... 258 3e-69
BF518868 similar to GP|12541097|em unnamed protein product {Arab... 204 5e-53
TC93430 weakly similar to GP|10177112|dbj|BAB10402. Mlo protein-... 148 4e-36
TC81345 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtM... 142 3e-34
TC82927 similar to GP|12541095|emb|CAC25084. unnamed protein pro... 88 3e-33
AJ503520 weakly similar to GP|15290605|gb| seven transmembrane p... 122 3e-28
AA660856 similar to GP|12541099|em unnamed protein product {Arab... 107 1e-23
TC84667 similar to GP|12541095|emb|CAC25084. unnamed protein pro... 100 1e-21
AL387000 similar to GP|15290605|gb seven transmembrane protein M... 91 1e-19
TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtM... 83 1e-16
BQ124231 similar to GP|13784981|gb seven transmembrane protein M... 54 5e-15
TC77962 similar to PIR|S49848|S49848 probable narbonin - jack be... 31 1.0
TC93199 weakly similar to GP|9755448|gb|AAF98209.1| Unknown prot... 30 2.3
TC88302 similar to GP|8953714|dbj|BAA98077.1 contains similarity... 30 3.0
CA858513 weakly similar to PIR|S61428|S614 embryonic abundant pr... 29 5.1
TC86988 weakly similar to GP|17529294|gb|AAL38874.1 unknown prot... 29 5.1
BQ124242 similar to PIR|S12404|A330 conglutin delta precursor - ... 29 5.1
BI312369 29 5.1
BQ124315 similar to GP|23510634|emb hypothetical protein {Plasmo... 29 5.1
TC78043 weakly similar to GP|4753801|emb|CAB42052.1 nicotianamin... 29 5.1
>TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane protein
MLO2 {Oryza sativa} [Oryza sativa (indica
cultivar-group)], partial (47%)
Length = 944
Score = 258 bits (660), Expect = 3e-69
Identities = 130/257 (50%), Positives = 172/257 (66%), Gaps = 20/257 (7%)
Frame = +3
Query: 62 WLGKREKKALLEALEKVKAELMILGFISLLLTIGQNYIVRICISEKIANKMLPCPHKYIG 121
W K+ KKA+ EALEK+K ELM+LGFISLLLT G YI +ICI K+ + MLPC +
Sbjct: 60 WFHKKHKKAMSEALEKIKEELMLLGFISLLLTFGTKYIAKICIPNKLGDSMLPCKKGEVE 239
Query: 122 GDKKSNSRYL-----------------AADTPSFKCSRKGHEPLLSVNGLHQLHILIFFL 164
+ K + R L AA + CS KG PL+S +G+HQLHI IF L
Sbjct: 240 EESKDDRRRLLSFDFDENVVWRRGLAPAAGGDDY-CSNKGQVPLISQSGVHQLHIFIFVL 416
Query: 165 AVFHVFYSAVTMLLGRLKIRGWKEWEEETLTHGYEFANDAAKLRLTHETSFVRAHTSFWT 224
AVFH+FYS +T++L R K++ WK WE ET + Y+F +D ++ R H+T+FV+ H+ +
Sbjct: 417 AVFHIFYSVMTIVLARAKMQKWKAWETETSSVEYQFTHDPSRFRFAHQTTFVKRHSGWSE 596
Query: 225 RV---YIGCFFRQFYRSVRKVDYQTLRNGFINVHLAPGSKFNFQKYIKRSLEDDFKVVVG 281
+ +I FFRQF+ SV KVDY T+R+GFIN H AP SKF+F KYIKRS+EDDFKVVVG
Sbjct: 597 KPGIRWIVAFFRQFFASVSKVDYMTMRHGFINAHFAPDSKFDFHKYIKRSMEDDFKVVVG 776
Query: 282 VSPVLWASAVVFLLLNI 298
+S LWA A++FLL+N+
Sbjct: 777 ISVPLWAFAIIFLLMNV 827
>BF518868 similar to GP|12541097|em unnamed protein product {Arabidopsis
thaliana}, partial (20%)
Length = 378
Score = 204 bits (520), Expect = 5e-53
Identities = 97/125 (77%), Positives = 105/125 (83%), Gaps = 4/125 (3%)
Frame = +3
Query: 144 GHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLGRLKIRGWKEWEEETLTHGYEFAND 203
G EPL+SVNGLHQLHILIFFLAV HV YSAVTMLLGRLKIRGWK WE ET +HGYEFA D
Sbjct: 3 GQEPLISVNGLHQLHILIFFLAVLHVIYSAVTMLLGRLKIRGWKAWEAETSSHGYEFATD 182
Query: 204 AAKLRLTHETSFVRAHTSFWTRV----YIGCFFRQFYRSVRKVDYQTLRNGFINVHLAPG 259
++ RLTHETSFVR H +FWT++ YI CFFRQFYRSV K DY TLRNGFI VHL+PG
Sbjct: 183 PSRFRLTHETSFVRQHATFWTKIPIFFYISCFFRQFYRSVEKSDYLTLRNGFIAVHLSPG 362
Query: 260 SKFNF 264
SKFNF
Sbjct: 363 SKFNF 377
>TC93430 weakly similar to GP|10177112|dbj|BAB10402. Mlo protein-like
{Arabidopsis thaliana}, partial (15%)
Length = 775
Score = 148 bits (374), Expect = 4e-36
Identities = 76/118 (64%), Positives = 92/118 (77%), Gaps = 5/118 (4%)
Frame = +3
Query: 3 KRVFSCLCLFY----GGLSMAAS-DSRNSSRTLDQTPTWGVAAVCTVFILISIALEKSLH 57
+R+F CL LF G ++MAAS + SS+ LD+TPTW V VCTVFIL+SI LEKSLH
Sbjct: 294 QRLFLCLVLFAWLYNGSVAMAASSEDSGSSKDLDKTPTWAVGCVCTVFILVSIILEKSLH 473
Query: 58 KLGTWLGKREKKALLEALEKVKAELMILGFISLLLTIGQNYIVRICISEKIANKMLPC 115
K+GTWLG R KKALLEALEKVKAELMILGF+SLLLT GQ+Y+V++C+ + MLPC
Sbjct: 474 KIGTWLGDRHKKALLEALEKVKAELMILGFLSLLLTFGQSYMVKLCMPANATDIMLPC 647
>TC81345 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtMlo12).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (24%)
Length = 654
Score = 142 bits (358), Expect = 3e-34
Identities = 77/170 (45%), Positives = 101/170 (59%), Gaps = 18/170 (10%)
Frame = +2
Query: 28 RTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMILGF 87
RTL++TPTW VA VC V + ISI +E +H +G W K+ K AL EALEKVK ELM++GF
Sbjct: 143 RTLEETPTWAVAVVCFVLLAISIVIEHIIHAIGKWFKKKNKNALYEALEKVKGELMLMGF 322
Query: 88 ISLLLTIGQNYIVRICISEKIANKMLPC---PHKYIGGDKKS---------------NSR 129
ISLLLT Q+YI +ICISEK+ + PC K D+ S N R
Sbjct: 323 ISLLLTEYQDYISKICISEKVGSTWHPCSTPKTKTASNDENSESENHDRKLLEYFDPNPR 502
Query: 130 YLAADTPSFKCSRKGHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLG 179
+ A +C+ KG L+S G+H+LHI IF LA+FH+ +T+ LG
Sbjct: 503 RILATKGYDQCADKGKVALVSAYGIHELHIFIFVLAIFHILQRIITLALG 652
>TC82927 similar to GP|12541095|emb|CAC25084. unnamed protein product
{Arabidopsis thaliana}, partial (44%)
Length = 810
Score = 88.2 bits (217), Expect(2) = 3e-33
Identities = 41/124 (33%), Positives = 72/124 (58%), Gaps = 8/124 (6%)
Frame = +2
Query: 286 LWASAVVFLLLNID--------AYIIFRMAYSNMGILDSNYSRMALEISERHIVVQGMPL 337
LW V+FLLLNI+ A++ + S L+ +++A E++E+H ++G +
Sbjct: 350 LWLFVVIFLLLNINGWHTYFWIAFVPVILLLSVGTKLEHVITQLAHEVAEKHAAIEGDLV 529
Query: 338 VQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKVAF 397
VQ SD +FW +P +VL LIHF LFQN+F+I + W W ++G ++C + + ++
Sbjct: 530 VQPSDNHFWFHRPHIVLFLIHFILFQNAFEIAFFFWIWVTYGFDSCIMGQVRYIVPRLVI 709
Query: 398 GLYM 401
G ++
Sbjct: 710 GXFI 721
Score = 72.0 bits (175), Expect(2) = 3e-33
Identities = 35/60 (58%), Positives = 41/60 (68%)
Frame = +1
Query: 227 YIGCFFRQFYRSVRKVDYQTLRNGFINVHLAPGSKFNFQKYIKRSLEDDFKVVVGVSPVL 286
++ FF+QFY SV K DY TLR GFI H KFNF KY+ R+LEDDFK VVG+ VL
Sbjct: 91 WLQSFFKQFYGSVTKSDYVTLRLGFITTHCKTNPKFNFHKYMIRALEDDFKQVVGIRQVL 270
>AJ503520 weakly similar to GP|15290605|gb| seven transmembrane protein MLO2
{Oryza sativa} [Oryza sativa (indica cultivar-group)],
partial (16%)
Length = 339
Score = 122 bits (306), Expect = 3e-28
Identities = 57/104 (54%), Positives = 73/104 (69%)
Frame = +2
Query: 28 RTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMILGF 87
RTL +TPTW VAAVC VF++IS+ +E +HKL W K+ KKA+ EALEK+K ELM+LGF
Sbjct: 26 RTLKETPTWAVAAVCAVFVIISVLIEHGIHKLEKWFHKKHKKAMSEALEKIKEELMLLGF 205
Query: 88 ISLLLTIGQNYIVRICISEKIANKMLPCPHKYIGGDKKSNSRYL 131
ISLLLT G YI +ICI K+ + MLPC + + K + R L
Sbjct: 206 ISLLLTFGTKYIAKICIPNKLGDSMLPCKKGEVEEESKDDRRRL 337
>AA660856 similar to GP|12541099|em unnamed protein product {Arabidopsis
thaliana}, partial (29%)
Length = 633
Score = 107 bits (267), Expect = 1e-23
Identities = 61/182 (33%), Positives = 95/182 (51%), Gaps = 17/182 (9%)
Frame = +1
Query: 28 RTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMILGF 87
R+L +TPT+ VA+V TV + + +E+S+++ G WL K +KAL +LEK+K ELM+LG
Sbjct: 37 RSLAETPTYSVASVVTVMVFVCFLVERSIYRFGKWLKKTRRKALFASLEKIKEELMLLGL 216
Query: 88 ISLLLTIGQNYIVRICISEKI-ANKMLPCPHKYIGGDKKSNSRYLAADTPSF-------- 138
ISLLL +I IC++ + +++ C + D + ++ SF
Sbjct: 217 ISLLLAQSARWISEICVNSSLFSSRFYTCSEQ----DLPMIEHVMIVNSSSFSDETTIPK 384
Query: 139 --------KCSRKGHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLGRLKIRGWKEWE 190
+C +GHEP +S GL QLH +F L + V YS + + KI W WE
Sbjct: 385 GLYSGALHQCG-EGHEPFVSYEGLEQLHRFLFVLGITXVLYSCLAVGXAMSKIYSWCXWE 561
Query: 191 EE 192
+
Sbjct: 562 NQ 567
>TC84667 similar to GP|12541095|emb|CAC25084. unnamed protein product
{Arabidopsis thaliana}, partial (16%)
Length = 385
Score = 100 bits (250), Expect = 1e-21
Identities = 51/107 (47%), Positives = 72/107 (66%)
Frame = +3
Query: 30 LDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMILGFIS 89
L+ TPTW VA VCTV + IS+A+E+ LH G +L +++K+L EAL+K+K ELM+LGFIS
Sbjct: 6 LEFTPTWVVAGVCTVIVAISLAVERLLHYGGKFLKSKDQKSLYEALQKIKEELMLLGFIS 185
Query: 90 LLLTIGQNYIVRICISEKIANKMLPCPHKYIGGDKKSNSRYLAADTP 136
LLLT+ QN + +IC+ + MLPC + DKK + + A P
Sbjct: 186 LLLTVSQNRLTKICVPPGVLRHMLPCSLE----DKKESHSHSAFSFP 314
>AL387000 similar to GP|15290605|gb seven transmembrane protein MLO2 {Oryza
sativa} [Oryza sativa (indica cultivar-group)], partial
(24%)
Length = 464
Score = 90.9 bits (224), Expect(2) = 1e-19
Identities = 36/79 (45%), Positives = 55/79 (69%)
Frame = +1
Query: 321 MALEISERHIVVQGMPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSFGL 380
MA EI +R +V+G+P+V+ +++YFW +P ++ LIHF LFQN+FQI Y LWTWY F +
Sbjct: 10 MAQEIQDRTTIVRGVPVVEPNNKYFWFNRPHWIIFLIHFTLFQNAFQIAYFLWTWYEFTI 189
Query: 381 ENCFRLDYKLAIIKVAFGL 399
+CF + L + +V G+
Sbjct: 190 TSCFHENLPLIVTRVVLGV 246
Score = 23.9 bits (50), Expect(2) = 1e-19
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +2
Query: 404 *HRWVQG*KNQYLMNKHPRR*RNG 427
*HRW *+ +YL +K + +NG
Sbjct: 296 *HRWDLT*REEYLRSKQQKHLKNG 367
>TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtMlo12).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (42%)
Length = 1625
Score = 83.2 bits (204), Expect = 2e-16
Identities = 42/71 (59%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Frame = +3
Query: 239 VRKVDYQTLRNGFINVHLAPGS--KFNFQKYIKRSLEDDFKVVVGVSPVLWASAVVFLLL 296
+ +VDY LR+GFI HLAPG+ +F+FQKYI RSLE DFKVVVG+SP +W AV+FLL
Sbjct: 3 ISRVDYLALRHGFIMAHLAPGNDAEFDFQKYISRSLEKDFKVVVGISPTIWFFAVLFLLT 182
Query: 297 NI--DAYIIFR 305
N +++IFR
Sbjct: 183 NTHGKSFLIFR 215
Score = 73.9 bits (180), Expect(2) = 1e-16
Identities = 30/81 (37%), Positives = 49/81 (60%)
Frame = +2
Query: 319 SRMALEISERHIVVQGMPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITYILWTWYSF 378
++M L I +R V++G P+V+ D FW P L+L +IH LFQN+FQ+ + W+ Y F
Sbjct: 602 TKMGLRIQDRGEVIKGAPVVEPGDHLFWFNSPNLLLFIIHLVLFQNAFQLAFFSWSTYEF 781
Query: 379 GLENCFRLDYKLAIIKVAFGL 399
+ +CF +I+V+ G+
Sbjct: 782 SINSCFHRTTADNVIRVSVGI 844
Score = 30.4 bits (67), Expect(2) = 1e-16
Identities = 15/36 (41%), Positives = 21/36 (57%)
Frame = +3
Query: 393 IKVAFGLYMLL*HRWVQG*KNQYLMNKHPRR*RNGT 428
+ ++ L+ML HRWVQ * +LM + RNGT
Sbjct: 861 VAMSLCLFML*SHRWVQP*NQPFLMKDWQQHSRNGT 968
>BQ124231 similar to GP|13784981|gb seven transmembrane protein Mlo4 {Zea
mays}, partial (22%)
Length = 566
Score = 53.5 bits (127), Expect(2) = 5e-15
Identities = 24/53 (45%), Positives = 29/53 (54%)
Frame = +1
Query: 143 KGHEPLLSVNGLHQLHILIFFLAVFHVFYSAVTMLLGRLKIRGWKEWEEETLT 195
KGHE S L QLH +F L + HV YS V + L +KI W+ WE E T
Sbjct: 124 KGHESFASYESLEQLHRFVFVLGITHVSYSFVAVALAMIKIYSWRTWENEAKT 282
Score = 45.4 bits (106), Expect(2) = 5e-15
Identities = 25/82 (30%), Positives = 44/82 (53%), Gaps = 5/82 (6%)
Frame = +3
Query: 201 ANDAAKLRLTHETSFVRAHTSF-WTR----VYIGCFFRQFYRSVRKVDYQTLRNGFINVH 255
+ + +RL ++F+ S W+ V++ CF RQF+ S+ + DY LR GFI H
Sbjct: 321 STSTSSIRLKRLSTFIFHQASHPWSHHKILVWLLCFSRQFWSSISRTDYMALRLGFITTH 500
Query: 256 LAPGSKFNFQKYIKRSLEDDFK 277
P ++F Y+ S+++ F+
Sbjct: 501 DLP-LTYDFHNYMLPSMDEGFR 563
>TC77962 similar to PIR|S49848|S49848 probable narbonin - jack bean, partial
(18%)
Length = 1239
Score = 31.2 bits (69), Expect = 1.0
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -3
Query: 149 LSVNGLHQLHILIFFLAVFHVFYSAV 174
++ N H LHILIFFL H F++ V
Sbjct: 361 MASNSYHNLHILIFFLVFKHFFWAEV 284
>TC93199 weakly similar to GP|9755448|gb|AAF98209.1| Unknown protein
{Arabidopsis thaliana}, partial (4%)
Length = 738
Score = 30.0 bits (66), Expect = 2.3
Identities = 17/37 (45%), Positives = 19/37 (50%), Gaps = 3/37 (8%)
Frame = -1
Query: 221 SFWTRVYIGCFFRQFYRSVRKVDYQTLR---NGFINV 254
S W R YI CFFR SV +V LR N IN+
Sbjct: 564 SRWNRAYIRCFFRML*TSVLRVSNSRLREVSNNAINL 454
>TC88302 similar to GP|8953714|dbj|BAA98077.1 contains similarity to GTPase
activating protein~gene_id:F6N7.7 {Arabidopsis
thaliana}, complete
Length = 1198
Score = 29.6 bits (65), Expect = 3.0
Identities = 13/43 (30%), Positives = 26/43 (60%)
Frame = +3
Query: 233 RQFYRSVRKVDYQTLRNGFINVHLAPGSKFNFQKYIKRSLEDD 275
R+F +SV+K +Y+T++N + ++ A +F + K +E D
Sbjct: 213 REFLKSVKKSEYETIKNQWQSISSAQAKRFTKFRERKGLIEKD 341
>CA858513 weakly similar to PIR|S61428|S614 embryonic abundant protein group
3 precursor (clone PM10) - soybean, partial (19%)
Length = 869
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = -2
Query: 350 PQLVLHLIHFALFQNSFQITYIL 372
P L LHLIHF LFQ QI + L
Sbjct: 361 PYLPLHLIHFPLFQPITQILHYL 293
>TC86988 weakly similar to GP|17529294|gb|AAL38874.1 unknown protein
{Arabidopsis thaliana}, partial (66%)
Length = 1378
Score = 28.9 bits (63), Expect = 5.1
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = +2
Query: 445 PWMVAPLLDQQCSPLDQHFTVSRL 468
PW+ PL D+ C+ L Q+F +S L
Sbjct: 803 PWLSLPLKDKTCAKLIQYFELSEL 874
>BQ124242 similar to PIR|S12404|A330 conglutin delta precursor -
narrow-leaved blue lupine, partial (14%)
Length = 391
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = -2
Query: 350 PQLVLHLIHFALFQNSFQITYIL 372
P L LHLIHF LFQ QI + L
Sbjct: 336 PYLPLHLIHFPLFQPITQILHYL 268
>BI312369
Length = 328
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = -3
Query: 350 PQLVLHLIHFALFQNSFQITYIL 372
P L LHLIHF LFQ QI + L
Sbjct: 254 PYLPLHLIHFPLFQPITQILHYL 186
>BQ124315 similar to GP|23510634|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (1%)
Length = 681
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = -1
Query: 350 PQLVLHLIHFALFQNSFQITYIL 372
P L LHLIHF LFQ QI + L
Sbjct: 342 PYLPLHLIHFPLFQPITQILHYL 274
>TC78043 weakly similar to GP|4753801|emb|CAB42052.1 nicotianamine synthase
{Lycopersicon esculentum}, partial (83%)
Length = 1285
Score = 28.9 bits (63), Expect = 5.1
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +1
Query: 96 QNYIVRICISEKIANKMLPCPHKYIGGDKKSN 127
Q YI R+ E++ N +L CP +Y G KKSN
Sbjct: 571 Q*YIGRVK*FERL*NCLLSCPGRYGQGGKKSN 666
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.338 0.147 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,129,803
Number of Sequences: 36976
Number of extensions: 273955
Number of successful extensions: 2470
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 2444
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2462
length of query: 532
length of database: 9,014,727
effective HSP length: 101
effective length of query: 431
effective length of database: 5,280,151
effective search space: 2275745081
effective search space used: 2275745081
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC127019.14


