

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126794.6 - phase: 0
(926 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
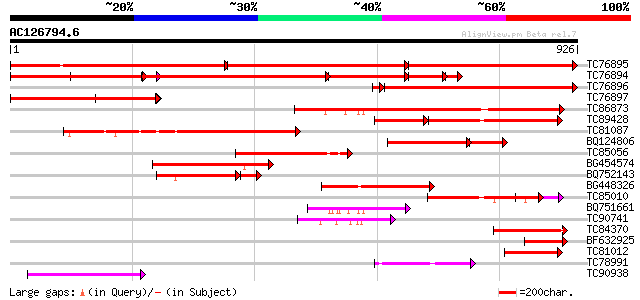
TC76895 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 588 0.0
TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 600 0.0
TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 610 e-180
TC76897 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 424 e-119
TC86873 homologue to GP|530207|gb|AAA66338.1|| heat shock protei... 362 e-100
TC89428 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus l... 225 1e-87
TC81087 homologue to GP|530207|gb|AAA66338.1|| heat shock protei... 304 1e-82
BQ124806 similar to SP|P42762|ERD1 ERD1 protein chloroplast pre... 214 6e-69
TC85056 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus l... 199 5e-51
BG454574 similar to SP|P42762|ERD1 ERD1 protein chloroplast pre... 187 1e-47
BQ752143 similar to PIR|T39572|T39 probable proteinase subunit -... 147 1e-44
BG448326 similar to PIR|T51523|T51 clpB heat shock protein-like ... 160 2e-39
TC85010 weakly similar to PIR|T40514|T40514 Chaperonin hsp78p - ... 145 7e-35
BQ751661 weakly similar to PIR|T39572|T39 probable proteinase su... 104 2e-22
TC90741 similar to PIR|T51523|T51523 clpB heat shock protein-lik... 93 4e-19
TC84370 weakly similar to PIR|T39572|T39572 probable proteinase ... 77 3e-14
BF632925 similar to GP|15425870|gb| heat shock protein 78 {Candi... 77 3e-14
TC81012 similar to PIR|T51523|T51523 clpB heat shock protein-lik... 75 1e-13
TC78991 similar to GP|9759268|dbj|BAB09589.1 101 kDa heat shock ... 71 2e-12
TC90938 similar to GP|21592785|gb|AAM64734.1 unknown {Arabidopsi... 68 2e-11
>TC76895 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], complete
Length = 3425
Score = 588 bits (1517), Expect(3) = 0.0
Identities = 307/359 (85%), Positives = 327/359 (90%)
Frame = +2
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+R LAQSINVPGLV G + + +KG+ +S+RSV+MM +RT R+ +SGLRT+N LD
Sbjct: 230 MARVLAQSINVPGLVVGHKQ-SQHKGSGKSKRSVKMMCASRTIGLRMPGFSGLRTINHLD 406
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
+MLRPG DFH+KV I + RA R R + +AMFERFTEKAIKVIMLAQEEARRLGHN
Sbjct: 407 TMLRPGLDFHAKVSMAISSRRA---RAKRIIPRAMFERFTEKAIKVIMLAQEEARRLGHN 577
Query: 121 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 180
FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR
Sbjct: 578 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 757
Query: 181 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGA 240
VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGE A
Sbjct: 758 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGESA 937
Query: 241 DSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 300
D+V ATVGSGSSNNK PTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP
Sbjct: 938 DNVTATVGSGSSNNKTPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 1117
Query: 301 CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKK 359
CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEER ++
Sbjct: 1118CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERFEE 1294
Score = 533 bits (1372), Expect(3) = 0.0
Identities = 270/297 (90%), Positives = 287/297 (95%)
Frame = +3
Query: 357 LKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY 416
LKKLMEEIKQSD+IILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY
Sbjct: 1287 LKKLMEEIKQSDDIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY 1466
Query: 417 RKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQ 476
RKHIEKDPALERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTD+AL+AAA+LS+Q
Sbjct: 1467 RKHIEKDPALERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDDALIAAAQLSYQ 1646
Query: 477 YISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAG 536
YISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEA+ L+KEVR+IVKEK+E VRNQ+FEKAG
Sbjct: 1647 YISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEAKELDKEVRKIVKEKEEFVRNQDFEKAG 1826
Query: 537 ELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVD 596
ELRDKEMDL+ QISAL+EK KEM+KAESEA D G +VTEVDIQHIV+SWTGIPVDKVS D
Sbjct: 1827 ELRDKEMDLRAQISALVEKGKEMSKAESEAADEGPIVTEVDIQHIVSSWTGIPVDKVSAD 2006
Query: 597 ESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGK 653
ESDRLLKMEDTLHKR+IGQ EAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVG+
Sbjct: 2007 ESDRLLKMEDTLHKRVIGQDEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGE 2177
Score = 511 bits (1315), Expect(3) = 0.0
Identities = 259/275 (94%), Positives = 269/275 (97%)
Frame = +1
Query: 652 GKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR 711
GKSELAKALA+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR
Sbjct: 2173 GKSELAKALAAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR 2352
Query: 712 PYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGG 771
PYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGG
Sbjct: 2353 PYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGG 2532
Query: 772 RRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIM 831
RRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIM
Sbjct: 2533 RRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIM 2712
Query: 832 LKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREI 891
L+EVF+RLK KEIEL VTERFR+RVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREI
Sbjct: 2713 LREVFQRLKNKEIELQVTERFRDRVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREI 2892
Query: 892 KEGDSVIVDADSDGNVIVLNGSTGAPDSLPDALPV 926
KEGDSVIVD DS+GNVIVLNG++G +SLP+AL +
Sbjct: 2893 KEGDSVIVDVDSEGNVIVLNGTSGTQESLPEALAI 2997
>TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (71%)
Length = 2413
Score = 600 bits (1548), Expect(4) = 0.0
Identities = 307/308 (99%), Positives = 308/308 (99%)
Frame = +2
Query: 216 RVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKL 275
RVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKL
Sbjct: 839 RVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKL 1018
Query: 276 DPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKV 335
DPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKV
Sbjct: 1019 DPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKV 1198
Query: 336 ITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANI 395
ITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANI
Sbjct: 1199 ITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANI 1378
Query: 396 LKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYE 455
LKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYE
Sbjct: 1379 LKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYE 1558
Query: 456 IHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKE 515
IHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKE
Sbjct: 1559 IHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKE 1738
Query: 516 VRQIVKEK 523
VRQIVKE+
Sbjct: 1739 VRQIVKEE 1762
Score = 413 bits (1061), Expect = e-115
Identities = 216/225 (96%), Positives = 216/225 (96%)
Frame = +1
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD
Sbjct: 193 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 372
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN
Sbjct: 373 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 552
Query: 121 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 180
FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR
Sbjct: 553 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 732
Query: 181 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADP 225
VLELSLEEARQLGHNYIGSEH LLGLLREGEGVAA G P
Sbjct: 733 VLELSLEEARQLGHNYIGSEHSLLGLLREGEGVAAPCS*KFGCRP 867
Score = 255 bits (652), Expect(4) = 0.0
Identities = 130/133 (97%), Positives = 132/133 (98%)
Frame = +3
Query: 521 KEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQH 580
+ KDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQH
Sbjct: 1755 RRKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQH 1934
Query: 581 IVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPI 640
IVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPI
Sbjct: 1935 IVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPI 2114
Query: 641 ASFIFSGPTGVGK 653
ASFIFSGPTGVG+
Sbjct: 2115 ASFIFSGPTGVGE 2153
Score = 129 bits (323), Expect(4) = 0.0
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +1
Query: 652 GKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR 711
GKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR
Sbjct: 2149 GKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRR 2328
Query: 712 PYT 714
PYT
Sbjct: 2329 PYT 2337
Score = 78.2 bits (191), Expect = 1e-14
Identities = 48/151 (31%), Positives = 88/151 (57%), Gaps = 3/151 (1%)
Frame = +1
Query: 100 TEKAIKVIMLAQEEARRLG-HNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVE 158
+ ++++++ + + RL ++ + T L ++ G +KVL +G N
Sbjct: 280 SRRSVRMMFTTRTASPRLSSYSGLRTLNSLDSMLRPGQDFHSKVLTQIGTNRAKG----- 444
Query: 159 KIIGRGSGFV--AVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAAR 216
GRGS V A+ FT +A +V+ L+ EEAR+LGHN++G+E +LLGL+ EG G+AA+
Sbjct: 445 ---GRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAK 615
Query: 217 VLENLGADPTNIRTQVIRMVGEGADSVGATV 247
VL+++G + + R +V +++G G+ V +
Sbjct: 616 VLKSMGINLKDARVEVEKIIGRGSGFVAVEI 708
Score = 53.1 bits (126), Expect(4) = 0.0
Identities = 25/28 (89%), Positives = 25/28 (89%)
Frame = +2
Query: 712 PYTVVLFDEIEKAHPDVFNMMLQILEDG 739
P VVLFDEI KAHPDVFNMMLQILEDG
Sbjct: 2330 PTRVVLFDEIXKAHPDVFNMMLQILEDG 2413
>TC76896 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (35%)
Length = 1348
Score = 610 bits (1574), Expect(2) = e-180
Identities = 313/315 (99%), Positives = 314/315 (99%)
Frame = +1
Query: 612 IIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAM 671
+IGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAK LASYYFGSEEAM
Sbjct: 61 VIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKTLASYYFGSEEAM 240
Query: 672 IRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNM 731
IRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNM
Sbjct: 241 IRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNM 420
Query: 732 MLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIK 791
MLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIK
Sbjct: 421 MLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIK 600
Query: 792 SLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTER 851
SLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTER
Sbjct: 601 SLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTER 780
Query: 852 FRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN 911
FRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN
Sbjct: 781 FRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLN 960
Query: 912 GSTGAPDSLPDALPV 926
GSTGAPDSLPDALPV
Sbjct: 961 GSTGAPDSLPDALPV 1005
Score = 39.3 bits (90), Expect(2) = e-180
Identities = 18/19 (94%), Positives = 19/19 (99%)
Frame = +3
Query: 593 VSVDESDRLLKMEDTLHKR 611
VSVDESDRLLKME+TLHKR
Sbjct: 3 VSVDESDRLLKMEETLHKR 59
>TC76897 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (18%)
Length = 826
Score = 424 bits (1091), Expect = e-119
Identities = 221/246 (89%), Positives = 230/246 (92%), Gaps = 1/246 (0%)
Frame = +2
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSR-RSVRMMFTTRTASPRLSSYSGLRTLNSL 59
MSR LAQSINVPGLVAGRRHVNNN G+ RSR RSVRMM+ T+TA+ RLS +SGLR LNSL
Sbjct: 89 MSRVLAQSINVPGLVAGRRHVNNNNGSVRSRPRSVRMMYATQTATMRLSGFSGLRPLNSL 268
Query: 60 DSMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGH 119
DSM RP QDFHSKVLTQIGTNRA+GGRGSRCVT+AMFERFTEKAIKVIMLAQEEARRLGH
Sbjct: 269 DSMFRPRQDFHSKVLTQIGTNRARGGRGSRCVTRAMFERFTEKAIKVIMLAQEEARRLGH 448
Query: 120 NFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAK 179
NFVGTEQILLGLIGEGTGI AKVLKSMGINLKD+RVEVEKIIGRGSGFVAVEIPFTPRAK
Sbjct: 449 NFVGTEQILLGLIGEGTGIGAKVLKSMGINLKDSRVEVEKIIGRGSGFVAVEIPFTPRAK 628
Query: 180 RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEG 239
RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNI T VI M GEG
Sbjct: 629 RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIXTXVIXMXGEG 808
Query: 240 ADSVGA 245
AD+ GA
Sbjct: 809 ADNXGA 826
Score = 73.2 bits (178), Expect = 4e-13
Identities = 42/110 (38%), Positives = 68/110 (61%), Gaps = 2/110 (1%)
Frame = +2
Query: 140 AKVLKSMGINLKDARVEVEKIIGRGSGFV--AVEIPFTPRAKRVLELSLEEARQLGHNYI 197
+KVL +G N GRGS V A+ FT +A +V+ L+ EEAR+LGHN++
Sbjct: 302 SKVLTQIGTNRARG--------GRGSRCVTRAMFERFTEKAIKVIMLAQEEARRLGHNFV 457
Query: 198 GSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGADSVGATV 247
G+E +LLGL+ EG G+ A+VL+++G + + R +V +++G G+ V +
Sbjct: 458 GTEQILLGLIGEGTGIGAKVLKSMGINLKDSRVEVEKIIGRGSGFVAVEI 607
>TC86873 homologue to GP|530207|gb|AAA66338.1|| heat shock protein {Glycine
max}, partial (53%)
Length = 1475
Score = 362 bits (929), Expect = e-100
Identities = 205/499 (41%), Positives = 302/499 (60%), Gaps = 58/499 (11%)
Frame = +2
Query: 466 ALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEK-------EVRQ 518
A+V AA+LS +YI+ R LPDKAIDL+DEA + VR+Q PEE LE+ E+
Sbjct: 2 AIVVAAQLSSRYITGRHLPDKAIDLVDEACANVRVQLDSQPEEIDNLERKRMQLEVELHA 181
Query: 519 IVKEKDEAVRNQEFE---KAGELRDKEMDLKT----------QISALIEKNKEMNKAESE 565
+ KEKD+A + + + + +LRDK LK +I L +K +E+ A E
Sbjct: 182 LEKEKDKASKARLVDVRRELDDLRDKLQPLKMKYSKEKERIDEIRRLKQKREELLFALQE 361
Query: 566 AG----------------------------------DVGALVTEV----DIQHIVASWTG 587
A D ++TE I +V+ WTG
Sbjct: 362 AERRYDLARAADLRYGAIEEVETAIKNLEGSTDGNTDENLMLTETVGPDQIAEVVSRWTG 541
Query: 588 IPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSG 647
IPV ++ +E RL+ + D LH R++GQ +AV A++ A+ R+R GL P +P SF+F G
Sbjct: 542 IPVTRLGQNEKARLVGLGDRLHTRVVGQDQAVNAVAEAVLRSRAGLGRPQQPTGSFLFLG 721
Query: 648 PTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEA 707
PTGVGK+ELAKALA F E ++R+DMSE+ME+H+VS+LIG+PPGYVG+ EGGQLTEA
Sbjct: 722 PTGVGKTELAKALAEQLFDDENQLVRIDMSEYMEQHSVSRLIGAPPGYVGHEEGGQLTEA 901
Query: 708 VRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVI 767
VRRRPY+VVLFDE+EKAH VFN +LQ+L+DGRLTD +GRTVDF+NT++IMTSN+G+ +
Sbjct: 902 VRRRPYSVVLFDEVEKAHTSVFNTLLQVLDDGRLTDGQGRTVDFRNTVIIMTSNLGAEHL 1081
Query: 768 EKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEI 827
G + + V +E++++FRPE LNRLDE++VF L+ +++++
Sbjct: 1082LSG----------LSGKCTMQAARDRVMQEVRRHFRPELLNRLDEVVVFDPLSHEQLRKV 1231
Query: 828 ADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKML 887
A + +K+V RL + I L+VT+ + ++ E Y+P YGARP+R+ + + + ++ ++
Sbjct: 1232ARLQMKDVASRLAERGIALAVTDAALDYILAESYDPVYGARPIRKWLEKKVVTELSRMLI 1411
Query: 888 AREIKEGDSVIVDADSDGN 906
EI E + +DA G+
Sbjct: 1412REEIDENTTGYIDAGPKGS 1468
>TC89428 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus lunatus},
partial (32%)
Length = 1227
Score = 225 bits (574), Expect(2) = 1e-87
Identities = 115/218 (52%), Positives = 160/218 (72%)
Frame = +2
Query: 685 VSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDS 744
VS+L+G+PPGYVGY EGGQLTE VRRRPY+VVLFDEIEKAH DVFN++LQ+L+DGR+TDS
Sbjct: 317 VSRLVGAPPGYVGYEEGGQLTEVVRRRPYSVVLFDEIEKAHHDVFNILLQLLDDGRITDS 496
Query: 745 KGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRP 804
+GRTV F N +LIMTSN+GS I + D+K + Y+++K V E +Q FRP
Sbjct: 497 QGRTVSFTNCVLIMTSNIGSHHILE-----TLSSTQDDKIAVYDQMKRQVVELARQTFRP 661
Query: 805 EFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPS 864
EF+NR+DE IVF+ L E+ +I ++ ++ V RLK K+I+L TE + + G++P+
Sbjct: 662 EFMNRIDEYIVFQPLDSSEISKIVELQMERVKGRLKQKKIDLHYTEEAVKLLGVLGFDPN 841
Query: 865 YGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 902
+GARP++R I +L+E+ +A +L + KE DS+IVDAD
Sbjct: 842 FGARPVKRVIQQLVENEIAMGVLRGDFKEEDSIIVDAD 955
Score = 117 bits (293), Expect(2) = 1e-87
Identities = 53/87 (60%), Positives = 73/87 (82%)
Frame = +1
Query: 597 ESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSEL 656
E ++L+ +E LHKR+IGQ AV++++ AIRR+R GL +PNRPIASF+F GPTGVGK+EL
Sbjct: 52 EREKLVFLEQVLHKRVIGQDIAVKSVADAIRRSRAGLSDPNRPIASFMFMGPTGVGKTEL 231
Query: 657 AKALASYYFGSEEAMIRLDMSEFMERH 683
KALA+Y F +E A++R+DMSE+ME+H
Sbjct: 232 GKALANYLFNTENALVRIDMSEYMEKH 312
>TC81087 homologue to GP|530207|gb|AAA66338.1|| heat shock protein {Glycine
max}, partial (41%)
Length = 1318
Score = 304 bits (778), Expect = 1e-82
Identities = 171/398 (42%), Positives = 251/398 (62%), Gaps = 11/398 (2%)
Frame = +2
Query: 88 SRCVTKAMF----ERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVL 143
S + K+ F E+FT K + + A E A GH + + L+ + GI + +
Sbjct: 152 SHSINKSSFTMNPEKFTHKTNEALAGAHELAMTSGHAQITPLHLASILVSDPNGIFFQAI 331
Query: 144 KSMGINLKDARVEVEKIIGRGSGFVAV------EIPFTPRAKRVLELSLEEARQLGHNYI 197
++ + AR VE+++ + + E+P + + + + + G ++
Sbjct: 332 SNVA-GEESARA-VERVLKQALKKLPSQSPPPDEVPGSTALIKAIRRAQAAQKSRGDTHL 505
Query: 198 GSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMP 257
+ L+LG+L + + + + G + ++T+V ++ G+ G V S S +
Sbjct: 506 AVDQLILGILEDSQ--IGDLFKEAGVAVSRVKTEVEKLRGKD----GKKVESASGDTNFQ 667
Query: 258 TLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGL 317
L+ YG +L + A GKLDPV+GR +I RV +IL RRTKNNP LIGEPGVGKTA+ EGL
Sbjct: 668 ALKTYGRDLVEQA--GKLDPVIGRDEEIRRVVRILSRRTKNNPVLIGEPGVGKTAVVEGL 841
Query: 318 AQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDE 376
AQRI GDVP + ++I LDMG LVAG KYRGEFEERLK +++E+++++ ++ILFIDE
Sbjct: 842 AQRIVRGDVPSNLADVRLIALDMGALVAGAKYRGEFEERLKAVLKEVEEAEGKVILFIDE 1021
Query: 377 VHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVP 436
+H ++GAG EG++DAAN+ KP LARG+L+CIGATTL+EYRK++EKD A ERRFQ V V
Sbjct: 1022IHLVLGAGRTEGSMDAANLFKPMLARGQLRCIGATTLEEYRKYVEKDAAFERRFQQVYVA 1201
Query: 437 EPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELS 474
EP+VP+TI IL+GL+ERYE HH +R D A+V AA+LS
Sbjct: 1202EPSVPDTISILRGLKERYEGHHGVRIQDRAIVVAAQLS 1315
>BQ124806 similar to SP|P42762|ERD1 ERD1 protein chloroplast precursor.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (20%)
Length = 679
Score = 214 bits (546), Expect(2) = 6e-69
Identities = 101/136 (74%), Positives = 123/136 (90%)
Frame = +2
Query: 618 AVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMS 677
AV AISR ++R+RVGL++P RPIA+ +F GPTGVGK+ELAK+LA+ YFGSE MIRLDMS
Sbjct: 2 AVSAISRFVKRSRVGLQDPGRPIATLLFCGPTGVGKTELAKSLAACYFGSETNMIRLDMS 181
Query: 678 EFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILE 737
E+MERH+VSKL+GSPPGYVGY EGG LTEA+RR+P+TVVLFDEIEKAHPD+FN++LQ++E
Sbjct: 182 EYMERHSVSKLLGSPPGYVGYGEGGILTEAIRRKPFTVVLFDEIEKAHPDIFNILLQLME 361
Query: 738 DGRLTDSKGRTVDFKN 753
DG LTDS+GR V FKN
Sbjct: 362 DGHLTDSQGRRVSFKN 409
Score = 65.9 bits (159), Expect(2) = 6e-69
Identities = 34/61 (55%), Positives = 43/61 (69%)
Frame = +3
Query: 752 KNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLD 811
K L++MTSNVGSS I KG L D+K +SY+ +KS+V EEL+ YFRPE LNR+D
Sbjct: 405 KMHLVVMTSNVGSSAIAKGRHNSMGFLISDDKPTSYSGLKSMVIEELRTYFRPELLNRID 584
Query: 812 E 812
E
Sbjct: 585 E 587
>TC85056 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus lunatus},
partial (25%)
Length = 743
Score = 199 bits (505), Expect = 5e-51
Identities = 99/191 (51%), Positives = 144/191 (74%)
Frame = +1
Query: 369 EIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALER 428
+IILFIDE+HT++GAGA GA+DA N+LKP L RGEL+CIGATTL+EYRK+IEKDPALER
Sbjct: 19 QIILFIDEIHTVVGAGATSGAMDAGNLLKPMLGRGELRCIGATTLNEYRKYIEKDPALER 198
Query: 429 RFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAI 488
RFQ V +P+V +TI IL+GLRERYE+HH ++ +D ALV+AA L+ +YI++RFLPDKAI
Sbjct: 199 RFQQVFCCQPSVEDTISILRGLRERYELHHGVKISDSALVSAAVLADRYITERFLPDKAI 378
Query: 489 DLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQ 548
DL+DEA ++++++ P E +++ V ++ E ++ + +KA +++ L+
Sbjct: 379 DLVDEAAAKLKMEITSKPTELDEIDRAVLKL--EMEKLSLKSDTDKAS--KERLSKLEND 546
Query: 549 ISALIEKNKEM 559
+S L +K KE+
Sbjct: 547 LSLLKQKQKEL 579
>BG454574 similar to SP|P42762|ERD1 ERD1 protein chloroplast precursor.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (19%)
Length = 672
Score = 187 bits (476), Expect = 1e-47
Identities = 95/202 (47%), Positives = 136/202 (67%), Gaps = 5/202 (2%)
Frame = +1
Query: 234 RMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILG 293
+ + + G+ + +K L ++ +LT A G +DPV+GR+ +++R+ QIL
Sbjct: 67 KSIPQRRSGAGSAAKTKDKKDKKNALSQFCVDLTARASVGLIDPVIGREVEVQRIIQILC 246
Query: 294 RRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEF 353
R+TK+NP L+GE GVGKTAIAEGLA I+ +V + K+V++LD+GLL+AG K RGE
Sbjct: 247 RKTKSNPILLGEAGVGKTAIAEGLAILISRAEVAPFLLTKRVMSLDVGLLMAGAKERGEL 426
Query: 354 EERLKKLMEEIKQSDEIILFIDEVHTLI-----GAGAAEGAIDAANILKPALARGELQCI 408
E+R+ KL+++I +S ++ILFIDEVHTL+ G G D AN+LKP+L RG+ QCI
Sbjct: 427 EDRVTKLIKDIIESGDVILFIDEVHTLVQSGTTGRGNKGSGFDIANLLKPSLGRGQFQCI 606
Query: 409 GATTLDEYRKHIEKDPALERRF 430
+TT+DEYR H EKD AL RF
Sbjct: 607 ASTTIDEYRLHFEKDKALAXRF 672
>BQ752143 similar to PIR|T39572|T39 probable proteinase subunit - fission
yeast (Schizosaccharomyces pombe), partial (18%)
Length = 774
Score = 147 bits (371), Expect(2) = 1e-44
Identities = 77/144 (53%), Positives = 103/144 (71%), Gaps = 8/144 (5%)
Frame = +2
Query: 241 DSVGATVGSGSSNNKMPTLEEYGTNLTK-------LAEEGKLDPVVGRQPQIERVTQILG 293
D+V G+ ++K EE NL K LA + K+DPV+GR+ +I RV +IL
Sbjct: 236 DAVSQIRGTKRVDSKNADTEEEHENLAKFTIDMTALARDKKIDPVIGREEEIRRVVRILS 415
Query: 294 RRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEF 353
RRTKNNP LIGEPGVGKT + EGLAQRI N DVP+ ++ K+++LD+G LVAG+KYRGEF
Sbjct: 416 RRTKNNPVLIGEPGVGKTTVVEGLAQRIVNRDVPDNLKACKLLSLDVGALVAGSKYRGEF 595
Query: 354 EERLKKLMEEIKQSDE-IILFIDE 376
EER+K +++EI+ S E I+LF+DE
Sbjct: 596 EERMKGVLKEIEDSKEMIVLFVDE 667
Score = 52.0 bits (123), Expect(2) = 1e-44
Identities = 24/36 (66%), Positives = 31/36 (85%), Gaps = 1/36 (2%)
Frame = +1
Query: 377 VHTLIGAGAA-EGAIDAANILKPALARGELQCIGAT 411
+H L+GAG++ EG +DAAN+LKP LARG+L CIGAT
Sbjct: 667 IHLLMGAGSSGEGGMDAANLLKPMLARGQLHCIGAT 774
>BG448326 similar to PIR|T51523|T51 clpB heat shock protein-like -
Arabidopsis thaliana, partial (22%)
Length = 646
Score = 160 bits (405), Expect = 2e-39
Identities = 85/186 (45%), Positives = 128/186 (68%), Gaps = 2/186 (1%)
Frame = +2
Query: 510 RGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMD-LKTQI-SALIEKNKEMNKAESEAG 567
+ +++E+ ++ E +A R + +A EL+ ++ L+ Q+ SA E ++ MN +S
Sbjct: 98 QSIKEEIDRVNLEIQQAEREYDLNRAAELKYGSLNSLQRQLESAEKELDEYMNSGKSMLR 277
Query: 568 DVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIR 627
+ VT DI IV+ WTGIPV K+ E ++LL +E+ LHKR++GQ AV+A++ AI+
Sbjct: 278 EE---VTGSDIAEIVSKWTGIPVSKLQQSEREKLLYLEEVLHKRVVGQDPAVKAVAEAIQ 448
Query: 628 RARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSK 687
R+R GL +P+RPIASF+F GPTGVGK+ELA LASY F +E A++ +DMSE ME+H S+
Sbjct: 449 RSRAGLSDPHRPIASFMFMGPTGVGKTELAXTLASYMFNTEXALVXIDMSESMEKHAXSR 628
Query: 688 LIGSPP 693
LIG+PP
Sbjct: 629 LIGAPP 646
>TC85010 weakly similar to PIR|T40514|T40514 Chaperonin hsp78p - fission
yeast (Schizosaccharomyces pombe), partial (23%)
Length = 904
Score = 145 bits (366), Expect = 7e-35
Identities = 78/203 (38%), Positives = 124/203 (60%), Gaps = 13/203 (6%)
Frame = +3
Query: 683 HTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLT 742
HT+S+LIG+P GYVGY + GQLTEAVRR+PY V+LFDE EKAH D+ ++LQ+L++G LT
Sbjct: 9 HTISRLIGAPSGYVGYEDAGQLTEAVRRKPYAVLLFDEFEKAHRDISALLLQVLDEGYLT 188
Query: 743 DSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRI----KSLVTEEL 798
D++G VDF+NT++++TSN+G+ ++ +G D + K+++ I + V + +
Sbjct: 189 DAQGHKVDFRNTIIVLTSNLGADIL------VGADPLHPYKETADGEIDPDVRKAVMDVV 350
Query: 799 KQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKT---------KEIELSVT 849
+ PEFLNR+D IVF++L +++I DI + V + + L+
Sbjct: 351 AANYAPEFLNRIDSFIVFKRLALEALRDIVDISSRRVADSPRR*THLP*RT*SRS*LACG 530
Query: 850 ERFRERVVDEGYNPSYGARPLRR 872
R R +V E P++ R +R
Sbjct: 531 ARLRPQVRSEAAKPAHHQRDWKR 599
Score = 53.1 bits (126), Expect = 5e-07
Identities = 26/78 (33%), Positives = 43/78 (54%)
Frame = +1
Query: 827 IADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKM 886
++ L E+ RL K I L V R+ + + GY+P +GARPL R I + + +A+K+
Sbjct: 436 LSTFRLGELQTRLDDKRISLDVPNHVRDWLAERGYDPKFGARPLNRLITNEIGNGLADKI 615
Query: 887 LAREIKEGDSVIVDADSD 904
+ E+K G++ V D
Sbjct: 616 IRGELKMGETATVAIKED 669
>BQ751661 weakly similar to PIR|T39572|T39 probable proteinase subunit -
fission yeast (Schizosaccharomyces pombe), partial (22%)
Length = 708
Score = 104 bits (259), Expect = 2e-22
Identities = 82/234 (35%), Positives = 115/234 (49%), Gaps = 66/234 (28%)
Frame = +2
Query: 487 AIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIV-------KEKDEA-----------VR 528
AIDL+DEA + VR+ PE LE+++RQI+ KEKDEA +
Sbjct: 5 AIDLVDEAAAAVRVARESQPEIIDSLERKLRQIMIEIAALEKEKDEASQTRLVQARKDAK 184
Query: 529 NQEFE------------KAGE------LRDKEMDLKTQISA------------------- 551
N E E K GE L+ E++ + + +A
Sbjct: 185 NVEEELEPLRQKYLSEIKRGEQIHLAKLKLDELEKRLEDAANAGDHAKAADLQYGAIPEQ 364
Query: 552 -LIEKNKEMNKAESEAG------DVGALVTEV----DIQHIVASWTGIPVDKVSVDESDR 600
+ K E KA ++A D GA+VT+V I IV+ WTGIPV ++ E ++
Sbjct: 365 RAVIKELEAKKAAADAALNATGQDHGAMVTDVVTADHINEIVSRWTGIPVTRLKTTEKEK 544
Query: 601 LLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKS 654
L+ ME L K ++GQ EAV+A+S AIR R GL NPN+P SF+F GP+G GK+
Sbjct: 545 LIHMEKQLGKIVVGQREAVQAVSNAIRLQRSGLSNPNQP-PSFLFCGPSGTGKT 703
>TC90741 similar to PIR|T51523|T51523 clpB heat shock protein-like -
Arabidopsis thaliana, partial (24%)
Length = 710
Score = 93.2 bits (230), Expect = 4e-19
Identities = 69/219 (31%), Positives = 110/219 (49%), Gaps = 59/219 (26%)
Frame = +1
Query: 471 AELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLP-------EEARGLEKEVRQIVKEK 523
A LS +YIS RFLPDKAIDL+DEA ++++++ P LE E ++ +
Sbjct: 52 AILSDRYISGRFLPDKAIDLVDEAAAKLKMEITSKPTALDEINRSVLKLEMERLSLMNDT 231
Query: 524 DEAVRNQEF----------EKAGELRDK---EMDLKTQISALIEK----NKEMNKAESEA 566
D+A +++ EK GEL ++ E + T++ ++ E+ N E+ +AE E
Sbjct: 232 DKASKDRLNRLETELSLLKEKQGELTEQWEHEKSVMTRLQSIKEEIDRVNLEIQQAEREY 411
Query: 567 ----------GDVGAL-------------------------VTEVDIQHIVASWTGIPVD 591
G + +L VT DI IV+ WTGIP+
Sbjct: 412 DLNRAAELKYGSLNSLQRQLEGAEKELHEYMSSGKSMLREEVTGNDIGEIVSKWTGIPIS 591
Query: 592 KVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRAR 630
K+ E ++LL ++D LHKR++GQ AV+A++ AI+R+R
Sbjct: 592 KLQQSEREKLLYLQDELHKRVVGQDPAVKAVAEAIQRSR 708
>TC84370 weakly similar to PIR|T39572|T39572 probable proteinase subunit -
fission yeast (Schizosaccharomyces pombe), partial (13%)
Length = 643
Score = 77.0 bits (188), Expect = 3e-14
Identities = 40/122 (32%), Positives = 78/122 (63%), Gaps = 2/122 (1%)
Frame = +3
Query: 791 KSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLK--TKEIELSV 848
+ LV L+ PEFLNR++ ++F +LT+ E+++I DI L E+ +RL+ +++ + V
Sbjct: 12 RELVMNALRNCSLPEFLNRINSTVIFNRLTRREIRKIVDIRLLEIQKRLEGNGRKVYIDV 191
Query: 849 TERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVI 908
+E ++ + + GY+P+YGARPL R I + + + +A +L +++G++ V+ +G +
Sbjct: 192 SEEAKDYLGNAGYSPAYGARPLSRLIEKEVLNKLAILILRGNVRDGENARVEL-INGRIT 368
Query: 909 VL 910
VL
Sbjct: 369 VL 374
>BF632925 similar to GP|15425870|gb| heat shock protein 78 {Candida
albicans}, partial (2%)
Length = 525
Score = 77.0 bits (188), Expect = 3e-14
Identities = 33/70 (47%), Positives = 53/70 (75%)
Frame = -3
Query: 842 KEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDA 901
+EI+ V+E ++ V EGY+P+YGA PLR+AI+ L+ + +AE +LA + KEGD+V +D
Sbjct: 520 EEIDFEVSESVKDLVCKEGYDPTYGASPLRKAIVNLIANPLAEALLAEKCKEGDTVFIDL 341
Query: 902 DSDGNVIVLN 911
D++GN +V+N
Sbjct: 340 DANGNTLVIN 311
>TC81012 similar to PIR|T51523|T51523 clpB heat shock protein-like -
Arabidopsis thaliana, partial (11%)
Length = 577
Score = 75.1 bits (183), Expect = 1e-13
Identities = 35/95 (36%), Positives = 64/95 (66%)
Frame = +3
Query: 808 NRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGA 867
NR+DE IVFR L + ++ I + L+ V +R+ +++++ VTE + + GY+PSYGA
Sbjct: 3 NRVDEYIVFRPLDRDQISSIVRLQLERVQKRVADRKMKIRVTEPAIQLLGSLGYDPSYGA 182
Query: 868 RPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 902
RP++R I + +E+ +A+ +L E KE D++++D +
Sbjct: 183 RPVKRVIQQNVENELAKGILRGEFKEEDTILIDTE 287
>TC78991 similar to GP|9759268|dbj|BAB09589.1 101 kDa heat shock protein;
HSP101-like protein {Arabidopsis thaliana}, partial
(13%)
Length = 1370
Score = 71.2 bits (173), Expect = 2e-12
Identities = 47/167 (28%), Positives = 85/167 (50%), Gaps = 1/167 (0%)
Frame = +2
Query: 596 DESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGL-KNPNRPIASFIFSGPTGVGKS 654
D RLLK TL +++ Q +A AI+ A+ + ++G K ++ +F+GP +GK
Sbjct: 320 DSFKRLLK---TLTEKVWWQQDAASAIATAVTQCKLGNGKRRSKGDTWLMFTGPDRIGKK 490
Query: 655 ELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYT 714
+A AL+ GS +I L + + G T ++ E +RR P++
Sbjct: 491 RMAAALSELVSGSNPIVISLAQRRGDGDSNAHQ-------FRGKTVLDRIVETIRRNPHS 649
Query: 715 VVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSN 761
V++ ++I++A+ + + + +E GR DS GR + N + I+TSN
Sbjct: 650 VIMLEDIDEANTLLRGNIKRAMEQGRFPDSHGREISLGNVMFILTSN 790
>TC90938 similar to GP|21592785|gb|AAM64734.1 unknown {Arabidopsis
thaliana}, partial (53%)
Length = 691
Score = 67.8 bits (164), Expect = 2e-11
Identities = 51/199 (25%), Positives = 96/199 (47%), Gaps = 5/199 (2%)
Frame = +2
Query: 29 RSRRSVRMMFTTRTASPRLSSYSG--LRTLNSLDSMLRPGQDFHSKVLTQIGTNRAKGGR 86
RS S+R++ +P +S+ G +R + S+ FH + L +
Sbjct: 71 RSNPSLRLIQNLNITNPN-NSWLGTKIRVQHCSVSVKSSVSVFHRRPLAATVSFSLPTSN 247
Query: 87 GSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSM 146
R +++ KAIK ++ EAR+L GTE +++G++ EGT +A+K L++
Sbjct: 248 PERVSPGQEIPKWSSKAIKSFAMSAIEARKLKFTTTGTEALIMGILIEGTNLASKFLRAN 427
Query: 147 GINLKDARVEVEKIIGRGSGFVAV--EIPFTPRAKRVLELSLE-EARQLGHNYIGSEHLL 203
GI L R E+ KI+G F P T A+R L+ +++ + + + + + H++
Sbjct: 428 GITLFKVRDEIVKILGEVDPFTETPERPPMTDDAQRALDWAVDRKLKSVDGGEVTTAHII 607
Query: 204 LGLLREGEGVAARVLENLG 222
LG+ + + ++L LG
Sbjct: 608 LGIWSDVDSPGHKILSKLG 664
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.136 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,141,243
Number of Sequences: 36976
Number of extensions: 206009
Number of successful extensions: 1229
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 1188
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1211
length of query: 926
length of database: 9,014,727
effective HSP length: 105
effective length of query: 821
effective length of database: 5,132,247
effective search space: 4213574787
effective search space used: 4213574787
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126794.6


