

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126792.7 - phase: 0
(253 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
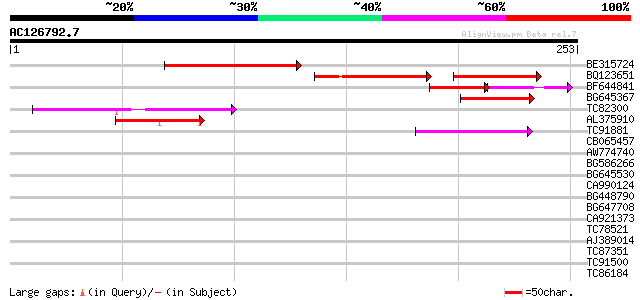
Score E
Sequences producing significant alignments: (bits) Value
BE315724 115 2e-26
BQ123651 49 7e-15
BF644841 similar to GP|18175662|gb| putative receptor protein ki... 44 6e-08
BG645367 53 1e-07
TC82300 52 3e-07
AL375910 45 2e-05
TC91881 42 3e-04
CB065457 30 0.016
AW774740 34 0.062
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 33 0.081
BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinas... 30 1.2
CA990124 weakly similar to PIR|A96697|A96 protein F1N21.18 [impo... 28 2.6
BG448790 28 2.6
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 28 3.4
CA921373 weakly similar to PIR|T14519|T14 probable S-receptor ki... 28 4.4
TC78521 PIR|A25642|HSZM4 histone H4 - maize, complete 27 5.8
AJ389014 PIR|A25642|HS histone H4 - maize, partial (98%) 27 5.8
TC87351 similar to GP|10176988|dbj|BAB10220. GTP-binding protein... 27 7.5
TC91500 similar to SP|Q9BBS8|RPOC_LOTJA DNA-directed RNA polymer... 27 7.5
TC86184 similar to GP|21553481|gb|AAM62574.1 putative ribosomal ... 27 9.9
>BE315724
Length = 392
Score = 115 bits (287), Expect = 2e-26
Identities = 56/61 (91%), Positives = 59/61 (95%)
Frame = -2
Query: 70 DVVLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHII 129
+VVLVG+SREEVNGRLETWRQALEAY F LSRSKTEYMECNFSGRRS+STLEVKVGDHII
Sbjct: 184 NVVLVGESREEVNGRLETWRQALEAYEFRLSRSKTEYMECNFSGRRSRSTLEVKVGDHII 5
Query: 130 P 130
P
Sbjct: 4 P 2
>BQ123651
Length = 654
Score = 48.5 bits (114), Expect(2) = 7e-15
Identities = 22/39 (56%), Positives = 26/39 (66%)
Frame = -2
Query: 199 WAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRNDIIRER 237
WA HE ++S A+ RMLRWM G TR D+IRND I R
Sbjct: 248 WAE*CHHERRISEAKKRMLRWMCGNTRRDKIRNDSISVR 132
Score = 48.5 bits (114), Expect(2) = 7e-15
Identities = 27/52 (51%), Positives = 34/52 (64%)
Frame = -3
Query: 137 YLESFVQNDGEIEADVSHRIQVGWLKWRRASCVLCDKKVPFKLKGKFYRTAV 188
YL S +Q + + DV++RIQ +K R VLCDKKVP K+KGKFY T V
Sbjct: 421 YLGSVIQK*QK-QRDVNNRIQARLMK*RSPLGVLCDKKVPLKIKGKFYLTTV 269
>BF644841 similar to GP|18175662|gb| putative receptor protein kinase
{Arabidopsis thaliana}, partial (1%)
Length = 676
Score = 43.9 bits (102), Expect(2) = 6e-08
Identities = 18/27 (66%), Positives = 22/27 (80%)
Frame = +2
Query: 188 VRSALLYGTECWAVKSQHENQVSVAEM 214
+R A+LY ECWAVK+QHEN+VS EM
Sbjct: 158 LRPAMLYAMECWAVKNQHENKVSYVEM 238
Score = 29.6 bits (65), Expect(2) = 6e-08
Identities = 18/38 (47%), Positives = 22/38 (57%)
Frame = +3
Query: 214 MRMLRWMSGKTRHDRIRNDIIRERERESGGSTYSRKVG 251
M+ML W+ GKTR +N+ I RES G SRK G
Sbjct: 243 MKMLCWICGKTRRYPFQNNNI----RESLGIVNSRKYG 344
>BG645367
Length = 739
Score = 52.8 bits (125), Expect = 1e-07
Identities = 21/33 (63%), Positives = 28/33 (84%)
Frame = +1
Query: 202 KSQHENQVSVAEMRMLRWMSGKTRHDRIRNDII 234
K+QH+N++S +EMRML WM GKTRH RIRN+ +
Sbjct: 640 KNQHKNKISTSEMRMLHWMCGKTRHGRIRNETL 738
>TC82300
Length = 589
Score = 51.6 bits (122), Expect = 3e-07
Identities = 38/99 (38%), Positives = 51/99 (51%), Gaps = 8/99 (8%)
Frame = +2
Query: 11 MYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIF--------TLVLDVLTEHIQELAP 62
MYE + SV T G EDFPIT G++ GS LSPY T +V+T
Sbjct: 299 MYERGTASVTTLGGNVEDFPITKGVN*GSVLSPYFSIGSGFVY*T*AREVVT------GS 460
Query: 63 RCMLFADDVVLVGDSREEVNGRLETWRQALEAYGFCLSR 101
+ ML +++VLV +S E +N L+ Q LE Y L+R
Sbjct: 461 QGMLVGNEIVLVEES*EAINKILKIR*QVLETYDNRLTR 577
>AL375910
Length = 244
Score = 45.4 bits (106), Expect = 2e-05
Identities = 28/48 (58%), Positives = 32/48 (66%), Gaps = 8/48 (16%)
Frame = -1
Query: 48 LVLDVLTEHIQELAPRCM------LFADDVVLVGDSREEVNG--RLET 87
+ LDVLTE+IQELA RCM F DD VL G+ REE+ G RLET
Sbjct: 181 IFLDVLTENIQELALRCMFFFFLFFFTDDAVLHGELREELMGG*RLET 38
Score = 30.0 bits (66), Expect = 0.89
Identities = 28/79 (35%), Positives = 35/79 (43%), Gaps = 4/79 (5%)
Frame = -2
Query: 28 DFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELAPRCMLFADDVVLVGDSREEVNGR--- 84
DF TIG HQ F + L ++ C F+ + + S E GR
Sbjct: 231 DFRRTIGFHQVVNHKFLSFWMYLQKTSKS*H*DV--CFFFSFFFLRMMQSSMEN*GRN*W 58
Query: 85 -LETWRQALEAYGFCLSRS 102
+ RQALEAYGFCLS S
Sbjct: 57 VVRDLRQALEAYGFCLSGS 1
>TC91881
Length = 860
Score = 41.6 bits (96), Expect = 3e-04
Identities = 19/52 (36%), Positives = 31/52 (59%)
Frame = +3
Query: 182 KFYRTAVRSALLYGTECWAVKSQHENQVSVAEMRMLRWMSGKTRHDRIRNDI 233
++Y +++ +CWAVKSQ E S+A+MRML + G ++I+N I
Sbjct: 309 RYYYCTSLKLVIFRIDCWAVKSQ*ERMTSIAKMRMLW*ICGHIGRNKIKNKI 464
Score = 36.2 bits (82), Expect = 0.012
Identities = 17/35 (48%), Positives = 22/35 (62%)
Frame = +1
Query: 90 QALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKV 124
Q L FCL+RSK EY+ C FS + +L+VKV
Sbjct: 16 QTLNFNDFCLTRSKLEYIGCRFSRGHTNVSLDVKV 120
>CB065457
Length = 831
Score = 30.0 bits (66), Expect(2) = 0.016
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = +3
Query: 152 VSHRIQVGWLKWRRAS 167
V+H IQ GWLKW+RAS
Sbjct: 48 VNH*IQDGWLKWQRAS 95
Score = 24.6 bits (52), Expect(2) = 0.016
Identities = 11/14 (78%), Positives = 11/14 (78%)
Frame = +1
Query: 135 FKYLESFVQNDGEI 148
FKYL S V NDGEI
Sbjct: 1 FKYLGSKVHNDGEI 42
>AW774740
Length = 739
Score = 33.9 bits (76), Expect = 0.062
Identities = 15/15 (100%), Positives = 15/15 (100%)
Frame = +1
Query: 239 RESGGSTYSRKVGRK 253
RESGGSTYSRKVGRK
Sbjct: 4 RESGGSTYSRKVGRK 48
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 33.5 bits (75), Expect = 0.081
Identities = 31/110 (28%), Positives = 49/110 (44%), Gaps = 17/110 (15%)
Frame = -3
Query: 34 GLHQGSTLSPYIFTLVLDVLTEHIQE-----------LAPRC-----MLFADDVVLVGDS 77
GL QG LSPY+F L +VL+ Q+ +A C +LFADD + G S
Sbjct: 385 GLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLLFADDTMFFGKS 206
Query: 78 REEVNG-RLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGD 126
L + A G C++ +K+ FS + S++ ++ G+
Sbjct: 205 NASSCAILLSIMDKYRAASGRCIN*TKS---AITFSSKTSQAIIDRVKGE 65
>BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinase root
isoform (EC 2.7.1.137) (PI3-kinase) (PtdIns-3-kinase)
(PI3K), partial (7%)
Length = 821
Score = 29.6 bits (65), Expect = 1.2
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +1
Query: 184 YRTAVRSALLYGTECWAVK 202
YRT +R + Y T+CW VK
Sbjct: 757 YRTVIRPMIFYETKCWGVK 813
>CA990124 weakly similar to PIR|A96697|A96 protein F1N21.18 [imported] -
Arabidopsis thaliana, partial (72%)
Length = 537
Score = 28.5 bits (62), Expect = 2.6
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = +2
Query: 159 GWLKWRRASCVLCDKKVPFKLKGKFYRTAVRSALLYGTEC 198
G + W ++SC LC K+ KF + VRS L C
Sbjct: 272 GSIWWGKSSCKLCSYKI*TNCANKFSKNMVRSQPLSTIPC 391
>BG448790
Length = 652
Score = 28.5 bits (62), Expect = 2.6
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +2
Query: 82 NGRLETWRQALEAYGFCLSRSKTEYMECNF 111
N +L+ W +E + F S SK E +EC F
Sbjct: 422 NEKLKVWSHIVETHSFG*SXSKIENIECKF 511
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 28.1 bits (61), Expect = 3.4
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = +1
Query: 34 GLHQGSTLSPYIFTLVLDVLT 54
GL QG LSPY+F L +VL+
Sbjct: 13 GLRQGDPLSPYLFILCANVLS 75
>CA921373 weakly similar to PIR|T14519|T14 probable S-receptor kinase (EC
2.7.1.-) SFR1 - wild cabbage, partial (13%)
Length = 779
Score = 27.7 bits (60), Expect = 4.4
Identities = 9/14 (64%), Positives = 10/14 (71%)
Frame = +2
Query: 160 WLKWRRASCVLCDK 173
WL WRR C+L DK
Sbjct: 422 WLTWRRKKCILTDK 463
>TC78521 PIR|A25642|HSZM4 histone H4 - maize, complete
Length = 579
Score = 27.3 bits (59), Expect = 5.8
Identities = 23/95 (24%), Positives = 45/95 (47%), Gaps = 7/95 (7%)
Frame = -3
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFT----LVLDVLTEHIQELAPRCMLFADD 70
+S++ + +G++ +H+G+ L P + + + D+L E +Q+ + + D
Sbjct: 397 SSSTSESIEGSSLPLQSIDNIHRGNCLPPSVLSVCDRITDDILQEDLQDTSSFFIDQTTD 218
Query: 71 VVLVGDSREEVNGRL-ETWRQALEAYG--FCLSRS 102
+ SR+ NGRL +T E + FC S S
Sbjct: 217 PLHTTSSRQSTNGRLSDTLDVVAENFSVPFCTSLS 113
>AJ389014 PIR|A25642|HS histone H4 - maize, partial (98%)
Length = 568
Score = 27.3 bits (59), Expect = 5.8
Identities = 23/87 (26%), Positives = 41/87 (46%), Gaps = 7/87 (8%)
Frame = -2
Query: 23 DGTTEDFPITIGLHQGSTLSPYIFT----LVLDVLTEHIQELAPRCMLFADDVVLVGDSR 78
+G++ F +H+G+ L P + + + D+L E +Q+ + + D + SR
Sbjct: 357 EGSSLPFQSIDNIHRGNCLPPSMLSVCDRITDDILQEDLQDTSGFFIDQTTDPLHTTSSR 178
Query: 79 EEVNGRL-ETWRQALEAYG--FCLSRS 102
+ NGRL +T E + FC S S
Sbjct: 177 QSTNGRLSDTLDVVAENFSVPFCTSLS 97
>TC87351 similar to GP|10176988|dbj|BAB10220. GTP-binding protein-like
{Arabidopsis thaliana}, partial (81%)
Length = 2266
Score = 26.9 bits (58), Expect = 7.5
Identities = 14/34 (41%), Positives = 17/34 (49%)
Frame = -3
Query: 110 NFSGRRSKSTLEVKVGDHIIPQVTRFKYLESFVQ 143
+F GRRSK + +K G I RFK F Q
Sbjct: 782 HFIGRRSKRAINIKEGFRFIHNKNRFKLRCIFAQ 681
>TC91500 similar to SP|Q9BBS8|RPOC_LOTJA DNA-directed RNA polymerase beta'
chain (EC 2.7.7.6). {Lotus japonicus}, partial (22%)
Length = 1004
Score = 26.9 bits (58), Expect = 7.5
Identities = 8/27 (29%), Positives = 17/27 (62%)
Frame = +3
Query: 160 WLKWRRASCVLCDKKVPFKLKGKFYRT 186
WL+WR C++ ++VP ++ + + T
Sbjct: 237 WLRWRIDQCIMSSREVPIEVHYESFGT 317
>TC86184 similar to GP|21553481|gb|AAM62574.1 putative ribosomal protein
{Arabidopsis thaliana}, partial (97%)
Length = 554
Score = 26.6 bits (57), Expect = 9.9
Identities = 16/47 (34%), Positives = 22/47 (46%)
Frame = -2
Query: 87 TWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTLEVKVGDHIIPQVT 133
TW Q + CLSR + C+FS S++TLE +P T
Sbjct: 349 TWYQTKQQLSKCLSRCSSLLPRCSFSS-LSETTLETARNVAQVPHPT 212
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,709,582
Number of Sequences: 36976
Number of extensions: 109481
Number of successful extensions: 537
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 536
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 537
length of query: 253
length of database: 9,014,727
effective HSP length: 94
effective length of query: 159
effective length of database: 5,538,983
effective search space: 880698297
effective search space used: 880698297
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126792.7


