

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126788.1 - phase: 0 /pseudo
(981 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
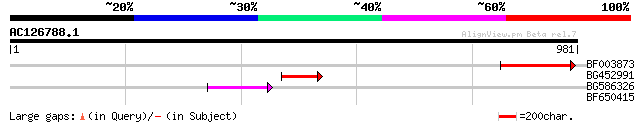
Score E
Sequences producing significant alignments: (bits) Value
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 107 2e-23
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 52 9e-07
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 50 6e-06
BF650415 similar to GP|20196875|gb unknown protein {Arabidopsis ... 31 2.7
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 107 bits (267), Expect = 2e-23
Identities = 69/129 (53%), Positives = 80/129 (61%)
Frame = +3
Query: 850 VH*NLRS*LQSLLAHIRYLRESGQWLIGWVCPLIFRICMTCSTCRSCGSMYRILRILFLG 909
V *+ RS*L L IRY +E +W I W IF ICM CR+ GSM+RI +
Sbjct: 12 VL*SQRS*L*DSLVRIRYQKELERWRIEWDYHRIF*ICMMSFMCRNFGSMFRIHLM*SRV 191
Query: 910 MTCKLETTLRWKPYR*ELMIGR*NP*EVKRYHL*ESFGVERLEKV*LGSWKVRCGSCTQS 969
M CK ETTLR + YR* LMI +* *E +RY L*ESFG ER+ K * GS +VR S QS
Sbjct: 192 MMCKSETTLR*RLYR*GLMIEK*RH*EARRYLL*ESFGTERMVKA*RGSLRVRWWSLIQS 371
Query: 970 CLFEVRFRG 978
CL EV FRG
Sbjct: 372 CLHEVNFRG 398
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 52.4 bits (124), Expect = 9e-07
Identities = 34/72 (47%), Positives = 44/72 (60%)
Frame = +1
Query: 470 LNSLET*VLFANCHPRVCSWVCRKSIVNS*IVSEKRNRLMSSLWI**LIVMKLRIVISKL 529
LNS ET*V F C RV +W C +S N IVS++ R M S I* L +++L++VI KL
Sbjct: 70 LNSSET*VWFVKCRHRV*NWGC*RSTTNFWIVSKRLRRWM*SW*I*CLGIIRLKMVILKL 249
Query: 530 MNKVC*SSDAEF 541
M K C S E+
Sbjct: 250 MIKECCSFGIEY 285
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 49.7 bits (117), Expect = 6e-06
Identities = 48/113 (42%), Positives = 57/113 (49%)
Frame = +3
Query: 342 FTVMRLSWV*EVF*CKMERW*RMLLGS*GCMKRIILLTIYSWLRWFLF*RFGGTTCMVPD 401
F RLS * +* M R GS* M+ I WLR + *RFG TCMVP
Sbjct: 54 FIQTRLSLD*VAY*PSMRRSSPTRQGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMVPR 233
Query: 402 SKCSVTIRA*NIYLIRRS*T*GRGDG*SY*RIMILG*TIIRVKPM*LQML*VG 454
+ TI+ *+I+L S*T*GRG G + * I IIR K + Q L* G
Sbjct: 234 FRYIRTIKV*SIFLPSLS*T*GRGGGWNS*LTTI*TSLIIREKLIW*QTL*AG 392
>BF650415 similar to GP|20196875|gb unknown protein {Arabidopsis thaliana},
partial (82%)
Length = 666
Score = 30.8 bits (68), Expect = 2.7
Identities = 17/39 (43%), Positives = 25/39 (63%)
Frame = -1
Query: 222 KRRSCMPNCQSASFG*VK*VFLAILFLVVVLQWILQKLM 260
+RRSC +C +A F ++ +FL LFL + L +LQ LM
Sbjct: 174 ERRSCSFSCSAALFCSLETIFLCFLFLNLSLFLLLQILM 58
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.373 0.166 0.653
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,129,858
Number of Sequences: 36976
Number of extensions: 539713
Number of successful extensions: 7435
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1592
Number of HSP's successfully gapped in prelim test: 272
Number of HSP's that attempted gapping in prelim test: 5723
Number of HSP's gapped (non-prelim): 2125
length of query: 981
length of database: 9,014,727
effective HSP length: 105
effective length of query: 876
effective length of database: 5,132,247
effective search space: 4495848372
effective search space used: 4495848372
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126788.1


