

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126787.14 - phase: 0
(370 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
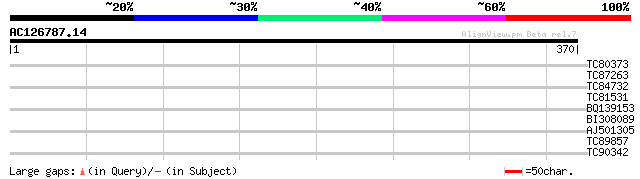
Score E
Sequences producing significant alignments: (bits) Value
TC80373 weakly similar to PIR|T51373|T51373 hypothetical protein... 33 0.18
TC87263 similar to GP|9759298|dbj|BAB09804.1 emb|CAB82953.1~gene... 32 0.51
TC84732 homologue to GP|14334996|gb|AAK59762.1 At1g05230/YUP8H12... 30 1.5
TC81531 homologue to GP|3925363|gb|AAC79430.1| homeodomain prote... 30 1.9
BQ139153 similar to GP|19310589|gb unknown protein {Arabidopsis ... 29 2.5
BI308089 similar to GP|22136748|gb unknown protein {Arabidopsis ... 29 3.3
AJ501305 weakly similar to GP|19310589|gb unknown protein {Arabi... 27 9.6
TC89857 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220 ... 27 9.6
TC90342 similar to PIR|C96757|C96757 hypothetical protein T18K17... 27 9.6
>TC80373 weakly similar to PIR|T51373|T51373 hypothetical protein F1N13_40 -
Arabidopsis thaliana, partial (35%)
Length = 899
Score = 33.1 bits (74), Expect = 0.18
Identities = 25/96 (26%), Positives = 40/96 (41%), Gaps = 8/96 (8%)
Frame = +1
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESV- 169
W W +ECE ++++ G ++ VGDS R SLL L+ + E V
Sbjct: 307 WRWKP----NECELPLFNATQFLEIVRGKKMVFVGDSVGRNQLQSLLCLLSQVSQPEDVS 474
Query: 170 -----RSFLFKRH--SDYHIDIDEMGLKLDFIWAPY 198
FKR+ +DY+ + +W+PY
Sbjct: 475 HKYTTNVIYFKRYFFADYNFTLGN-------LWSPY 561
>TC87263 similar to GP|9759298|dbj|BAB09804.1
emb|CAB82953.1~gene_id:MPH15.5~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (65%)
Length = 2229
Score = 31.6 bits (70), Expect = 0.51
Identities = 35/128 (27%), Positives = 53/128 (41%), Gaps = 23/128 (17%)
Frame = +2
Query: 122 CEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSE---------------RV 166
C RL ++DLL G ++ VGDS R SL+ ++ +S R
Sbjct: 971 CSLPRLDGHRMLDLLRGKRLVFVGDSLNRNMWESLICILKNSVKDKKKVYEANGRVHFRG 1150
Query: 167 ESVRSFLFKRHSDYHIDID-EMGLKLDFIWAPYPNNLT-------DLVMRFKKKHLYPDV 218
E+ SF+FK DY ++ + L W N T DLV R ++ D+
Sbjct: 1151 EASYSFVFK---DYKFSVELFVSPFLVQEWEMPDKNGTKKETLRLDLVGRSSDQYKDADI 1321
Query: 219 LVMGSGLW 226
+V +G W
Sbjct: 1322 IVFNTGHW 1345
>TC84732 homologue to GP|14334996|gb|AAK59762.1 At1g05230/YUP8H12_16
{Arabidopsis thaliana}, partial (35%)
Length = 808
Score = 30.0 bits (66), Expect = 1.5
Identities = 19/57 (33%), Positives = 24/57 (41%), Gaps = 1/57 (1%)
Frame = +3
Query: 195 WAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLR-NSVTS 250
W P P +R + D+L G + M HI N D G + LLR NS S
Sbjct: 303 WLPVPPKRVFEFLRDENSRSEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSANS 473
>TC81531 homologue to GP|3925363|gb|AAC79430.1| homeodomain protein {Malus x
domestica}, partial (25%)
Length = 888
Score = 29.6 bits (65), Expect = 1.9
Identities = 18/55 (32%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = +3
Query: 194 IWAPY-PNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLLRNS 247
+W P P L D +R ++ D+L G + M HI D+G + LLR S
Sbjct: 6 VWLPVSPQRLFDF-LRDERLRSEWDILSNGGPMQEMAHIAKGQDHGNCVSLLRAS 167
>BQ139153 similar to GP|19310589|gb unknown protein {Arabidopsis thaliana},
partial (31%)
Length = 673
Score = 29.3 bits (64), Expect = 2.5
Identities = 16/51 (31%), Positives = 23/51 (44%)
Frame = +3
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV 160
+W W SEC+ R + + + + VGDS AR SLL L+
Sbjct: 420 YWRWKP----SECKLPRFEPNTFLQFIENKHMAFVGDSMARNQLESLLCLL 560
>BI308089 similar to GP|22136748|gb unknown protein {Arabidopsis thaliana},
partial (49%)
Length = 849
Score = 28.9 bits (63), Expect = 3.3
Identities = 15/58 (25%), Positives = 27/58 (45%)
Frame = +2
Query: 107 YDKFWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSE 164
Y + W W +C+ R + ++L+ G + +GDS AR S+L ++ E
Sbjct: 65 YYENWRWKPF----QCDIPRFDPRKFLELMRGKTLAFIGDSVARNQMESMLCILWQVE 226
>AJ501305 weakly similar to GP|19310589|gb unknown protein {Arabidopsis
thaliana}, partial (27%)
Length = 620
Score = 27.3 bits (59), Expect = 9.6
Identities = 15/51 (29%), Positives = 23/51 (44%)
Frame = +2
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV 160
+W W +EC+ R + + L + VGDS AR SLL ++
Sbjct: 389 YWRWKP----NECKLPRFEPNTFLQLSKNKHIAFVGDSLARNQLESLLCML 529
>TC89857 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220
{Arabidopsis thaliana}, partial (34%)
Length = 952
Score = 27.3 bits (59), Expect = 9.6
Identities = 10/40 (25%), Positives = 22/40 (55%)
Frame = -1
Query: 330 SLSLNCGIKCTDDGMHYDGAVYEAELHIMFNALLIESHQK 369
S+ + G KC ++ +H D + A + ++ + +E HQ+
Sbjct: 877 SILITIGAKCNNNTLHLDKLCF*ANQELKYSGM*LEHHQQ 758
>TC90342 similar to PIR|C96757|C96757 hypothetical protein T18K17.20
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1533
Score = 27.3 bits (59), Expect = 9.6
Identities = 19/88 (21%), Positives = 35/88 (39%)
Frame = +3
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W EC R ++ +L +M +GDS R S++ LV + +
Sbjct: 342 WRWQP----HECNLPRFDPLKLLHMLRNKRMMFIGDSLQRGQFESMICLV--QSVIPEGK 503
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
L + ++E +++ WAP+
Sbjct: 504 KSLQRIPPMKIFRVEEFNATIEYYWAPF 587
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,348,886
Number of Sequences: 36976
Number of extensions: 184086
Number of successful extensions: 795
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 795
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 795
length of query: 370
length of database: 9,014,727
effective HSP length: 98
effective length of query: 272
effective length of database: 5,391,079
effective search space: 1466373488
effective search space used: 1466373488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126787.14


