

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
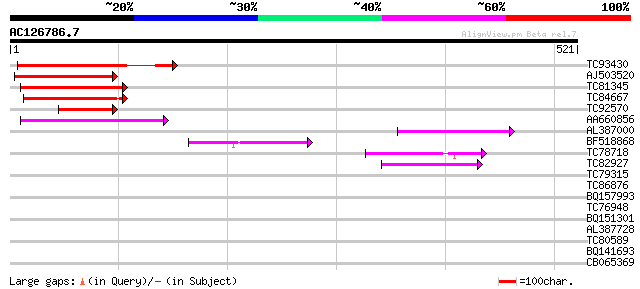
Query= AC126786.7 - phase: 0 /pseudo
(521 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93430 weakly similar to GP|10177112|dbj|BAB10402. Mlo protein-... 134 1e-31
AJ503520 weakly similar to GP|15290605|gb| seven transmembrane p... 119 2e-27
TC81345 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtM... 105 4e-23
TC84667 similar to GP|12541095|emb|CAC25084. unnamed protein pro... 95 7e-20
TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane... 75 6e-14
AA660856 similar to GP|12541099|em unnamed protein product {Arab... 71 9e-13
AL387000 similar to GP|15290605|gb seven transmembrane protein M... 68 8e-12
BF518868 similar to GP|12541097|em unnamed protein product {Arab... 52 4e-07
TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtM... 48 1e-05
TC82927 similar to GP|12541095|emb|CAC25084. unnamed protein pro... 44 1e-04
TC79315 similar to PIR|B84847|B84847 hypothetical protein At2g41... 33 0.35
TC86876 similar to GP|16973296|emb|CAC80857. C-type MADS box pro... 30 2.3
BQ157993 similar to GP|13122418|dbj putative glycin-rich protein... 30 3.0
TC76948 similar to SP|Q08935|ROC1_NICSY 29 kDa ribonucleoprotein... 29 3.9
BQ151301 similar to PIR|S34666|S346 glycine-rich protein - commo... 26 4.6
AL387728 similar to GP|14091574|gb| membrane protein Mlo2 {Arabi... 29 5.0
TC80589 similar to PIR|B86468|B86468 probable DNA polymerase typ... 28 8.6
BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Sch... 28 8.6
CB065369 homologue to PIR|A36930|DAP protocatechuate 3 4-dioxyge... 28 8.6
>TC93430 weakly similar to GP|10177112|dbj|BAB10402. Mlo protein-like
{Arabidopsis thaliana}, partial (15%)
Length = 775
Score = 134 bits (336), Expect = 1e-31
Identities = 68/147 (46%), Positives = 90/147 (60%)
Frame = +3
Query: 8 GDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMV 67
G S+ LD TPTWAV VC + +++SI+LEK +HK + +R K ALLEALEK+KAELM+
Sbjct: 375 GSSKDLDKTPTWAVGCVCTVFILVSIILEKSLHKIGTWLGDRHKKALLEALEKVKAELMI 554
Query: 68 LGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNI 127
LGF+SLLLTFGQ+Y+ K+C+P ++ MLPCQ A R A +
Sbjct: 555 LGFLSLLLTFGQSYMVKLCMPANATDIMLPCQSRATRDAKN------------------- 677
Query: 128 PAGEGESKGEHHRRLLSYERRFLGGGG 154
+ EHHR+LL YERR+ G
Sbjct: 678 ------DEDEHHRKLLYYERRYXSXDG 740
>AJ503520 weakly similar to GP|15290605|gb| seven transmembrane protein MLO2
{Oryza sativa} [Oryza sativa (indica cultivar-group)],
partial (16%)
Length = 339
Score = 119 bits (299), Expect = 2e-27
Identities = 59/95 (62%), Positives = 71/95 (74%)
Frame = +2
Query: 5 GGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAE 64
GG R L TPTWAVAAVCA+ VIIS+L+E IHK F ++ K A+ EALEKIK E
Sbjct: 8 GGAPGERTLKETPTWAVAAVCAVFVIISVLIEHGIHKLEKWFHKKHKKAMSEALEKIKEE 187
Query: 65 LMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQ 99
LM+LGFISLLLTFG YI+K+CIP K ++MLPC+
Sbjct: 188 LMLLGFISLLLTFGTKYIAKICIPNKLGDSMLPCK 292
>TC81345 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtMlo12).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (24%)
Length = 654
Score = 105 bits (262), Expect = 4e-23
Identities = 51/98 (52%), Positives = 70/98 (71%)
Frame = +2
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
R L+ TPTWAVA VC +++ ISI++E +IH F+++ K+AL EALEK+K ELM++GF
Sbjct: 143 RTLEETPTWAVAVVCFVLLAISIVIEHIIHAIGKWFKKKNKNALYEALEKVKGELMLMGF 322
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADH 108
ISLLLT Q+YISK+CI K +T PC +TA +
Sbjct: 323 ISLLLTEYQDYISKICISEKVGSTWHPCSTPKTKTASN 436
>TC84667 similar to GP|12541095|emb|CAC25084. unnamed protein product
{Arabidopsis thaliana}, partial (16%)
Length = 385
Score = 94.7 bits (234), Expect = 7e-20
Identities = 46/96 (47%), Positives = 64/96 (65%)
Frame = +3
Query: 13 LDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGFIS 72
L+ TPTW VA VC +IV IS+ +E+L+H + + + +L EAL+KIK ELM+LGFIS
Sbjct: 6 LEFTPTWVVAGVCTVIVAISLAVERLLHYGGKFLKSKDQKSLYEALQKIKEELMLLGFIS 185
Query: 73 LLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADH 108
LLLT QN ++K+C+P MLPC L ++ H
Sbjct: 186 LLLTVSQNRLTKICVPPGVLRHMLPCS-LEDKKESH 290
>TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane protein
MLO2 {Oryza sativa} [Oryza sativa (indica
cultivar-group)], partial (47%)
Length = 944
Score = 75.1 bits (183), Expect = 6e-14
Identities = 35/54 (64%), Positives = 44/54 (80%)
Frame = +3
Query: 46 FEERKKHALLEALEKIKAELMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQ 99
F ++ K A+ EALEKIK ELM+LGFISLLLTFG YI+K+CIP K ++MLPC+
Sbjct: 63 FHKKHKKAMSEALEKIKEELMLLGFISLLLTFGTKYIAKICIPNKLGDSMLPCK 224
>AA660856 similar to GP|12541099|em unnamed protein product {Arabidopsis
thaliana}, partial (29%)
Length = 633
Score = 71.2 bits (173), Expect = 9e-13
Identities = 44/137 (32%), Positives = 70/137 (50%), Gaps = 1/137 (0%)
Frame = +1
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
R L TPT++VA+V ++V + L+E+ I++F ++ ++ AL +LEKIK ELM+LG
Sbjct: 37 RSLAETPTYSVASVVTVMVFVCFLVERSIYRFGKWLKKTRRKALFASLEKIKEELMLLGL 216
Query: 71 ISLLLTFGQNYISKVCIPVK-YSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIPA 129
ISLLL +IS++C+ +S+ C +H E + +
Sbjct: 217 ISLLLAQSARWISEICVNSSLFSSRFYTCSEQDLPMIEHVMIVNSSSFSDETTIPKGLYS 396
Query: 130 GEGESKGEHHRRLLSYE 146
G GE H +SYE
Sbjct: 397 GALHQCGEGHEPFVSYE 447
>AL387000 similar to GP|15290605|gb seven transmembrane protein MLO2 {Oryza
sativa} [Oryza sativa (indica cultivar-group)], partial
(24%)
Length = 464
Score = 68.2 bits (165), Expect = 8e-12
Identities = 46/108 (42%), Positives = 59/108 (54%)
Frame = +3
Query: 357 QVFLV*MASISSLSPPLCPLSECI*ADLFLVDMV*IWVGILLL*R*QSYDC*SCSWGGCS 416
QVFLV* + L + EC LFLVD+V*++ ILL * + C WG +
Sbjct: 75 QVFLV*SPPLDHLLDSFYLIPECFPNSLFLVDLV*VYNHILLP*ELATNSDKGCPWGSLT 254
Query: 417 ICLQLYHTSSLCTRHTDGINDEAVNI*RANIKGIAKLA*KCNA*ETIK 464
+QL++ SSLCT TDGI+ E NI* AN K K+A *E +K
Sbjct: 255 SPMQLHNISSLCTSDTDGISHEERNI*GANNKST*KMAKGSKR*EEVK 398
>BF518868 similar to GP|12541097|em unnamed protein product {Arabidopsis
thaliana}, partial (20%)
Length = 378
Score = 52.4 bits (124), Expect = 4e-07
Identities = 46/116 (39%), Positives = 60/116 (51%), Gaps = 2/116 (1%)
Frame = +1
Query: 165 YISYIFSFSSWLSSM*YSVPLQ*HLVEQRLKVGKNGSKTI--WSMKMP*MILEDSGLLMR 222
Y SY S+ S S M*Y+V Q + R + G++G + + M +P +IL+DS L M+
Sbjct: 34 YTSYTSSYFS*PSFM*YTVLSQCFSGD*RYEDGRHGRQRLRLTDMSLP-LILQDSVLHMK 210
Query: 223 HPL*GITTVSGQKHLLLSILYASFDSS*DRFEGQTT*QCDMDLFLSI*HLEVSLIF 278
H L + G K SIL AS DSS TT* C DL S H EV+L F
Sbjct: 211 HHLLDNMPLFGLKFQSSSILVASLDSSIGLSRNLTT*LCATDLSRSTSHREVNLTF 378
>TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtMlo12).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (42%)
Length = 1625
Score = 47.8 bits (112), Expect = 1e-05
Identities = 40/115 (34%), Positives = 60/115 (51%), Gaps = 4/115 (3%)
Frame = +1
Query: 328 ASSNYN*NGTRHTRKTCSSAGDTSCTSL*QVFLV*MASISSLSPPLCPLSECI*ADLFLV 387
A++++N +GT+ +R S+ G T + FLV + SSL C LSEC+ +F +
Sbjct: 586 ATNDHNKDGTKDSR*RRSNQGCTCG*ARRSPFLVQ*SQPSSLYNSSCSLSECLSTCIFFL 765
Query: 388 DMV*IWVGILLL*R*QSYDC----*SCSWGGCSICLQLYHTSSLCTRHTDGINDE 438
+ +*I LL +C *S S +I + L H +SLC+ HTDG N E
Sbjct: 766 EYI*ILYKFLL----PQNNCR*CH*SLSRDFNTISM*LCHFASLCSSHTDGFNHE 918
>TC82927 similar to GP|12541095|emb|CAC25084. unnamed protein product
{Arabidopsis thaliana}, partial (44%)
Length = 810
Score = 44.3 bits (103), Expect = 1e-04
Identities = 32/93 (34%), Positives = 46/93 (49%)
Frame = +1
Query: 342 KTCSSAGDTSCTSL*QVFLV*MASISSLSPPLCPLSECI*ADLFLVDMV*IWVGILLL*R 401
K CS SCT++ LV AS SL L PL +C * +F +DM IW+ +L
Sbjct: 499 KACSHRR*LSCTTIR*SLLVSSASHRSLLDSLYPLPKCF*DSIFFLDMGYIWI*LLYNGT 678
Query: 402 *QSYDC*SCSWGGCSICLQLYHTSSLCTRHTDG 434
+ + WG S + L H ++L + +TDG
Sbjct: 679 SSLHCSKARYWGIYSGTMXLQHPTTLRSXYTDG 777
>TC79315 similar to PIR|B84847|B84847 hypothetical protein At2g41870
[imported] - Arabidopsis thaliana, partial (57%)
Length = 1293
Score = 32.7 bits (73), Expect = 0.35
Identities = 19/33 (57%), Positives = 20/33 (60%), Gaps = 2/33 (6%)
Frame = -2
Query: 148 RFLGGGGGGP--GCKPWTGYISYIFSFSSWLSS 178
R +GGGGGG G K W IS I S SSW SS
Sbjct: 251 RLVGGGGGGRRGGVKAWISLISVICS-SSWFSS 156
>TC86876 similar to GP|16973296|emb|CAC80857. C-type MADS box protein {Malus x
domestica}, partial (85%)
Length = 2050
Score = 30.0 bits (66), Expect = 2.3
Identities = 34/150 (22%), Positives = 53/150 (34%), Gaps = 12/150 (8%)
Frame = +1
Query: 27 IIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGFISLLLTFGQNYIS--K 84
II++ +L F S + KK L+ + +++L F+S+ +FG
Sbjct: 1126 IIIVPHFILRFSSFSFCSQIQG-KKMVTLQTTTSMCCFMIMLLFLSVTKSFGDLVTDHKN 1302
Query: 85 VCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIPAGEGESKGEHHRRLLS 144
+ K PC ++ P P+ P T + P P P G G +
Sbjct: 1303 LTTVTKDEVKCTPCGQVSSPPPPSPPPPSPPPASTNNCPPPPSPPSSGGGSGSTYYSPPP 1482
Query: 145 YERRFL----------GGGGGGPGCKPWTG 164
+ GGGGGG P TG
Sbjct: 1483 PSVYYYSSPPPPASTGGGGGGGLYYPPPTG 1572
>BQ157993 similar to GP|13122418|dbj putative glycin-rich protein {Oryza
sativa (japonica cultivar-group)}, partial (5%)
Length = 1045
Score = 29.6 bits (65), Expect = 3.0
Identities = 17/49 (34%), Positives = 23/49 (46%), Gaps = 6/49 (12%)
Frame = -3
Query: 135 KGEHHRRLLSYERRFLGGGGGGPGCK------PWTGYISYIFSFSSWLS 177
KG R ++ GGGGG G + +TG++S FS WLS
Sbjct: 206 KGGGGRDRFDLKKEIKAGGGGGRGSRLRGMIQGYTGWVSSSFSVPFWLS 60
>TC76948 similar to SP|Q08935|ROC1_NICSY 29 kDa ribonucleoprotein A
chloroplast precursor (CP29A). [Wood tobacco] {Nicotiana
sylvestris}, partial (61%)
Length = 1374
Score = 29.3 bits (64), Expect = 3.9
Identities = 22/70 (31%), Positives = 31/70 (43%), Gaps = 1/70 (1%)
Frame = -2
Query: 89 VKYSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIPAGEGESKGEHHRRLLSYERR 148
+ +++ PC P + +A A P T P+ P G G +R S +R
Sbjct: 773 ITFASKTCPCSPNNDSSAKLSTPQARLPT*TRLSPSEEPPLG-GPLGRSPPKRGFSPKRE 597
Query: 149 F-LGGGGGGP 157
F L GGGGGP
Sbjct: 596 FSLRGGGGGP 567
>BQ151301 similar to PIR|S34666|S346 glycine-rich protein - common tobacco,
partial (22%)
Length = 530
Score = 25.8 bits (55), Expect(2) = 4.6
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = -2
Query: 130 GEGESKGEHHRRLLSYERRFLGGGGGG 156
G G +G H+ Y R +GGGGGG
Sbjct: 187 GGGGGRGGHN-----YRPRLMGGGGGG 122
Score = 21.6 bits (44), Expect(2) = 4.6
Identities = 9/26 (34%), Positives = 13/26 (49%)
Frame = -1
Query: 151 GGGGGGPGCKPWTGYISYIFSFSSWL 176
GGGGGG G + ++F W+
Sbjct: 140 GGGGGGGG----VVFFFFVFGGGGWV 75
>AL387728 similar to GP|14091574|gb| membrane protein Mlo2 {Arabidopsis
thaliana}, partial (14%)
Length = 465
Score = 28.9 bits (63), Expect = 5.0
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +3
Query: 416 SICLQLYHTSSLCTRHTDGINDE 438
+I + L H +SLC+ HTDG N E
Sbjct: 66 TISM*LCHFASLCSSHTDGFNHE 134
>TC80589 similar to PIR|B86468|B86468 probable DNA polymerase type I
54894-56354 [imported] - Arabidopsis thaliana, partial
(67%)
Length = 1416
Score = 28.1 bits (61), Expect = 8.6
Identities = 19/41 (46%), Positives = 24/41 (58%), Gaps = 7/41 (17%)
Frame = +1
Query: 125 PNIPAGEGESKGEHHRRLL-SYE---RRFL---GGGGGGPG 158
P I +GE E+ R LL SY+ R+F+ GGGGGG G
Sbjct: 106 PVIAVFDGERGSEYRRNLLPSYKANRRKFVARGGGGGGGSG 228
>BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Schistosoma
japonicum}, partial (12%)
Length = 1226
Score = 28.1 bits (61), Expect = 8.6
Identities = 20/42 (47%), Positives = 24/42 (56%), Gaps = 2/42 (4%)
Frame = -2
Query: 116 PQGTEHEPTPNI--PAGEGESKGEHHRRLLSYERRFLGGGGG 155
P+G +PTP I P EGE KGE R+ +R GGGGG
Sbjct: 118 PKGKNTKPTPRITKPEREGE-KGE--RKEKRGGKRGKGGGGG 2
>CB065369 homologue to PIR|A36930|DAP protocatechuate 3 4-dioxygenase (EC
1.13.11.3) beta chain [validated] - Pseudomonas putida,
partial (71%)
Length = 526
Score = 28.1 bits (61), Expect = 8.6
Identities = 22/57 (38%), Positives = 26/57 (45%), Gaps = 2/57 (3%)
Frame = -1
Query: 101 LAERTADHPGEPALEP--QGTEHEPTPNIPAGEGESKGEHHRRLLSYERRFLGGGGG 155
LA+R A HPG P P Q P N+ AG+ RR LS + R L G G
Sbjct: 331 LADRRAHHPGRPCARPVRQAGSQHPGGNL-AGQ-------RRRPLSPQERPLPGPAG 185
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.354 0.158 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,001,564
Number of Sequences: 36976
Number of extensions: 291494
Number of successful extensions: 6625
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 2463
Number of HSP's successfully gapped in prelim test: 176
Number of HSP's that attempted gapping in prelim test: 1676
Number of HSP's gapped (non-prelim): 5064
length of query: 521
length of database: 9,014,727
effective HSP length: 100
effective length of query: 421
effective length of database: 5,317,127
effective search space: 2238510467
effective search space used: 2238510467
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126786.7


