

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
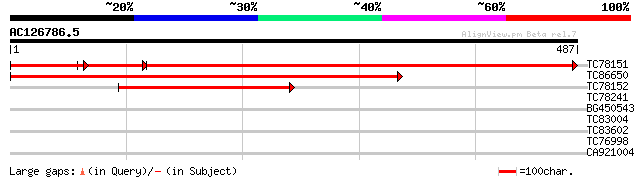
TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10 ... 734 0.0
TC86650 similar to PIR|C84428|C84428 probable ribophorin I [impo... 427 e-120
TC78152 similar to PIR|T04482|T04482 ribophorin I homolog - barl... 310 8e-85
TC78241 similar to GP|13899121|gb|AAK48982.1 Unknown protein {Ar... 31 1.2
BG450543 weakly similar to GP|20198135|g putative chromosome ass... 29 4.6
TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 29 4.6
TC83602 similar to GP|22136892|gb|AAM91790.1 unknown protein {Ar... 28 6.1
TC76998 similar to GP|11994729|dbj|BAB03045. gene_id:F5N5.17~pir... 28 6.1
CA921004 weakly similar to GP|7110148|gb DNA repair-recombinatio... 28 7.9
>TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10 {Arabidopsis
thaliana}, partial (88%)
Length = 2171
Score = 734 bits (1894), Expect(3) = 0.0
Identities = 369/370 (99%), Positives = 370/370 (99%)
Frame = +1
Query: 118 EHYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSS 177
+HYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSS
Sbjct: 805 KHYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSS 984
Query: 178 LWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVI 237
LWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVI
Sbjct: 985 LWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVI 1164
Query: 238 DTLVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQV 297
DTLVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQV
Sbjct: 1165 DTLVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQV 1344
Query: 298 YYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQV 357
YYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQV
Sbjct: 1345 YYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQV 1524
Query: 358 HSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQ 417
HSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQ
Sbjct: 1525 HSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQ 1704
Query: 418 AAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEI 477
AAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEI
Sbjct: 1705 AAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEI 1884
Query: 478 DDLMDLIDEI 487
DDLMDLIDEI
Sbjct: 1885 DDLMDLIDEI 1914
Score = 134 bits (336), Expect(3) = 0.0
Identities = 67/67 (100%), Positives = 67/67 (100%)
Frame = +2
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL
Sbjct: 452 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 631
Query: 61 ENTKLQG 67
ENTKLQG
Sbjct: 632 ENTKLQG 652
Score = 113 bits (283), Expect(3) = 0.0
Identities = 54/60 (90%), Positives = 57/60 (95%)
Frame = +3
Query: 59 KLENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITE 118
K++N +GSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITE
Sbjct: 633 KIQN--FRGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITE 806
>TC86650 similar to PIR|C84428|C84428 probable ribophorin I [imported] -
Arabidopsis thaliana, partial (94%)
Length = 1782
Score = 427 bits (1097), Expect = e-120
Identities = 205/338 (60%), Positives = 259/338 (75%), Gaps = 1/338 (0%)
Frame = +1
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+++L + TH L+PFP +I+Q++ QL+ F++SA LSPY VK Q+ +K P AR+ES+T +
Sbjct: 427 LEVLYILTHSLEPFPVEISQSESQLVYFRDSAILLSPYHVKQQTTFIKTPTARVESFTVV 606
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
E TK G+ELKYGPY+ P+SY P+++HFENN PFAV +EL REIEISHWG++QITE Y
Sbjct: 607 EPTKRAGTELKYGPYDEQTPYSYSPVLVHFENNNPFAVVEELEREIEISHWGSLQITERY 786
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
LVH GA+ KG FSR++YQ R G ++F+ L AKLPPR HSVYYRDEIGNIS+S L
Sbjct: 787 KLVHAGARHKGVFSRVEYQTRSGASGVASFKHLLAKLPPRVHSVYYRDEIGNISSSHLRT 966
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLF-GVDGKRFLNISFGSPINELVIDT 239
D K+ELE EPRYPLFGGWK+ F +GYGLPLQDFLF DG+R+LN ++G P+ + V+D
Sbjct: 967 DFLKSELEFEPRYPLFGGWKSTFLLGYGLPLQDFLFESPDGRRYLNFTYGCPLAQTVVDK 1146
Query: 240 LVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYY 299
L++KVVLPEGSKD + +PF V + ETK+S+LDI GR VVVLEK N VPEHN FQVYY
Sbjct: 1147LIIKVVLPEGSKDPAAVIPFQVDQHLETKYSYLDIVGRTVVVLEKRNVVPEHNIPFQVYY 1326
Query: 300 KFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKT 337
FN + ML EPLML S FF LF AS+ Y+H DLSI K+
Sbjct: 1327NFNPIFMLAEPLMLASVFFLLFAASVTYLHMDLSIRKS 1440
>TC78152 similar to PIR|T04482|T04482 ribophorin I homolog - barley
(fragment), partial (31%)
Length = 680
Score = 310 bits (794), Expect = 8e-85
Identities = 151/151 (100%), Positives = 151/151 (100%)
Frame = +3
Query: 94 QPFAVAKELVREIEISHWGNVQITEHYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRL 153
QPFAVAKELVREIEISHWGNVQITEHYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRL
Sbjct: 3 QPFAVAKELVREIEISHWGNVQITEHYNLVHGGAQSKGEFSRLDYQARPYVRGASAFRRL 182
Query: 154 TAKLPPRAHSVYYRDEIGNISTSSLWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQD 213
TAKLPPRAHSVYYRDEIGNISTSSLWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQD
Sbjct: 183 TAKLPPRAHSVYYRDEIGNISTSSLWGDSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQD 362
Query: 214 FLFGVDGKRFLNISFGSPINELVIDTLVVKV 244
FLFGVDGKRFLNISFGSPINELVIDTLVVKV
Sbjct: 363 FLFGVDGKRFLNISFGSPINELVIDTLVVKV 455
>TC78241 similar to GP|13899121|gb|AAK48982.1 Unknown protein {Arabidopsis
thaliana}, partial (56%)
Length = 786
Score = 30.8 bits (68), Expect = 1.2
Identities = 23/85 (27%), Positives = 42/85 (49%), Gaps = 10/85 (11%)
Frame = +1
Query: 374 SLRDLSRTGDIQGCKATRKSVDSSLKE---------LTKELKSPV-TFLQSSPQAAQILP 423
SL + R G + C+A K VDS++K+ K KS + T + +A ++L
Sbjct: 178 SLNTVQRRGVVVVCEAAPKKVDSAIKKALQAERRRVYNKARKSEIRTRTKKVLEALELLK 357
Query: 424 KVEEVIAKERDLQEKLMAKHSTVVD 448
K + A+E EK++ + +++D
Sbjct: 358 KKSDAQAEEITSIEKMIGEAYSIID 432
>BG450543 weakly similar to GP|20198135|g putative chromosome associated
protein {Arabidopsis thaliana}, partial (10%)
Length = 682
Score = 28.9 bits (63), Expect = 4.6
Identities = 23/110 (20%), Positives = 50/110 (44%)
Frame = +1
Query: 371 LEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIA 430
L ++DL G I G + ++ L L +++ + L + I P + ++
Sbjct: 82 LSXDVKDLQ--GKISGNRHAKEEAAEQLAILENKIQGSMDELNN------ISPLYDNLVQ 237
Query: 431 KERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDL 480
KE+D+ + +M + + Y+K+ + ++ A + L++E DDL
Sbjct: 238 KEKDITKGIMEREKKLSILYQKQGRATQFSSKAARDKW----LQKETDDL 375
>TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (78%)
Length = 1250
Score = 28.9 bits (63), Expect = 4.6
Identities = 22/82 (26%), Positives = 39/82 (46%), Gaps = 3/82 (3%)
Frame = +1
Query: 404 ELKSPVTFLQSSPQAAQILPKVEEVIAKERDL---QEKLMAKHSTVVDCYEKKLGGREIE 460
EL++P S+PQ +LP E + ER+L + K + S K+ E+
Sbjct: 685 ELRNPSPISTSAPQPCGVLPP-EAWMQNERELKRERRKQSNRESARRSRLRKQAEAEELA 861
Query: 461 NRIASHQQKITALKQEIDDLMD 482
R+ + + ALK E+++L +
Sbjct: 862 RRVDALTAENLALKSEMNELAE 927
>TC83602 similar to GP|22136892|gb|AAM91790.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 1167
Score = 28.5 bits (62), Expect = 6.1
Identities = 28/130 (21%), Positives = 61/130 (46%), Gaps = 2/130 (1%)
Frame = +2
Query: 324 SIVYMHADLSISKTSASYLAKIQWEE--VQATIHQVHSIVSRCLTTHDKLEASLRDLSRT 381
++V + A L +++ S +++ E+ ++ +++ H I+ + D LE L + +
Sbjct: 233 TVVELEAQLEEVRSAPSKKDELEKEKAVLEGDVNKFHKIIEEFGSRIDPLERVLMEKEKQ 412
Query: 382 GDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMA 441
+ + + R + E +ELKS V + + + K E+ A ERD E +A
Sbjct: 413 LEAKVVENER------IVEENEELKSKVELQTFNARDVDRMKK--ELQAAERDASEAELA 568
Query: 442 KHSTVVDCYE 451
+++ C+E
Sbjct: 569 RNAWEEKCWE 598
>TC76998 similar to GP|11994729|dbj|BAB03045.
gene_id:F5N5.17~pir||G71408~similar to unknown protein
{Arabidopsis thaliana}, partial (62%)
Length = 1704
Score = 28.5 bits (62), Expect = 6.1
Identities = 17/67 (25%), Positives = 32/67 (47%)
Frame = +1
Query: 14 FPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTKLQGSELKYG 73
FP + Q +L F ++ P+ +K+Q ++ ++P RI+ K +L + G
Sbjct: 283 FPNYLLQNLEKLFKF*RFLNFVMPFTMKIQPIDSQIPAERIKPVVKSRLKRLFERQFS-G 459
Query: 74 PYENLPP 80
+N PP
Sbjct: 460 VLKNPPP 480
>CA921004 weakly similar to GP|7110148|gb DNA repair-recombination protein
{Arabidopsis thaliana}, partial (11%)
Length = 809
Score = 28.1 bits (61), Expect = 7.9
Identities = 20/53 (37%), Positives = 30/53 (55%)
Frame = +1
Query: 425 VEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEI 477
V + +ER+LQ +L K D E +L EIEN+ + +QKI AL +E+
Sbjct: 358 VSHLDERERELQIRLEGKIKQR-DEREFELKKSEIENKALNVEQKIRALNREM 513
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,730,848
Number of Sequences: 36976
Number of extensions: 187854
Number of successful extensions: 1073
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1065
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1071
length of query: 487
length of database: 9,014,727
effective HSP length: 100
effective length of query: 387
effective length of database: 5,317,127
effective search space: 2057728149
effective search space used: 2057728149
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126786.5


