

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.6 - phase: 0
(135 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
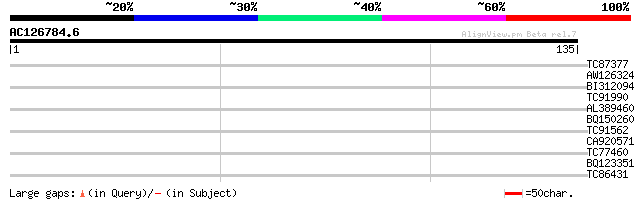
Score E
Sequences producing significant alignments: (bits) Value
TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like prot... 34 0.015
AW126324 similar to GP|20161818|db putative laccase {Oryza sativ... 27 2.5
BI312094 similar to PIR|D86397|D863 protein T7N9.12 [imported] -... 27 2.5
TC91990 weakly similar to SP|Q25504|CU16_MANSE Larval cuticle pr... 27 2.5
AL389460 similar to SP|O46157|RL13_ 60S ribosomal protein L13. [... 27 3.2
BQ150260 27 3.2
TC91562 similar to PIR|T07172|T07172 subtilisin-like proteinase ... 26 4.2
CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {A... 26 5.5
TC77460 similar to GP|6728993|gb|AAF26990.1| fatty acid multifun... 25 7.1
BQ123351 25 9.3
TC86431 similar to SP|Q8W425|PSD6_ORYSA 26S proteasome non-ATPas... 25 9.3
>TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (54%)
Length = 2071
Score = 34.3 bits (77), Expect = 0.015
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = +1
Query: 40 QYFKPKPTGVVGEDGLPLQQSHFRGRLLEGTTLPLPH 76
++F P TG VG GL L + H ++ GT++ PH
Sbjct: 811 KHFSPGWTGAVGPTGLALDKRHVNFNIISGTSMSCPH 921
>AW126324 similar to GP|20161818|db putative laccase {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 535
Score = 26.9 bits (58), Expect = 2.5
Identities = 14/32 (43%), Positives = 17/32 (52%), Gaps = 3/32 (9%)
Frame = -1
Query: 105 TYWNHDSVPS---NNDDFSRAFHWIPVAEAVS 133
T WN + VP N FSR F WI +A + S
Sbjct: 172 TSWNANGVPLHRINQIIFSRVFSWIKIANSSS 77
>BI312094 similar to PIR|D86397|D863 protein T7N9.12 [imported] - Arabidopsis
thaliana, partial (14%)
Length = 728
Score = 26.9 bits (58), Expect = 2.5
Identities = 16/47 (34%), Positives = 20/47 (42%)
Frame = -2
Query: 78 YSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFH 124
YSGF K N + NS + + W HD+ P DFS H
Sbjct: 607 YSGF---KCNVA*TQNSMRSKIV------WKHDTAPCKFGDFSGCIH 494
>TC91990 weakly similar to SP|Q25504|CU16_MANSE Larval cuticle protein 16/17
precursor. [Tobacco hawkmoth Tobacco hornworm] {Manduca
sexta}, partial (82%)
Length = 436
Score = 26.9 bits (58), Expect = 2.5
Identities = 14/44 (31%), Positives = 21/44 (46%)
Frame = +1
Query: 9 IDLKRDGIEDLSGRVHLLPCCIKHDGPTEVSQYFKPKPTGVVGE 52
+DLK + L+ RV+ + PT+ S + P PT V E
Sbjct: 145 LDLKPQTVSKLTNRVN*RKLWTRRTSPTQSSLFAVPTPTSTVKE 276
>AL389460 similar to SP|O46157|RL13_ 60S ribosomal protein L13. [Humus
earthworm] {Lumbricus rubellus}, partial (18%)
Length = 454
Score = 26.6 bits (57), Expect = 3.2
Identities = 15/64 (23%), Positives = 33/64 (51%)
Frame = -3
Query: 72 LPLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDFSRAFHWIPVAEA 131
LPLP Y G + +++ + ++ + S+ N + + ++ + P + D F+W A
Sbjct: 392 LPLPWPYDGLHVHQQDGLVGASFTQNSICANSLFFISNSTEPYSPLD-PCLFNW*DCALL 216
Query: 132 VSCL 135
+SC+
Sbjct: 215 LSCI 204
>BQ150260
Length = 519
Score = 26.6 bits (57), Expect = 3.2
Identities = 13/24 (54%), Positives = 13/24 (54%)
Frame = -1
Query: 41 YFKPKPTGVVGEDGLPLQQSHFRG 64
YF PKP G GE LP Q F G
Sbjct: 135 YFNPKPWGKKGEFFLPKAQLKFGG 64
>TC91562 similar to PIR|T07172|T07172 subtilisin-like proteinase (EC
3.4.21.-) 2 - tomato, partial (36%)
Length = 1062
Score = 26.2 bits (56), Expect = 4.2
Identities = 9/30 (30%), Positives = 16/30 (53%)
Frame = +2
Query: 47 TGVVGEDGLPLQQSHFRGRLLEGTTLPLPH 76
+G+ G LP+ + +L GT++ PH
Sbjct: 155 SGLTGPSSLPIDHRRVKFNILSGTSMSCPH 244
>CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {Arabidopsis
thaliana}, partial (48%)
Length = 827
Score = 25.8 bits (55), Expect = 5.5
Identities = 9/27 (33%), Positives = 16/27 (58%)
Frame = +2
Query: 94 SWETSVTFNDITYWNHDSVPSNNDDFS 120
+W + + F D +WN + S+N D+S
Sbjct: 359 TWPSFLQF*DRVFWNVSVISSSNKDYS 439
>TC77460 similar to GP|6728993|gb|AAF26990.1| fatty acid multifunctional
protein (AtMFP2) {Arabidopsis thaliana}, partial (98%)
Length = 2594
Score = 25.4 bits (54), Expect = 7.1
Identities = 19/68 (27%), Positives = 27/68 (38%), Gaps = 1/68 (1%)
Frame = +2
Query: 61 HFRGRL-LEGTTLPLPHNYSGFVLGKKNSVESSNSWETSVTFNDITYWNHDSVPSNNDDF 119
H G + G TL P Y+ L +N +E+S+S VP +
Sbjct: 2132 HLTGEVSYSGLTLLDPSTYTQGCLNGQNYMENSSS----------------PVPIWLQEL 2263
Query: 120 SRAFHWIP 127
R FHW+P
Sbjct: 2264 PRGFHWVP 2287
>BQ123351
Length = 725
Score = 25.0 bits (53), Expect = 9.3
Identities = 9/21 (42%), Positives = 14/21 (65%)
Frame = +3
Query: 82 VLGKKNSVESSNSWETSVTFN 102
+ GKK+S E + W T++T N
Sbjct: 33 IRGKKDS*EGGHDWSTNLTLN 95
>TC86431 similar to SP|Q8W425|PSD6_ORYSA 26S proteasome non-ATPase
regulatory subunit 6, partial (97%)
Length = 1517
Score = 25.0 bits (53), Expect = 9.3
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -2
Query: 4 GGEAKIDLKRDGIEDLSGRVHLLPCC 29
G E KI K ++ + +VHLLP C
Sbjct: 562 GNEIKIHAKETKLQCIKYQVHLLPNC 485
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.136 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,027,507
Number of Sequences: 36976
Number of extensions: 72584
Number of successful extensions: 344
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 344
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 344
length of query: 135
length of database: 9,014,727
effective HSP length: 86
effective length of query: 49
effective length of database: 5,834,791
effective search space: 285904759
effective search space used: 285904759
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 53 (25.0 bits)
Medicago: description of AC126784.6


