

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
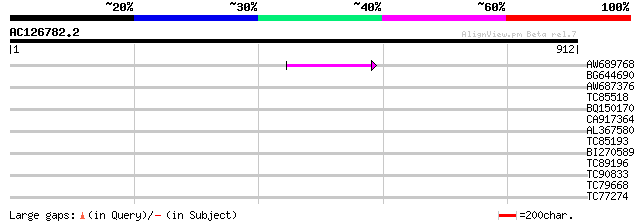
Query= AC126782.2 - phase: 0 /pseudo
(912 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 64 3e-10
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 35 0.13
AW687376 33 0.65
TC85518 similar to PIR|T45902|T45902 inorganic pyrophosphatase-l... 31 2.5
BQ150170 30 3.2
CA917364 30 4.2
AL367580 30 5.5
TC85193 similar to GP|14423502|gb|AAK62433.1 Unknown protein {Ar... 30 5.5
BI270589 similar to PIR|C96829|C96 unknown protein F19K16.21 [im... 30 5.5
TC89196 similar to GP|19401700|gb|AAL87667.1 transcription facto... 30 5.5
TC90833 similar to PIR|T05154|T05154 hypothetical protein F18E5.... 29 7.2
TC79668 similar to GP|9955513|emb|CAC05452.1 putative protein {A... 29 7.2
TC77274 similar to GP|22136956|gb|AAM91707.1 unknown protein {Ar... 29 9.3
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 63.5 bits (153), Expect = 3e-10
Identities = 55/144 (38%), Positives = 69/144 (47%)
Frame = +2
Query: 446 LFGPCGFLDIKKNLTVLLRGIKPDL*VMVQVNRLALTMEKLLAQL*NQLLFAQFSALLHP 505
L GF+++KK LL +KPD + V V L + KL *N L FS LL
Sbjct: 182 LLAASGFIELKKTQMDLLTNLKPD*WLRVSVKHLDVITLKLSLL**NLSLSGSFSPLLSL 361
Query: 506 DLGKFISWMSRMLFFMVN*KKQCTCTNLWDLKILKTPIMYAFFANHSMGLNKLLELGVSD 565
GKF +S M F MV K++C C NL DLK+L + A NHSM +K G+
Sbjct: 362 INGKFNKLISTMPF*MVFFKRKCICLNLRDLKLL-INLWCAS*TNHSMA*SKHHVHGMRX 538
Query: 566 LPTTCHLLVFLKANVITLFSCTRK 589
P LV LK VI + T K
Sbjct: 539 *PVHKFSLVSLKVVVIHHYLSTIK 610
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 35.0 bits (79), Expect = 0.13
Identities = 25/77 (32%), Positives = 37/77 (47%)
Frame = -3
Query: 502 LLHPDLGKFISWMSRMLFFMVN*KKQCTCTNLWDLKILKTPIMYAFFANHSMGLNKLLEL 561
LLH WM R+ M K++C +NL DLK+ + IM + H M +KL E
Sbjct: 306 LLHSWGSSCTKWM*RVHLLMEISKRRCLSSNLLDLKMQRYQIMCSD*IRHYMV*SKLQEH 127
Query: 562 GVSDLPTTCHLLVFLKA 578
G+ + C +V +A
Sbjct: 126 GMKGCQSFC*RMVSKEA 76
>AW687376
Length = 400
Score = 32.7 bits (73), Expect = 0.65
Identities = 19/58 (32%), Positives = 32/58 (54%), Gaps = 1/58 (1%)
Frame = -2
Query: 110 LKN*EHTSTHNFKEKLKQFNVTTAVNMSIVIFKK-CVKLMELFFVCLVLTHLLKMVNL 166
+K E HN + Q + T ++++SI+IF CV LM +F+C ++ H K+ L
Sbjct: 399 IKMIECIKLHNHQHLPSQHH-TNSISLSIIIFSSNCVSLM*YYFICNIVQH*HKITQL 229
>TC85518 similar to PIR|T45902|T45902 inorganic pyrophosphatase-like protein
- Arabidopsis thaliana, partial (87%)
Length = 1620
Score = 30.8 bits (68), Expect = 2.5
Identities = 24/74 (32%), Positives = 39/74 (52%), Gaps = 2/74 (2%)
Frame = +2
Query: 298 LNLIHTLS*TMSCPLILYNI*WTK-PKLAHLILNWL-ISQTKSLLLVQTHQISPHLHNLT 355
L LIH ++* + L L++I W K K A L +NW + L+ THQ+S H+ +
Sbjct: 713 LLLIHGMT*RLDQELHLFSIVWLKLAKEARLNMNWTRKADLLRLIGFFTHQLSIHITMVL 892
Query: 356 PLVPSNKIFLPYFN 369
P K+ +P+ +
Sbjct: 893 FQEPFVKMKIPWMS 934
>BQ150170
Length = 994
Score = 30.4 bits (67), Expect = 3.2
Identities = 18/37 (48%), Positives = 21/37 (56%)
Frame = +1
Query: 568 TTCHLLVFLKANVITLFSCTRKTLIWHIFYSRLMILF 604
T HL LK V+TL KTLIW IF+ +ILF
Sbjct: 10 THSHLHFHLK*LVLTLNIIYFKTLIWLIFFPLFLILF 120
>CA917364
Length = 699
Score = 30.0 bits (66), Expect = 4.2
Identities = 24/95 (25%), Positives = 45/95 (47%), Gaps = 1/95 (1%)
Frame = +1
Query: 278 LFVVMSYL-MKILSHMQSCTFLNLIHTLS*TMSCPLILYNI*WTKPKLAHLILNWLISQT 336
++V +S L +K +SH+ ++ + + CPL+L+ + W LI
Sbjct: 262 IWVSLSRLSLKRISHLSFMHYVGVWSLFRVLLVCPLLLWRL-WPSV---------LILLE 411
Query: 337 KSLLLVQTHQISPHLHNLTPLVPSNKIFLPYFNQL 371
+ + V HQ++PH N+ V ++F P+ QL
Sbjct: 412 RKIAGVHGHQVAPHPQNMH--VYFGRVFKPFLLQL 510
>AL367580
Length = 479
Score = 29.6 bits (65), Expect = 5.5
Identities = 16/50 (32%), Positives = 24/50 (48%), Gaps = 5/50 (10%)
Frame = -2
Query: 59 VCCHLIFYIVMYGPLLF*VQLVTNIMFFSWIII-----QIFCGLFQCHIN 103
+CC I Y+V++G LL N+ F W+ + + C F C IN
Sbjct: 391 ICCQNIIYVVIFGDLL-------NVPLFPWLHLFDKCKRCIC*HFVCIIN 263
>TC85193 similar to GP|14423502|gb|AAK62433.1 Unknown protein {Arabidopsis
thaliana}, partial (73%)
Length = 1351
Score = 29.6 bits (65), Expect = 5.5
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = +2
Query: 397 TPMLQNPLYLGTLCLHLKTQIGKWSWMM 424
+P+ QNP + LC +L ++ KW W++
Sbjct: 68 SPISQNPNFFSFLC*NLHFELKKWLWLL 151
>BI270589 similar to PIR|C96829|C96 unknown protein F19K16.21 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 651
Score = 29.6 bits (65), Expect = 5.5
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 5/47 (10%)
Frame = -2
Query: 273 CQIKLLFVVMSYLMKILSHMQSCTFLNLIHTLS-----*TMSCPLIL 314
C ++LL V +SY+ + + S TFL++ L T SCPL L
Sbjct: 566 CLLQLLLVFLSYVSNLFLKLSSTTFLSVSFLLCRKIFLPTFSCPLSL 426
>TC89196 similar to GP|19401700|gb|AAL87667.1 transcription factor RAU1
{Oryza sativa}, partial (63%)
Length = 1312
Score = 29.6 bits (65), Expect = 5.5
Identities = 24/105 (22%), Positives = 53/105 (49%), Gaps = 6/105 (5%)
Frame = -3
Query: 501 ALLHPDLGKFISWMSRMLFFMVN*KKQCTCTNLWDLKILKTPIMYAFFANHSMGLNK--L 558
A+L +LG+ W+ ++ K + ++++ + +P++ AFF ++ L++ L
Sbjct: 875 AILEEELGRLK*WVKPSFWYP---KSTF*VSKAFEVENMNSPLLEAFFGALNVELSQ*SL 705
Query: 559 LELGVSDLPTTCHLLVFLKANVITLF----SCTRKTLIWHIFYSR 599
++LG+ D +FL ++ F SC+ T+ WH+ +R
Sbjct: 704 VKLGM*DFVIQLSFPMFL---LLIQFA*RTSCSFSTITWHLDRAR 579
>TC90833 similar to PIR|T05154|T05154 hypothetical protein F18E5.80 -
Arabidopsis thaliana, partial (19%)
Length = 803
Score = 29.3 bits (64), Expect = 7.2
Identities = 29/108 (26%), Positives = 52/108 (47%), Gaps = 8/108 (7%)
Frame = -3
Query: 187 HLHFGTML*KWQHIS*IFYQAKLLILSHLLKCCITKILLTITYECLV---VYVFLSFLHQ 243
HL F + K+ + IF + L +LS K + I I+ ++ V +F FLH
Sbjct: 660 HLLFYVIWEKFIPVKHIFKNS-LTLLSLCHKAMLNTIFNNISPFLIIHIKVLIFNHFLHN 484
Query: 244 KYTNF----NLVPQNVCFLRIRQIIEAINV*IYCQIKLLF-VVMSYLM 286
++ NL+ +CFL+I I I++ C +K + +++S+ M
Sbjct: 483 MISSIIQIINLIISFMCFLKIIFKIHIIHIINRCHLKFIHNLIISFFM 340
>TC79668 similar to GP|9955513|emb|CAC05452.1 putative protein {Arabidopsis
thaliana}, partial (75%)
Length = 757
Score = 29.3 bits (64), Expect = 7.2
Identities = 18/45 (40%), Positives = 26/45 (57%), Gaps = 3/45 (6%)
Frame = -1
Query: 116 TSTHNFKEKLK---QFNVTTAVNMSIVIFKKCVKLMELFFVCLVL 157
TS FK+ L QFN+ + + I+K CV L++ FF+CL L
Sbjct: 583 TSL*TFKDSLLLACQFNIC*LL---LNIYKPCVNLLQQFFLCLFL 458
>TC77274 similar to GP|22136956|gb|AAM91707.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2013
Score = 28.9 bits (63), Expect = 9.3
Identities = 32/124 (25%), Positives = 56/124 (44%), Gaps = 15/124 (12%)
Frame = +2
Query: 32 ENYILIFVIHVLLVNMLN--------CLS*I-LIIEVCCHLIFYIVMYGPLLF*VQLVTN 82
E+ L+ +I VLLV ++ CL+ I +CC L FY+V P L N
Sbjct: 476 ESLDLVIMIQVLLVLVVLS*F*V*FLCLTKIGFRFWLCCFLYFYVVKVNPWL----SCDN 643
Query: 83 IMFFSWIIIQIFCGLFQCHINP------KFIQFLKN*EHTSTHNFKEKLKQFNVTTAVNM 136
+F+ ++I F + NP +F F N ++ + +++ +NV+ ++
Sbjct: 644 ALFYCFLI--EFTNRDLSNSNPLR*T*TRFYLFFNNLKNIYKKHSFQQINTYNVSLHLSF 817
Query: 137 SIVI 140
IVI
Sbjct: 818 VIVI 829
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.346 0.153 0.515
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,631,711
Number of Sequences: 36976
Number of extensions: 635668
Number of successful extensions: 7349
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 3132
Number of HSP's successfully gapped in prelim test: 511
Number of HSP's that attempted gapping in prelim test: 4045
Number of HSP's gapped (non-prelim): 3999
length of query: 912
length of database: 9,014,727
effective HSP length: 105
effective length of query: 807
effective length of database: 5,132,247
effective search space: 4141723329
effective search space used: 4141723329
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126782.2


