

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126781.3 + phase: 0
(77 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
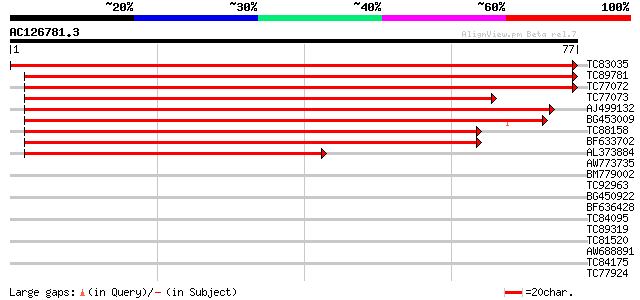
Score E
Sequences producing significant alignments: (bits) Value
TC83035 similar to GP|8843819|dbj|BAA97367.1 light-inducible pro... 155 2e-39
TC89781 similar to GP|8843819|dbj|BAA97367.1 light-inducible pro... 123 1e-29
TC77072 similar to PIR|T48186|T48186 light-inducible protein ATL... 90 1e-19
TC77073 similar to PIR|T48186|T48186 light-inducible protein ATL... 79 3e-16
AJ499132 weakly similar to PIR|T48186|T48 light-inducible protei... 75 6e-15
BG453009 weakly similar to PIR|T48186|T48 light-inducible protei... 72 3e-14
TC88158 weakly similar to PIR|T46062|T46062 LS1-like protein - A... 58 8e-10
BF633702 56 2e-09
AL373884 weakly similar to PIR|T48186|T481 light-inducible prote... 45 4e-06
AW773735 homologue to GP|13487785|gb ADP-glucose pyrophosphoryla... 28 0.85
BM779002 weakly similar to GP|3258570|gb| Unknown protein {Arabi... 26 2.5
TC92963 homologue to SP|O04865|PLD_VIGUN Phospholipase D precurs... 26 3.2
BG450922 similar to PIR|H86204|H86 hypothetical protein [importe... 25 4.2
BF636428 25 4.2
TC84095 25 4.2
TC89319 similar to PIR|T02350|T02350 hypothetical protein T8F5.5... 25 5.5
TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD460... 25 7.2
AW688891 similar to PIR|B96837|B968 hypothetical protein T21F11.... 25 7.2
TC84175 similar to GP|19896|emb|CAA41713.1|| photosystem II 23 k... 24 9.4
TC77924 similar to PIR|H84807|H84807 probable phospholipid cytid... 24 9.4
>TC83035 similar to GP|8843819|dbj|BAA97367.1 light-inducible protein
ATLS1-like {Arabidopsis thaliana}, complete
Length = 665
Score = 155 bits (393), Expect = 2e-39
Identities = 77/77 (100%), Positives = 77/77 (100%)
Frame = +2
Query: 1 MCILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYI 60
MCILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYI
Sbjct: 134 MCILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYI 313
Query: 61 KFSDVQPSFVGFNGSTF 77
KFSDVQPSFVGFNGSTF
Sbjct: 314 KFSDVQPSFVGFNGSTF 364
>TC89781 similar to GP|8843819|dbj|BAA97367.1 light-inducible protein
ATLS1-like {Arabidopsis thaliana}, complete
Length = 610
Score = 123 bits (308), Expect = 1e-29
Identities = 62/75 (82%), Positives = 67/75 (88%)
Frame = +2
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
IL+NGGV IAFGGTEEPAAYGELISIGGL P+VNAKLSSTIAQI+QT L+I SSRFYIKF
Sbjct: 218 ILLNGGVPIAFGGTEEPAAYGELISIGGLGPSVNAKLSSTIAQILQTKLYIDSSRFYIKF 397
Query: 63 SDVQPSFVGFNGSTF 77
D + SF GFNGSTF
Sbjct: 398 YDSERSFFGFNGSTF 442
>TC77072 similar to PIR|T48186|T48186 light-inducible protein ATLS1 -
Arabidopsis thaliana, complete
Length = 779
Score = 90.1 bits (222), Expect = 1e-19
Identities = 42/75 (56%), Positives = 58/75 (77%)
Frame = +1
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
I++ G V I+FGGTE+ AAYGEL+SIGGL+P VN KLS+ IA I++T L + +RF++KF
Sbjct: 211 IVLKGSVPISFGGTEQEAAYGELVSIGGLNPDVNKKLSAAIAAILETKLSVPKTRFFLKF 390
Query: 63 SDVQPSFVGFNGSTF 77
D + S G+NG+TF
Sbjct: 391 YDTKGSNFGWNGTTF 435
>TC77073 similar to PIR|T48186|T48186 light-inducible protein ATLS1 -
Arabidopsis thaliana, partial (90%)
Length = 772
Score = 79.0 bits (193), Expect = 3e-16
Identities = 36/64 (56%), Positives = 50/64 (77%)
Frame = +2
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
I++ G V I+FGGTE+ AAYGEL+SIGGL+P VN KLS+ IA I++T L + +RF++KF
Sbjct: 188 IVLKGSVPISFGGTEQEAAYGELVSIGGLNPDVNKKLSAAIAAILETKLSVPKTRFFLKF 367
Query: 63 SDVQ 66
D +
Sbjct: 368 YDTK 379
>AJ499132 weakly similar to PIR|T48186|T48 light-inducible protein ATLS1 -
Arabidopsis thaliana, partial (97%)
Length = 687
Score = 74.7 bits (182), Expect = 6e-15
Identities = 35/72 (48%), Positives = 49/72 (67%)
Frame = +1
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
+ + G + I FG TEEPAAYGE ++IG L+P +N KLS+ IA ++ T L + SRFY+KF
Sbjct: 166 VSLEGSIPICFGETEEPAAYGEFVAIGILNPDLNKKLSAEIALVLHTMLLVPKSRFYLKF 345
Query: 63 SDVQPSFVGFNG 74
D++ G NG
Sbjct: 346 YDIEGYNCGLNG 381
>BG453009 weakly similar to PIR|T48186|T48 light-inducible protein ATLS1 -
Arabidopsis thaliana, partial (88%)
Length = 656
Score = 72.4 bits (176), Expect = 3e-14
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Frame = +1
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
+ + G + I FG TEEPAAYGE ++IG L+P +N KLS+ IA ++ T L + SRFY+KF
Sbjct: 145 VSLEGSIPICFGETEEPAAYGEFVAIGILNPDLNKKLSAEIALVLHTMLLVPKSRFYLKF 324
Query: 63 SDVQ-PSFVGFN 73
D++ P F N
Sbjct: 325 YDIEVPKFTVLN 360
>TC88158 weakly similar to PIR|T46062|T46062 LS1-like protein - Arabidopsis
thaliana, partial (84%)
Length = 621
Score = 57.8 bits (138), Expect = 8e-10
Identities = 27/62 (43%), Positives = 41/62 (65%)
Frame = +1
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
+L+ V I+F G +EPAAY E+IS+GG++ V L +TI I+Q+ L I +RF++K
Sbjct: 232 VLLKSSVPISFEGNKEPAAYAEIISMGGINKEVKKNLIATIGTILQSKLSIPRTRFFLKV 411
Query: 63 SD 64
D
Sbjct: 412 FD 417
>BF633702
Length = 361
Score = 56.2 bits (134), Expect = 2e-09
Identities = 23/62 (37%), Positives = 39/62 (62%)
Frame = +2
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
+ V + + FGGT EP A+ E I IG + N +++ I+++++++LH+ RFYIKF
Sbjct: 170 VSVQANLPMTFGGTAEPTAFAEYICIGTIGGQANEGITAKISEVVESSLHVSKDRFYIKF 349
Query: 63 SD 64
D
Sbjct: 350 VD 355
>AL373884 weakly similar to PIR|T48186|T481 light-inducible protein ATLS1 -
Arabidopsis thaliana, partial (54%)
Length = 283
Score = 45.4 bits (106), Expect = 4e-06
Identities = 20/41 (48%), Positives = 29/41 (69%)
Frame = +1
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTI 43
+++ G V I+F +EPAAYGEL+S+GG++ V L STI
Sbjct: 160 VILKGSVPISFESNKEPAAYGELVSMGGINSEVKKNLISTI 282
>AW773735 homologue to GP|13487785|gb ADP-glucose pyrophosphorylase large
subunit CagpL2 {Cicer arietinum}, partial (37%)
Length = 602
Score = 27.7 bits (60), Expect = 0.85
Identities = 14/45 (31%), Positives = 26/45 (57%)
Frame = +3
Query: 26 ISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+ IGG ++ +S+ I I+TN+H ++ +F++ QPS V
Sbjct: 15 VPIGGCYRLIDIPMSNCINSGIRTNIHFNTVQFFLS----QPSLV 137
>BM779002 weakly similar to GP|3258570|gb| Unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 717
Score = 26.2 bits (56), Expect = 2.5
Identities = 15/51 (29%), Positives = 26/51 (50%)
Frame = +3
Query: 6 NGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSS 56
NGGVAI F E+ + + GG + +N ++ + + I+ + IH S
Sbjct: 561 NGGVAICFSACEDCQMAADTAAFGGKE--MNGVMTYLLTKNIREHF*IHMS 707
>TC92963 homologue to SP|O04865|PLD_VIGUN Phospholipase D precursor (EC
3.1.4.4) (PLD) (Choline phosphatase), partial (28%)
Length = 741
Score = 25.8 bits (55), Expect = 3.2
Identities = 20/75 (26%), Positives = 30/75 (39%), Gaps = 8/75 (10%)
Frame = +2
Query: 5 VNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQ--------TNLHIHSS 56
++GG A F T E AA LIS G D ++ + I+ N + S
Sbjct: 380 IDGGAAFGFPDTPEDAARAGLIS--GKDNIIDRSIQDAYINAIRRAKNFIYIENQYFLGS 553
Query: 57 RFYIKFSDVQPSFVG 71
F D++P +G
Sbjct: 554 SFAWSGEDIKPEDIG 598
>BG450922 similar to PIR|H86204|H86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (30%)
Length = 669
Score = 25.4 bits (54), Expect = 4.2
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = +1
Query: 33 PTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFVGFNGSTF 77
P N LSS+ + ++ N +IHS+ S+++P V + TF
Sbjct: 130 PRNNTTLSSSSSLLVDNNNNIHSNSLKTTSSNLKPIVVIGHPPTF 264
>BF636428
Length = 621
Score = 25.4 bits (54), Expect = 4.2
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = -3
Query: 41 STIAQIIQTNLHIHSSRFYIKFSDVQPSFVGFN 73
S I QIIQ +LHIH+S +K V + FN
Sbjct: 313 SRIEQIIQHSLHIHTS--ILKLETVIVTLATFN 221
>TC84095
Length = 793
Score = 25.4 bits (54), Expect = 4.2
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = -1
Query: 35 VNAKLSSTIAQIIQTNLHIHSSRFYIKFS-DVQPSFV 70
VN+++S TI T H H+ RF+IK + + P FV
Sbjct: 748 VNSRVSFTIFSSTITPHHKHTPRFFIKSNPQILP*FV 638
>TC89319 similar to PIR|T02350|T02350 hypothetical protein T8F5.5 -
Arabidopsis thaliana, partial (53%)
Length = 1436
Score = 25.0 bits (53), Expect = 5.5
Identities = 13/47 (27%), Positives = 23/47 (48%)
Frame = -3
Query: 26 ISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFVGF 72
+S+GG+ P + + A LH H++RF ++ P F+ F
Sbjct: 726 LSVGGVCPNTSVSSTFLCAPAKLGRLHAHTTRFPLRRQCCHP-FISF 589
>TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD46010.1
[imported] - Arabidopsis thaliana, partial (39%)
Length = 1259
Score = 24.6 bits (52), Expect = 7.2
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = -3
Query: 9 VAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQII 47
+ IAFGGT+ IG D T+ +SST+A+++
Sbjct: 978 IPIAFGGTD----------IGAEDGTLIHDVSSTVAKLV 892
>AW688891 similar to PIR|B96837|B968 hypothetical protein T21F11.15
[imported] - Arabidopsis thaliana, partial (18%)
Length = 645
Score = 24.6 bits (52), Expect = 7.2
Identities = 11/29 (37%), Positives = 13/29 (43%)
Frame = +3
Query: 48 QTNLHIHSSRFYIKFSDVQPSFVGFNGST 76
QTN H H +Y + PSF F T
Sbjct: 168 QTNNHHHDYHYYHHHHHLHPSFAPFGNYT 254
>TC84175 similar to GP|19896|emb|CAA41713.1|| photosystem II 23 kDa
polypeptide {Nicotiana tabacum}, partial (80%)
Length = 703
Score = 24.3 bits (51), Expect = 9.4
Identities = 14/43 (32%), Positives = 21/43 (48%)
Frame = +1
Query: 20 AAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
A +GE +S GG DP A A I++T+ + + Y F
Sbjct: 556 AFFGETLSEGGFDPNAVA-----TANILETSTPVIDGKKYYGF 669
>TC77924 similar to PIR|H84807|H84807 probable phospholipid
cytidylyltransferase [imported] - Arabidopsis thaliana,
partial (86%)
Length = 1691
Score = 24.3 bits (51), Expect = 9.4
Identities = 9/32 (28%), Positives = 20/32 (62%)
Frame = -2
Query: 34 TVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDV 65
++ ++ I I+T +HI+++RF+I S +
Sbjct: 298 SLTQSITMAIMHHIKTTIHINTNRFFIPSSTI 203
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,053,514
Number of Sequences: 36976
Number of extensions: 19532
Number of successful extensions: 141
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 77
length of database: 9,014,727
effective HSP length: 53
effective length of query: 24
effective length of database: 7,054,999
effective search space: 169319976
effective search space used: 169319976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC126781.3


