

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
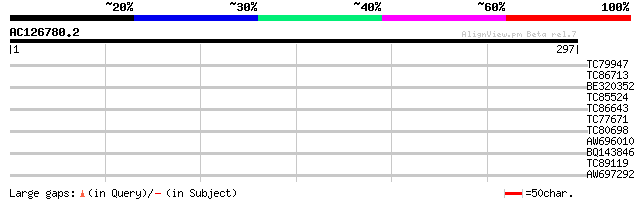
Query= AC126780.2 - phase: 0 /pseudo
(297 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79947 similar to GP|15450948|gb|AAK96745.1 Unknown protein {Ar... 38 0.004
TC86713 similar to SP|P17092|RR17_PEA 30S ribosomal protein S17 ... 29 1.9
BE320352 weakly similar to EGAD|142626|15 specific tissue protei... 29 1.9
TC85524 similar to SP|P93508|CRTC_RICCO Calreticulin precursor. ... 28 3.3
TC86643 similar to GP|5922612|dbj|BAA84613.1 EST AU078118(E3904)... 28 4.3
TC77671 27 7.3
TC80698 similar to GP|10177409|dbj|BAB10540. contains similarity... 27 7.3
AW696010 similar to GP|21553967|gb| unknown {Arabidopsis thalian... 27 7.3
BQ143846 27 9.5
TC89119 similar to PIR|T51958|T51958 hypothetical protein [impor... 27 9.5
AW697292 homologue to GP|22136708|gb putative isoamylase {Arabid... 27 9.5
>TC79947 similar to GP|15450948|gb|AAK96745.1 Unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 845
Score = 38.1 bits (87), Expect = 0.004
Identities = 22/59 (37%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Frame = +2
Query: 100 GCILGLRVSPIHRRKGVGLKLVTSIEEWM-LTNGADYAFLATEKNNNASKNLFTNKCNY 157
G I L V HR+ G+ KL+T+ + M GA+Y L K+N A+ NL+T Y
Sbjct: 296 GHITSLAVLRTHRKLGIATKLMTAAQNAMEQVFGAEYVSLHVRKSNRAAFNLYTETLGY 472
>TC86713 similar to SP|P17092|RR17_PEA 30S ribosomal protein S17
chloroplast precursor (CS17) (Fragment). [Garden pea]
{Pisum sativum}, partial (64%)
Length = 619
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = -2
Query: 136 AFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHPPT 171
+ + T ++N N TN N F SLI+ LHP T
Sbjct: 363 SLILTNRSNFTKLNNITNFENVFRIMSLILLLHPNT 256
>BE320352 weakly similar to EGAD|142626|15 specific tissue protein 2 {Cicer
arietinum}, partial (61%)
Length = 466
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -3
Query: 210 DMDIILKEKLSLGTWVSYYKDEGFKLNIEDIITHKSTTSSSWIIF 254
D+++I ++G+W + D LNI DI++ T S + IF
Sbjct: 290 DINVIFSIS*NIGSWFKIFSDSLLFLNINDIVSAS*NTRSWFEIF 156
>TC85524 similar to SP|P93508|CRTC_RICCO Calreticulin precursor. [Castor
bean] {Ricinus communis}, partial (93%)
Length = 1699
Score = 28.5 bits (62), Expect = 3.3
Identities = 17/47 (36%), Positives = 25/47 (53%)
Frame = +1
Query: 175 TNHISKKDVKIDKISIDQAISFYTRILKTKELYPLDMDIILKEKLSL 221
TNH+ KKDV + DQ YT I++ Y + +D + K+ SL
Sbjct: 550 TNHLIKKDVPCE---TDQLTHVYTFIIRPDATYSILIDNVEKQTGSL 681
>TC86643 similar to GP|5922612|dbj|BAA84613.1 EST AU078118(E3904)
corresponds to a region of the predicted gene.~similar
to Arabidopsis thaliana, partial (39%)
Length = 1705
Score = 28.1 bits (61), Expect = 4.3
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = -2
Query: 229 KDEGFKLNIEDIITHKSTTSSSWIIFSLWNTCEAC 263
+D FK++ DI++HK TT + +LWN +C
Sbjct: 555 RDCFFKVDARDIVSHKKTTVG---LPTLWNNTGSC 460
>TC77671
Length = 1156
Score = 27.3 bits (59), Expect = 7.3
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = -3
Query: 132 GADYAFLATEKNNNASKNLFTNKCNYFNFTSLIIFLHP 169
G D+AF N+ S+ C F+F SL+ F HP
Sbjct: 692 GVDFAFSTWVSFNHPSR*TCLEFCAVFSF*SLVSFNHP 579
>TC80698 similar to GP|10177409|dbj|BAB10540. contains similarity to unknown
protein~gene_id:K24M7.18~pir||T05785 {Arabidopsis
thaliana}, partial (42%)
Length = 898
Score = 27.3 bits (59), Expect = 7.3
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +2
Query: 162 SLIIFLHPPTSFPTNHIS 179
S+++ L PP SFP NH+S
Sbjct: 119 SILLTLLPPPSFPLNHVS 172
>AW696010 similar to GP|21553967|gb| unknown {Arabidopsis thaliana}, partial
(7%)
Length = 208
Score = 27.3 bits (59), Expect = 7.3
Identities = 16/37 (43%), Positives = 21/37 (56%), Gaps = 5/37 (13%)
Frame = +3
Query: 159 NFTSLIIFLHPPTSFPT----NHI-SKKDVKIDKISI 190
NF + IF+HPPT + NHI SK + I +I I
Sbjct: 3 NFHFIFIFIHPPTPYLNLHYRNHITSKSNSHISQIQI 113
>BQ143846
Length = 983
Score = 26.9 bits (58), Expect = 9.5
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 162 SLIIFLHPPTSFPTNHI 178
S ++FLH PTS P +HI
Sbjct: 424 SCVLFLHYPTSIPQSHI 474
>TC89119 similar to PIR|T51958|T51958 hypothetical protein [imported] -
Picea mariana, partial (42%)
Length = 862
Score = 26.9 bits (58), Expect = 9.5
Identities = 16/59 (27%), Positives = 27/59 (45%), Gaps = 3/59 (5%)
Frame = -1
Query: 137 FLATEKNNNASKNLFTNKC---NYFNFTSLIIFLHPPTSFPTNHISKKDVKIDKISIDQ 192
F A++ SKN+F KC +Y + T +H T N + + I ++S+ Q
Sbjct: 298 FAASKTTYALSKNMFNKKCD*ESYNHVTITKKTMHSTTIICNNKFNANRIDIKQVSLSQ 122
>AW697292 homologue to GP|22136708|gb putative isoamylase {Arabidopsis
thaliana}, partial (13%)
Length = 343
Score = 26.9 bits (58), Expect = 9.5
Identities = 13/35 (37%), Positives = 22/35 (62%)
Frame = +2
Query: 207 YPLDMDIILKEKLSLGTWVSYYKDEGFKLNIEDII 241
+P+ MD+IL SL WV+ Y +GF+ ++ I+
Sbjct: 56 HPVVMDLILD---SLRHWVTEYHVDGFRFDLASIL 151
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,075,360
Number of Sequences: 36976
Number of extensions: 153364
Number of successful extensions: 936
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 932
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 936
length of query: 297
length of database: 9,014,727
effective HSP length: 95
effective length of query: 202
effective length of database: 5,502,007
effective search space: 1111405414
effective search space used: 1111405414
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126780.2


