

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
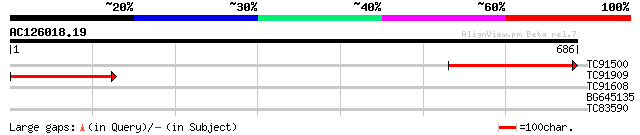
Query= AC126018.19 - phase: 0
(686 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC91500 similar to SP|Q9BBS8|RPOC_LOTJA DNA-directed RNA polymer... 328 3e-90
TC91909 homologue to PIR|S48842|S48842 DNA-directed RNA polymera... 276 1e-74
TC91608 similar to GP|14334894|gb|AAK59625.1 unknown protein {Ar... 31 1.8
BG645135 weakly similar to PIR|D84713|D84 probable dioxygenase [... 30 3.1
TC83590 similar to GP|9758365|dbj|BAB08866.1 GTP binding protein... 29 6.8
>TC91500 similar to SP|Q9BBS8|RPOC_LOTJA DNA-directed RNA polymerase beta'
chain (EC 2.7.7.6). {Lotus japonicus}, partial (22%)
Length = 1004
Score = 328 bits (842), Expect = 3e-90
Identities = 156/156 (100%), Positives = 156/156 (100%)
Frame = +3
Query: 531 PISVPTQDMLIGLYVLTSGNRRGICANRYNPFNCRNSKNEKISNNNSKYMKKKEPFFCNS 590
PISVPTQDMLIGLYVLTSGNRRGICANRYNPFNCRNSKNEKISNNNSKYMKKKEPFFCNS
Sbjct: 3 PISVPTQDMLIGLYVLTSGNRRGICANRYNPFNCRNSKNEKISNNNSKYMKKKEPFFCNS 182
Query: 591 YDAIGAYRQKRINLDSPFWLRWRIDQCIMSSREVPIEVHYESFGTYYEIYGHYLVIRSIK 650
YDAIGAYRQKRINLDSPFWLRWRIDQCIMSSREVPIEVHYESFGTYYEIYGHYLVIRSIK
Sbjct: 183 YDAIGAYRQKRINLDSPFWLRWRIDQCIMSSREVPIEVHYESFGTYYEIYGHYLVIRSIK 362
Query: 651 KEIRCIYIRTTVGHISFYREIEEAIQGFSRAYSYGI 686
KEIRCIYIRTTVGHISFYREIEEAIQGFSRAYSYGI
Sbjct: 363 KEIRCIYIRTTVGHISFYREIEEAIQGFSRAYSYGI 470
>TC91909 homologue to PIR|S48842|S48842 DNA-directed RNA polymerase (EC
2.7.7.6) beta chain - white mustard chloroplast, partial
(14%)
Length = 893
Score = 276 bits (707), Expect = 1e-74
Identities = 129/129 (100%), Positives = 129/129 (100%)
Frame = +2
Query: 1 MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI 60
MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI
Sbjct: 506 MIDQYKHQHLRIGSVSPEQISAWAKKILPNGEIVGEVTKPYTLHYKTNKPEKDGLFCERI 685
Query: 61 FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK 120
FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK
Sbjct: 686 FGPIKSGICACGNYRVIGDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLK 865
Query: 121 RLPSYIASL 129
RLPSYIASL
Sbjct: 866 RLPSYIASL 892
>TC91608 similar to GP|14334894|gb|AAK59625.1 unknown protein {Arabidopsis
thaliana}, partial (41%)
Length = 888
Score = 30.8 bits (68), Expect = 1.8
Identities = 25/104 (24%), Positives = 52/104 (49%), Gaps = 4/104 (3%)
Frame = +2
Query: 223 GEEGSADNENENEWEDRKVGRRKNFLVRRMELVKHFIRTNI-EPEWMVLSLLPVLPPELR 281
G+E +ADN++E E++D K + N L+ +++++ + + E E+ L E++
Sbjct: 209 GDEDAADNDDEVEFDDMK--EKDNILLAKLDVIDKRLEEKLAELEYTFGRKGKALEEEIK 382
Query: 282 PIIQIDGGKLMSSDINELYRRVIYRNN---TLIDLLTTSRSTPG 322
+ + +++ E RR ++R LID+ T + T G
Sbjct: 383 DLAE------ERNELTEKKRRPLFRKGFDVKLIDVNRTCKVTKG 496
>BG645135 weakly similar to PIR|D84713|D84 probable dioxygenase [imported] -
Arabidopsis thaliana, partial (44%)
Length = 812
Score = 30.0 bits (66), Expect = 3.1
Identities = 31/106 (29%), Positives = 45/106 (42%), Gaps = 7/106 (6%)
Frame = -3
Query: 573 SNNNSKYMKKKEPFFCNSYD-AIGAYRQKRINLDSPFWLRWRID----QCIMSSREVPIE 627
S+NNSK MK P+F ++ I AY I W W CI PI
Sbjct: 585 SDNNSKIMKPNIPYFLGIFNHTISAYGW*FI------WFEWMRSNSKTNCIS-----PIS 439
Query: 628 VHYESFGTYYEIYGHYLVIRSIK--KEIRCIYIRTTVGHISFYREI 671
G+ EI+ +L I S+K I ++++TT F ++I
Sbjct: 438 SLATKQGSIGEIHNFFL*ITSVKLFSYINILFMKTTDSMNHFIQDI 301
>TC83590 similar to GP|9758365|dbj|BAB08866.1 GTP binding protein-like
{Arabidopsis thaliana}, partial (49%)
Length = 1061
Score = 28.9 bits (63), Expect = 6.8
Identities = 18/74 (24%), Positives = 35/74 (46%)
Frame = -3
Query: 78 GDKKDQPKFCEQCGVEFVDSRVRRYQMGYIKLACPVTHVWYLKRLPSYIASLLDKPLKEL 137
G ++ ++C +C + S ++ GY K ACP+ V++ P+ + D+ L
Sbjct: 873 GTERRLLRYCFRCFLTD-SSSFSEHRFGYAKYACPLLFVFHPFPSPALLIDAQDESTFLL 697
Query: 138 EGLVYCDFSFARPV 151
+ +V F +PV
Sbjct: 696 DEVVGIPFVLMQPV 655
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,622,111
Number of Sequences: 36976
Number of extensions: 339923
Number of successful extensions: 1527
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1519
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1527
length of query: 686
length of database: 9,014,727
effective HSP length: 103
effective length of query: 583
effective length of database: 5,206,199
effective search space: 3035214017
effective search space used: 3035214017
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126018.19


