

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.1 + phase: 0 /pseudo
(596 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
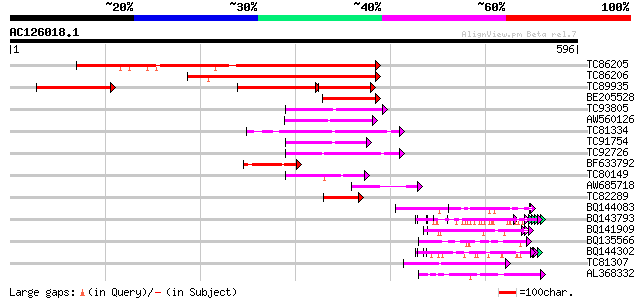
Score E
Sequences producing significant alignments: (bits) Value
TC86205 similar to PIR|A86289|A86289 probable ABC transporter [i... 402 e-112
TC86206 similar to GP|20522008|dbj|BAB92011. pleiotropic drug re... 317 8e-87
TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transp... 110 2e-46
BE205528 similar to GP|12382006|d putative ABC transporter prote... 96 3e-20
TC93805 weakly similar to GP|11994269|dbj|BAB01452. ABC transpor... 67 3e-11
AW560126 similar to GP|15292753|gb putative ABC transporter prot... 66 3e-11
TC81334 weakly similar to GP|19423992|gb|AAL87274.1 putative ABC... 63 3e-10
TC91754 similar to GP|20260310|gb|AAM13053.1 unknown protein {Ar... 62 8e-10
TC92726 similar to PIR|T04229|T04229 ABC-type transport protein ... 59 7e-09
BF633792 similar to PIR|T45888|T45 ABC transporter-like protein ... 58 9e-09
TC80149 similar to GP|22137204|gb|AAM91447.1 At2g01320/F10A8.20 ... 57 3e-08
AW685718 homologue to GP|20522008|d pleiotropic drug resistance ... 56 4e-08
TC82289 similar to GP|12382007|dbj|BAB21276. putative ABC transp... 56 4e-08
BQ144083 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 52 5e-07
BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 52 6e-07
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 50 3e-06
BQ135566 49 5e-06
BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza ... 48 9e-06
TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown prot... 48 1e-05
AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein... 47 2e-05
>TC86205 similar to PIR|A86289|A86289 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (52%)
Length = 2721
Score = 402 bits (1033), Expect = e-112
Identities = 230/361 (63%), Positives = 243/361 (66%), Gaps = 42/361 (11%)
Frame = +1
Query: 71 PSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFRCVVFHE---SDGFRIVPS--- 124
PSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF + F S FR + S
Sbjct: 1 PSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFVVLFFMSQMASGLFRAIASLGR 180
Query: 125 HRIPG*KHDCCQYIWVLCT-----SHISVIGWL-----------------------HSLK 156
+ I + L S + GW HS
Sbjct: 181 NMIVANTFGSFALLTFLSLGGFILSRKDIKGWWIWGYWISPLMYGQNALMANEFLGHS-- 354
Query: 157 NWHNATADLGKDYLDTRGFFPHAYWYWIGVGGIGWICLSFQRGVWCGSRCTRPI**A*C- 215
WHNATADLGKDYLDTRGFFPHAYWYWIGVGG+ F P
Sbjct: 355 -WHNATADLGKDYLDTRGFFPHAYWYWIGVGGLVGFVFLFNAAFGVALAVLGPFDKPSAT 531
Query: 216 -------NNN*RFRR*FIYRTRSEFTTYCSGRADSVTESSHGKKKGMVLPFEPHSITFDD 268
+++ + + R S SGR DSVTESSHGKKKGMVLPFEPHSITFDD
Sbjct: 532 ITEEDSEDDSSTVQEVELPRIES------SGRRDSVTESSHGKKKGMVLPFEPHSITFDD 693
Query: 269 IVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTG 328
IVYSVDMP EMKEQGV EDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTG
Sbjct: 694 IVYSVDMPAEMKEQGVTEDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTG 873
Query: 329 GYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTK 388
GYIDGDIKVSGYPKKQETFARISGYCE NDIHSPHVTV+ESLLYSAWLRLPSGVDSNT K
Sbjct: 874 GYIDGDIKVSGYPKKQETFARISGYCEQNDIHSPHVTVYESLLYSAWLRLPSGVDSNTRK 1053
Query: 389 V 389
+
Sbjct: 1054M 1056
>TC86206 similar to GP|20522008|dbj|BAB92011. pleiotropic drug resistance
like protein {Nicotiana tabacum}, partial (11%)
Length = 757
Score = 317 bits (812), Expect = 8e-87
Identities = 165/207 (79%), Positives = 173/207 (82%), Gaps = 5/207 (2%)
Frame = +3
Query: 188 GIGWICLSFQRGVWCGSRCT----RPI**A*CNNN*RFRR*FIYRTRSEFTTY-CSGRAD 242
GIGWIC+ FQRGVWCGSRCT + * +N+ ++ E SGR D
Sbjct: 9 GIGWICVPFQRGVWCGSRCT*AHLISLVQQ*PDNSEDDSSNYMTAQEVELPRIESSGRGD 188
Query: 243 SVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPG 302
SVT SSHGKKKGMVLPFEPHSITFDDIVYSVDMP EMKEQGV EDRLVLLKGVSGAFRPG
Sbjct: 189 SVTVSSHGKKKGMVLPFEPHSITFDDIVYSVDMPAEMKEQGVTEDRLVLLKGVSGAFRPG 368
Query: 303 VLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSP 362
VLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCE NDIHSP
Sbjct: 369 VLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEQNDIHSP 548
Query: 363 HVTVHESLLYSAWLRLPSGVDSNTTKV 389
HVTV+ESLLYSAWLRLPSGVDSNT K+
Sbjct: 549 HVTVYESLLYSAWLRLPSGVDSNTRKM 629
Score = 29.6 bits (65), Expect = 3.4
Identities = 16/34 (47%), Positives = 17/34 (49%)
Frame = +1
Query: 185 GVGGIGWICLSFQRGVWCGSRCTRPI**A*CNNN 218
GVGG+ F RPI**A*CNNN
Sbjct: 1 GVGGLAGFVFLFNAAFGVALAVPRPI**A*CNNN 102
>TC89935 similar to GP|16604683|gb|AAL24134.1 putative ABC transporter
protein {Arabidopsis thaliana}, partial (34%)
Length = 1244
Score = 110 bits (275), Expect(2) = 2e-46
Identities = 59/90 (65%), Positives = 72/90 (79%), Gaps = 3/90 (3%)
Frame = +3
Query: 240 RADSVTESSHG--KKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSG 297
R + ES+ G K+GMVLPF+P +++FD + Y VDMP EMKEQGV ++RL LL+ V+G
Sbjct: 798 RDNPTLESATGVAPKRGMVLPFQPLAMSFDSVNYYVDMPAEMKEQGVTDNRLQLLREVTG 977
Query: 298 AFRPGVLTALMGVSGAGKTTLMDVLA-GRK 326
AFRPGVLTALMGVSGAGKTTLMDV + GRK
Sbjct: 978 AFRPGVLTALMGVSGAGKTTLMDVFSLGRK 1067
Score = 103 bits (256), Expect = 2e-22
Identities = 42/83 (50%), Positives = 59/83 (70%)
Frame = +3
Query: 29 LFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWV 88
+ FT++ MFNG SE+ +TIA+LPV+YK RD LF+P W Y +P+++L+IP+S+ E +WV
Sbjct: 3 ILFTMIMNMFNGFSELPLTIARLPVFYKHRDHLFHPPWTYTLPNFLLRIPISIFEAIVWV 182
Query: 89 FLTYYVIGFDPNVGRMFKQFRCV 111
+TYY IGF P R FK V
Sbjct: 183 LITYYTIGFAPEASRFFKHLLLV 251
Score = 94.0 bits (232), Expect(2) = 2e-46
Identities = 41/60 (68%), Positives = 51/60 (84%)
Frame = +2
Query: 325 RKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDS 384
++ GGYI+GD+++SG+PK QETFARISGYCE DIHSP VTV ES++YSA+LRLP V S
Sbjct: 1061 KEEGGYIEGDVRISGFPKNQETFARISGYCEQTDIHSPQVTVRESVIYSAFLRLPREVSS 1240
>BE205528 similar to GP|12382006|d putative ABC transporter protein {Oryza
sativa (japonica cultivar-group)}, partial (14%)
Length = 617
Score = 96.3 bits (238), Expect = 3e-20
Identities = 43/60 (71%), Positives = 52/60 (86%)
Frame = +1
Query: 330 YIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKV 389
YI G+I +SG+PKKQETFARISGYCE NDIHSP+VTV+ESLLYSAWLRL +++ T K+
Sbjct: 1 YIGGNITISGFPKKQETFARISGYCEQNDIHSPYVTVYESLLYSAWLRLSPDINAETRKM 180
>TC93805 weakly similar to GP|11994269|dbj|BAB01452. ABC transporter-like
protein {Arabidopsis thaliana}, partial (15%)
Length = 649
Score = 66.6 bits (161), Expect = 3e-11
Identities = 40/108 (37%), Positives = 61/108 (56%), Gaps = 1/108 (0%)
Frame = +2
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYID-GDIKVSGYPKKQETFAR 349
+L+G++G +PG L A+MG SG GK+TL+D LAGR + G+I ++G+ KQE
Sbjct: 266 ILQGLTGYAKPGQLLAIMGPSGCGKSTLLDTLAGRLSSNTRQTGEILINGH--KQELSYG 439
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSEHVLYSE 397
S Y +D +TV E++ YSA L+LP+ + K + E
Sbjct: 440 TSAYVTQDDTLLTTLTVREAVFYSAQLQLPNTMSKEEKKERADITIKE 583
>AW560126 similar to GP|15292753|gb putative ABC transporter protein
{Arabidopsis thaliana}, partial (17%)
Length = 727
Score = 66.2 bits (160), Expect = 3e-11
Identities = 36/97 (37%), Positives = 58/97 (59%)
Frame = +3
Query: 290 VLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFAR 349
++L V+G PG + A++G SG+GK+TL++ LAGR + G I ++ K T R
Sbjct: 369 IILNNVTGIAYPGEILAILGPSGSGKSTLLNSLAGRLHTHNLTGTI-LANSSKLDRTILR 545
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNT 386
+G+ +DI PH+TV E+L++ + LRLP + T
Sbjct: 546 RTGFVTQDDILYPHLTVRETLVFCSMLRLPRQLTRET 656
>TC81334 weakly similar to GP|19423992|gb|AAL87274.1 putative ABC
transporter protein {Arabidopsis thaliana}, partial
(29%)
Length = 977
Score = 63.2 bits (152), Expect = 3e-10
Identities = 52/167 (31%), Positives = 80/167 (47%), Gaps = 1/167 (0%)
Frame = +2
Query: 250 GKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMG 309
G++KG+ L T+ D+ +V G + +L+G++G +P L A+MG
Sbjct: 47 GEEKGVCL-------TWKDLWVTVSAST-----GKTNESKSILQGLTGYAKPAQLLAIMG 190
Query: 310 VSGAGKTTLMDVLAGR-KTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHE 368
SG GK+TL+D LAGR + GDI ++G KQ S Y +D +TV E
Sbjct: 191 PSGCGKSTLLDALAGRLGSNTRQSGDILING--NKQALAYGTSAYVTQDDTLLTTLTVKE 364
Query: 369 SLLYSAWLRLPSGVDSNTTKVSEHVLYSEWIWLHLLTLLNEHDRGGG 415
++ YSA L+LP + + K E + L +N +R GG
Sbjct: 365 AVYYSAQLQLPDTMSNEEKKERADFTIRE---MGLQDAINTRNRKGG 496
>TC91754 similar to GP|20260310|gb|AAM13053.1 unknown protein {Arabidopsis
thaliana}, partial (25%)
Length = 663
Score = 61.6 bits (148), Expect = 8e-10
Identities = 37/91 (40%), Positives = 54/91 (58%), Gaps = 1/91 (1%)
Frame = +2
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI-DGDIKVSGYPKKQETFAR 349
LL G++G PG + A+MG SG+GK+TL+D LAGR + G++ ++G KK+
Sbjct: 200 LLNGLNGFAEPGRIMAIMGPSGSGKSTLLDTLAGRLAKNVVMTGNVFLNG--KKKTPGYG 373
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLPS 380
Y D+ +TV E++ YSA LRLPS
Sbjct: 374 FVAYVTQEDVLLGTLTVKETITYSAHLRLPS 466
>TC92726 similar to PIR|T04229|T04229 ABC-type transport protein F14M19.30 -
Arabidopsis thaliana, partial (20%)
Length = 585
Score = 58.5 bits (140), Expect = 7e-09
Identities = 41/126 (32%), Positives = 62/126 (48%), Gaps = 1/126 (0%)
Frame = +1
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYP-KKQETFAR 349
+LK VS P + A++G SGAGK+TL+D+L+ R+ + G + ++ P TF +
Sbjct: 187 ILKDVSLTAYPSEILAIVGPSGAGKSTLLDILSARRLP--LSGTLSLNSSPITNPSTFRK 360
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSEHVLYSEWIWLHLLTLLNE 409
+S Y +D P +TV E+ +SA L P +D S L +E HL
Sbjct: 361 LSAYVPQHDACLPMLTVSETFAFSASLLKPKTIDIAAIVSS---LLNELRLTHLANTRLN 531
Query: 410 HDRGGG 415
H GG
Sbjct: 532 HGLSGG 549
>BF633792 similar to PIR|T45888|T45 ABC transporter-like protein -
Arabidopsis thaliana, partial (3%)
Length = 355
Score = 58.2 bits (139), Expect = 9e-09
Identities = 33/61 (54%), Positives = 41/61 (67%)
Frame = +3
Query: 246 ESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLT 305
E+ GK MVLPF P SI F D+ Y VD P EMK+ G E +L LL ++GAF+PG+LT
Sbjct: 183 EAQTGK---MVLPFLPLSIAFKDVQYFVDTPPEMKKHGSNE-KLQLLCDITGAFKPGILT 350
Query: 306 A 306
A
Sbjct: 351 A 353
>TC80149 similar to GP|22137204|gb|AAM91447.1 At2g01320/F10A8.20
{Arabidopsis thaliana}, partial (44%)
Length = 1366
Score = 56.6 bits (135), Expect = 3e-08
Identities = 33/91 (36%), Positives = 53/91 (57%), Gaps = 3/91 (3%)
Frame = +3
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGG---YIDGDIKVSGYPKKQETF 347
LLK +SG +PG L A+MG SG+GKTTL++VLAG+ + ++ G ++ +G P + +
Sbjct: 402 LLKNLSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLSASPRLHLSGLLEFNGKPSSRNAY 581
Query: 348 ARISGYCEHNDIHSPHVTVHESLLYSAWLRL 378
Y D+ +TV E+L + L+L
Sbjct: 582 K--FAYVRQEDLFFSQLTVRETLSLAIELQL 668
>AW685718 homologue to GP|20522008|d pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (12%)
Length = 613
Score = 55.8 bits (133), Expect = 4e-08
Identities = 33/75 (44%), Positives = 38/75 (50%)
Frame = +2
Query: 360 HSPHVTVHESLLYSAWLRLPSGVDSNTTKVSEHVLYSEWIWLHLLTLLNEHDRGGGRFID 419
HSPHVTV+ESL+YSAWLRLP+ VDSNT K+ FI+
Sbjct: 2 HSPHVTVYESLVYSAWLRLPADVDSNTRKM---------------------------FIE 100
Query: 420 HAMLSAELTPLMTSL 434
M EL PL SL
Sbjct: 101 EVMELVELNPLRNSL 145
>TC82289 similar to GP|12382007|dbj|BAB21276. putative ABC transporter
protein {Oryza sativa (japonica cultivar-group)},
partial (25%)
Length = 1108
Score = 55.8 bits (133), Expect = 4e-08
Identities = 24/42 (57%), Positives = 30/42 (71%)
Frame = +3
Query: 331 IDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLY 372
I G+I +SG+PKKQETFARISGYCE ND H P + ++
Sbjct: 3 ICGNITISGFPKKQETFARISGYCEQNDRHWPDINAETRKMF 128
>BQ144083 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (7%)
Length = 467
Score = 52.4 bits (124), Expect = 5e-07
Identities = 54/161 (33%), Positives = 72/161 (44%), Gaps = 15/161 (9%)
Frame = +2
Query: 406 LLNEHDRGGGRFIDHAMLSAELTPLM--TSLTVFHFPSFAPLPPPFP-----AHLLHTS- 457
LL + RG R + ++ ++ L PL + + P F+ LP P A LLH S
Sbjct: 2 LLA*YRRGPLRSLLPSICASFLGPLALCNAFLFLYTPPFSLLPSSTPFFYLFARLLHFSF 181
Query: 458 LSHYPSGFSPYFTLLDPFF-FPSYPFTPSPLYSSFVFPPPFPMAVTRSPLPS------PA 510
L+H S +P+F PFF PS PF P S +FP FP+ PS P
Sbjct: 182 LAHKNS--TPFF--FPPFFPLPSLPFPSLPFSPSLLFPCSFPLPFLCLLCPSSLFPFFPL 349
Query: 511 ARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFS 551
P +FLPP P P FP LP S + ++FFS
Sbjct: 350 PPPSPPLFLPP-----PSPPFPLVSSFLPSLLSSAISLFFS 457
Score = 42.0 bits (97), Expect = 7e-04
Identities = 41/98 (41%), Positives = 48/98 (48%), Gaps = 7/98 (7%)
Frame = +3
Query: 462 PSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVT-----RSP-LPSPAARPPT 515
P F P F+ P PS PF P PL F+ P PF +V+ SP PSP P +
Sbjct: 201 PLSFFPPFSPSRPS--PSLPF-PFPLLFYFLVPFPFHFSVSYVLLLSSPSFPSPPPLPRS 371
Query: 516 AIFLPPITSSSPFPL-FPHSHKLLPLFPSPSPAVFFSP 552
PP S S FPL FPHS +P PSP +FSP
Sbjct: 372 FSPRPPRLSPS-FPLSFPHS------YPPPSP--YFSP 458
Score = 41.2 bits (95), Expect = 0.001
Identities = 33/90 (36%), Positives = 42/90 (46%), Gaps = 4/90 (4%)
Frame = +3
Query: 439 FPSFAPLPP----PFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFP 494
FP F+P P PFP LL L +P FS + LL PS+P +P PL SF
Sbjct: 213 FPPFSPSRPSPSLPFPFPLLFYFLVPFPFHFSVSYVLL--LSSPSFP-SPPPLPRSF--S 377
Query: 495 PPFPMAVTRSPLPSPAARPPTAIFLPPITS 524
P P PL P + PP + + P+ S
Sbjct: 378 PRPPRLSPSFPLSFPHSYPPPSPYFSPLPS 467
Score = 40.8 bits (94), Expect = 0.001
Identities = 34/90 (37%), Positives = 42/90 (45%), Gaps = 3/90 (3%)
Frame = +2
Query: 435 TVFHFPSFAPLPP-PFPAHLLHTSLSHYPSGFS-PYFTLLDPF-FFPSYPFTPSPLYSSF 491
T F FP F PLP PFP+ SL +P F P+ LL P FP +P P P F
Sbjct: 200 TPFFFPPFFPLPSLPFPSLPFSPSLL-FPCSFPLPFLCLLCPSSLFPFFPLPP-PSPPLF 373
Query: 492 VFPPPFPMAVTRSPLPSPAARPPTAIFLPP 521
+ PP P + S LPS + + F P
Sbjct: 374 LPPPSPPFPLVSSFLPSLLSSAISLFFSSP 463
Score = 40.8 bits (94), Expect = 0.001
Identities = 38/111 (34%), Positives = 46/111 (41%), Gaps = 3/111 (2%)
Frame = +1
Query: 454 LHTSLSHYP---SGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSPLPSPA 510
+H+SL P S F P FFF S F P F+F P FP V P PS
Sbjct: 103 VHSSLLSSPFVNSFFLPLCPSFAFFFFSS*EFHP------FLFSPLFPPPVPPLPFPSLF 264
Query: 511 ARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFFSPSCAVPLVLA 561
++FL P S SP F LPL P P SP+ + P + A
Sbjct: 265 PFSSISLFLSPSISLSPMSFFS-----LPLLSPPPP---LSPALSPPALPA 393
Score = 35.8 bits (81), Expect = 0.048
Identities = 45/130 (34%), Positives = 57/130 (43%), Gaps = 4/130 (3%)
Frame = +1
Query: 427 LTPLMTSLTVFHFPS--FAPLPPPFPAHLLHTSLSHYPSGFSPYFTL-LDPFFFPS-YPF 482
L P+ +SL F + F PL P F A +S +P FSP F + P FPS +PF
Sbjct: 94 LIPVHSSLLSSPFVNSFFLPLCPSF-AFFFFSS*EFHPFLFSPLFPPPVPPLPFPSLFPF 270
Query: 483 TPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFP 542
+ L+ S PM+ PL SP PP L P S P FP LFP
Sbjct: 271 SSISLFLSPSISLS-PMSFFSLPLLSPP--PP----LSPALSPPALPAFPPRF----LFP 417
Query: 543 SPSPAVFFSP 552
S +P + P
Sbjct: 418 SLTPILRHLP 447
>BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (19%)
Length = 1100
Score = 52.0 bits (123), Expect = 6e-07
Identities = 53/157 (33%), Positives = 59/157 (36%), Gaps = 46/157 (29%)
Frame = +3
Query: 440 PSFAPLPPP---FPAHLLHTSLSHYPSGFSPYFTL-LDP----FFFPSYPFTPSPLYSSF 491
P +P PPP P L S SH PS YF L L P FFFP P PL F
Sbjct: 195 PPPSPFPPPPFSSPPPLPPISSSHPPSLLLYYFILPLSPPTLSFFFPFSPPPSPPLILPF 374
Query: 492 V---------------FPPPFPMAVTR-----SPLPSPAARPPTAIFLPPITSSSPFPLF 531
+ FPP FP+ V SPLP PP+ + S FP F
Sbjct: 375 IYPISRSLLPLLLPPFFPPLFPLCVFPFPPFPSPLPFFRLSPPSPCPCSHLLSPFLFPTF 554
Query: 532 PH----------------SHKLLPL--FPSPSPAVFF 550
PH SH L P FP P P+ FF
Sbjct: 555 PHLRRPFSVS*PLLPLPRSHPLFPRRPFPLPPPSFFF 665
Score = 51.2 bits (121), Expect = 1e-06
Identities = 53/161 (32%), Positives = 59/161 (35%), Gaps = 42/161 (26%)
Frame = +3
Query: 439 FPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFF------------PSYPFTPSP 486
FP F P PPP P H +H P PY L+ P FF P PF P P
Sbjct: 54 FPPFPPPPPPIP-HNIHPPFPSPPPFPLPY--LIPPIFFLFLSL*RSFIPPPPSPFPPPP 224
Query: 487 LYSSFVFPPPFPMAVTRSP---------LP------------SPAARPPTAI-FLPPITS 524
S PPP P + P LP SP PP + F+ PI+
Sbjct: 225 FSS----PPPLPPISSSHPPSLLLYYFILPLSPPTLSFFFPFSPPPSPPLILPFIYPISR 392
Query: 525 S------SPF--PLFPHSHKLLPLFPSPSPAVFFSPSCAVP 557
S PF PLFP P FPSP P SP P
Sbjct: 393 SLLPLLLPPFFPPLFPLCVFPFPPFPSPLPFFRLSPPSPCP 515
Score = 51.2 bits (121), Expect = 1e-06
Identities = 49/138 (35%), Positives = 65/138 (46%), Gaps = 23/138 (16%)
Frame = +3
Query: 446 PPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPL----YSSFVFPPPFPMAV 501
PP FP LS +P P+F LL FFP +P P P+ + F PPPFP+
Sbjct: 3 PPLFP-------LSFFP----PFFPLL---FFPPFPPPPPPIPHNIHPPFPSPPPFPLPY 140
Query: 502 TRSPL---------------PSPAARPP--TAIFLPPITSSSPFPLFPHSHKLLPLFPSP 544
P+ PSP PP + LPPI+SS P L + + +LPL P P
Sbjct: 141 LIPPIFFLFLSL*RSFIPPPPSPFPPPPFSSPPPLPPISSSHPPSLLLY-YFILPLSP-P 314
Query: 545 SPAVF--FSPSCAVPLVL 560
+ + F FSP + PL+L
Sbjct: 315 TLSFFFPFSPPPSPPLIL 368
Score = 50.4 bits (119), Expect = 2e-06
Identities = 51/159 (32%), Positives = 63/159 (39%), Gaps = 22/159 (13%)
Frame = +2
Query: 427 LTPLMTSLTVFHFPSFAP-------LPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPS 479
L P SL + + P P LPPP P H L +PS L P F
Sbjct: 155 LFPFPFSLKIIYPPPPLPIPSPPLLLPPPPPPHFL------FPS------PLPSPILFHP 298
Query: 480 YPFTPSPLYSSFVFPPPF-----PMAVTRSPLPSPAARPPTAIFLPPITSSSPFPL---- 530
PF PL+ FPPP P+ ++ PLPS + P++ FL P S FPL
Sbjct: 299 SPFPSHPLFFFSFFPPPLPPPYTPLHISHFPLPSSS---PSSPFLSPPFPSLRFPLSSFS 469
Query: 531 FPHSHKL------LPLFPSPSPAVFFSPSCAVPLVLAVL 563
FP S LPLFP P S P + +L
Sbjct: 470 FPSSFLSLVPPLPLPLFPFTLPLSLPHLSSFAPSIFRLL 586
Score = 48.5 bits (114), Expect = 7e-06
Identities = 41/126 (32%), Positives = 53/126 (41%), Gaps = 6/126 (4%)
Frame = +2
Query: 427 LTPLMTSLTVFHFPSFAPLPPPFPAHLL-HTSLSHYPSGFSPYFTLLDPFFFPSYPFTPS 485
L+PL+ F P F+ LPP P H H S P SP + +P + F P
Sbjct: 11 LSPLLLP-PFFPSPLFSSLPPSSPPHSP*HPSPLPLPPSLSPTLS------YPPHLF-PF 166
Query: 486 PLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITS-----SSPFPLFPHSHKLLPL 540
P ++PPP P+ + PL P PP +F P+ S SPFP H
Sbjct: 167 PFSLKIIYPPP-PLPIPSPPLLLPPPPPPHFLFPSPLPSPILFHPSPFP--SHPLFFFSF 337
Query: 541 FPSPSP 546
FP P P
Sbjct: 338 FPPPLP 355
Score = 47.4 bits (111), Expect = 2e-05
Identities = 43/124 (34%), Positives = 50/124 (39%), Gaps = 31/124 (25%)
Frame = +1
Query: 461 YPSGFSPYFTLLDPFFFPS------YPFT---PSPLYSSFVFP----------------- 494
+PS SP F+L FF PS +P T PSP F +P
Sbjct: 13 FPSPSSPLFSL-SSFFLPSPLLPPPFPITSIPPSPPPLPFPYPILSPPSFSFSFLFKDHL 189
Query: 495 -PPFPMAVTRSPLPSPAARPPTAIFLPP----ITSSSPFPLFPHSHKLLPLFPSPSPAVF 549
PP P P P P PP + +PP I SS PFPL P LFP P P
Sbjct: 190 SPPPPPHSLPPPSPPPPPSPPFPLPIPPPFSYIISSFPFPLPP--SLFFFLFPPPPPPPL 363
Query: 550 FSPS 553
+SPS
Sbjct: 364 YSPS 375
Score = 45.1 bits (105), Expect = 8e-05
Identities = 41/124 (33%), Positives = 53/124 (42%), Gaps = 15/124 (12%)
Frame = +2
Query: 427 LTPLMTSLTVFHFP--SFAP----LPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSY 480
L P T L + HFP S +P L PPFP+ S +PS F +L+ P P +
Sbjct: 350 LPPPYTPLHISHFPLPSSSPSSPFLSPPFPSLRFPLSSFSFPSSF---LSLVPPLPLPLF 520
Query: 481 PFT---PSPLYSSFVFPPPF------PMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLF 531
PFT P SSF P F + SPLP P PP +F P +F
Sbjct: 521 PFTLPLSLPHLSSFA-PSIFRLLTLTSLTTIPSPLPPPPLPPPPPLFFFCFRRPPP*CVF 697
Query: 532 PHSH 535
P ++
Sbjct: 698 PKNY 709
Score = 42.4 bits (98), Expect = 5e-04
Identities = 45/140 (32%), Positives = 51/140 (36%), Gaps = 18/140 (12%)
Frame = +1
Query: 429 PLMTSLTVFHFPSFAPLPP-------PFPAHLLHTSLSHYPSGFSPY-------FTLLDP 474
PL SL F FP P PP PFPA LS +P F P+ F L P
Sbjct: 307 PLPPSLFFFLFPP-PPPPPLYSPSYIPFPAPFF---LSFFPLSFPPFSLFAFSPFLLFLP 474
Query: 475 FFF----PSYPFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPL 530
F P P P P+YS PPF + P + P + PI SS
Sbjct: 475 LFLSFACPPPPLAPVPIYSPPFSSPPFLICAVHFP-----SLDPYFPYHDPIPSS----- 624
Query: 531 FPHSHKLLPLFPSPSPAVFF 550
P PSPSP F
Sbjct: 625 --------PAAPSPSPPPLF 660
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 49.7 bits (117), Expect = 3e-06
Identities = 40/104 (38%), Positives = 45/104 (42%)
Frame = -2
Query: 440 PSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPM 499
PS P PPP P L+ SL+ P SP TL P F S PF P P PPP
Sbjct: 415 PSTFPPPPPCPIPSLYFSLAA-PLRSSPSLTL-PPSFCSSPPFLPPP-------PPP--- 272
Query: 500 AVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPS 543
P P P PP++ PP+ S P P P L P PS
Sbjct: 271 -----PSPPPPPPPPSSPLRPPLRSPPPLPPLP---PLPPFTPS 164
Score = 46.6 bits (109), Expect = 3e-05
Identities = 45/136 (33%), Positives = 60/136 (44%), Gaps = 21/136 (15%)
Frame = -1
Query: 436 VFHFPSFAPLPPPF----PAHLLHTSLSHYPSGFSPYFTLLD--PFFFPSY--PFTPSPL 487
++H P + P P ++ SLS +P F P +L P FFP PF P
Sbjct: 506 LYHLPIYYASIPAMMTLRPFVVIRFSLSLFPLHFPPASSLSHSFPLFFPCRPPPFLPLSH 327
Query: 488 YSSF-VFPPPFPMAVTRSPLPSPAARPPTAIF-----------LPPITSSSPF-PLFPHS 534
S F +F PP SPLPSP PP+++F PP +SPF P F
Sbjct: 326 ASPFLLFLPPLSSPSPSSPLPSPP--PPSSLFPSPTPPPLPPPPPPPPPASPFHPFFSSY 153
Query: 535 HKLLPLFPSPSPAVFF 550
H L P F P+++F
Sbjct: 152 HFLSPSF---LPSLYF 114
Score = 45.8 bits (107), Expect = 5e-05
Identities = 41/122 (33%), Positives = 53/122 (42%), Gaps = 15/122 (12%)
Frame = -1
Query: 440 PSFAPLPPPFPA-----HLLHTSLSHYPSGFS--PYFTLLDPFFFPSYPFTPSPLYSSFV 492
P F PLP P A H L +L H P ++ P L PF + + PL+
Sbjct: 575 PPFLPLPRPRAALILLIHHLVCALYHLPIYYASIPAMMTLRPFVVIRFSLSLFPLH---- 408
Query: 493 FPPP------FPMAVTRSPLPS-PAARP-PTAIFLPPITSSSPFPLFPHSHKLLPLFPSP 544
FPP FP+ P P P + P +FLPP++S SP P LFPSP
Sbjct: 407 FPPASSLSHSFPLFFPCRPPPFLPLSHASPFLLFLPPLSSPSPSSPLPSPPPPSSLFPSP 228
Query: 545 SP 546
+P
Sbjct: 227 TP 222
Score = 40.8 bits (94), Expect = 0.001
Identities = 48/130 (36%), Positives = 54/130 (40%), Gaps = 3/130 (2%)
Frame = -3
Query: 434 LTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVF 493
L V PS PLPPP + LS +P F P P FFP P P PL
Sbjct: 393 LPVPFLPSIFPLPPPS----VPPPLSRFPLPFVP-----PPPFFP-LPLLPPPL------ 262
Query: 494 PPPFPMAVTRSPLP---SPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSPSPAVFF 550
PP P+ PLP P+A P PP S FPL P LL + P P P +
Sbjct: 261 -PPPPLL----PLPLSDPPSAPP------PPSPPSPRFPLSP----LLLILPLPLPLL-- 133
Query: 551 SPSCAVPLVL 560
PS V L L
Sbjct: 132 -PSFVVFLPL 106
Score = 37.0 bits (84), Expect = 0.021
Identities = 43/136 (31%), Positives = 56/136 (40%), Gaps = 16/136 (11%)
Frame = -2
Query: 443 APLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTPSPLYSSFVFPPPFPMAVT 502
+P P A LL S S P P L PF PS+ L S FPP FP +
Sbjct: 1219 SPSPRLHLASLLSLS-SSTPRSSPP--RLAPPFSSPSFFLFCYLLSRSSPFPPAFPF*LP 1049
Query: 503 RSPLPS---PAARPP-------TAIFLPPITSSS---PFPLFPHSHKLLPLFPSPSPAVF 549
P P+ P A PP T + P+ S+S P P L P PS +
Sbjct: 1048 LLPPPALSRPLAAPPLSHLALLTLLLFAPLDSASCLAPSCAVPRRRALAPSLLLPSRSSS 869
Query: 550 FSP---SCAVPLVLAV 562
SP S A+PL++++
Sbjct: 868 LSPSHSSPALPLMISI 821
Score = 36.6 bits (83), Expect = 0.028
Identities = 32/83 (38%), Positives = 40/83 (47%)
Frame = -3
Query: 479 SYPFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLL 538
S PF P PL S + P PF ++ PLP P+ PP + F P+ P P FP L
Sbjct: 429 SLPFPP-PL-SPRLLPVPFLPSIF--PLPPPSVPPPLSRF--PLPFVPPPPFFP-----L 283
Query: 539 PLFPSPSPAVFFSPSCAVPLVLA 561
PL P P P P +PL L+
Sbjct: 282 PLLPPPLP-----PPPLLPLPLS 229
Score = 31.6 bits (70), Expect = 0.90
Identities = 26/77 (33%), Positives = 33/77 (42%)
Frame = -2
Query: 481 PFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPL 540
PF+PS +F PPP P+ L +P P+ + LPP SSP L P P
Sbjct: 424 PFSPS----TFPPPPPCPIPSLYFSLAAPLRSSPS-LTLPPSFCSSPPFLPPPPPPPSPP 260
Query: 541 FPSPSPAVFFSPSCAVP 557
P P P+ P P
Sbjct: 259 PPPPPPSSPLRPPLRSP 209
>BQ135566
Length = 930
Score = 48.9 bits (115), Expect = 5e-06
Identities = 51/139 (36%), Positives = 68/139 (48%), Gaps = 20/139 (14%)
Frame = +1
Query: 430 LMTSLTVFH-FPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFP----SY---- 480
++ +LT+F FP + P P+ LL +PS F P PFFFP SY
Sbjct: 199 IIQNLTLFQLFPIYLYHQTPPPSILLI-----FPSHFFPL-----PFFFPYSQTSYSPNT 348
Query: 481 PF-----TPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLP-PITSS--SPFPLFP 532
PF TP+P ++ PPPFP ++ S LP P PP LP P+ SS S FPL+
Sbjct: 349 PFPPKHHTPNPHFT----PPPFPFSILLS-LPIPTTSPP----LPHPLLSSLISLFPLYN 501
Query: 533 HSH---KLLPLFPSPSPAV 548
SH PL+ +P P +
Sbjct: 502 ISHIHTYTFPLYSAPPPVI 558
Score = 40.0 bits (92), Expect = 0.003
Identities = 41/126 (32%), Positives = 52/126 (40%), Gaps = 17/126 (13%)
Frame = +2
Query: 436 VFHFPSFAPLPPPFPAHLLHTSLSHY-------PSGFSPYFTLLDPFFFPSYPFTPS--P 486
+F P F P P FP H L+H+ P+ SP L PF +PS PF P P
Sbjct: 275 LFFRPIFFPSPFFFPIPKPHIPLTHHSHPNTTPPTPISPLPHSLFPFCYPS-PFQPHLLP 451
Query: 487 LYSSFVFPPPFPMAVTRSPLP----SP-AARPP---TAIFLPPITSSSPFPLFPHSHKLL 538
+ F P T SP+ SP A PP LPPI P+ + H +
Sbjct: 452 YHIPFYHPLSHSSHYTTSPISIHIHSPYTAHPPQLSPRSHLPPI-----IPVIKYKHPVT 616
Query: 539 PLFPSP 544
+ PSP
Sbjct: 617 HVNPSP 634
>BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 1047
Score = 48.1 bits (113), Expect = 9e-06
Identities = 49/138 (35%), Positives = 63/138 (45%), Gaps = 14/138 (10%)
Frame = +2
Query: 429 PLMTSLTVFHFPSFAPLPPPFPAHL-----LHTSLSHYPSGFSPYFTLLDPFFFPSYPFT 483
P +T L H P PP PA+ H +YP+ ++ F FP
Sbjct: 107 PWIT*LPPRHAPEV*SFKPPPPAYS*SPNPPHNKQKNYPAQYT--HPTHSHFIFP----L 268
Query: 484 PSPLYSS----FVFPPPFPMAVTRSPLPSPAARPPTA---IFLPPIT-SSSPFPL-FPHS 534
PSP Y S F+F P FP+++ PLP P+ RPP FL + S+PF L FP
Sbjct: 269 PSPSYLSSVVRFLFSPLFPLSI---PLPPPSYRPPPPSRFFFLSLLLFFSTPFALVFP-- 433
Query: 535 HKLLPLFPSPSPAVFFSP 552
L+ LFP SP FSP
Sbjct: 434 --LILLFPHYSPPPPFSP 481
Score = 43.5 bits (101), Expect = 2e-04
Identities = 49/145 (33%), Positives = 58/145 (39%), Gaps = 23/145 (15%)
Frame = +2
Query: 439 FPSFAPLPPPF-----PAHLLHTSLSHYPSGFSPYFTLLDPFF--FPSYPFTPSPLYSSF 491
FP PLPPP P+ SL + FS F L+ P FP Y SP
Sbjct: 320 FPLSIPLPPPSYRPPPPSRFFFLSLLLF---FSTPFALVFPLILLFPHY----SP----- 463
Query: 492 VFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFP----------HSH------ 535
PPPF R P P + P IFLPP SSPF LF HS
Sbjct: 464 --PPPFSPIFPRFPRPH-SFPPHFTIFLPP-PISSPFFLFSRISPLSISPIHSSIFFLHI 631
Query: 536 KLLPLFPSPSPAVFFSPSCAVPLVL 560
L P FP+P FF + +P ++
Sbjct: 632 PLFPFFPTPPSYPFFLLTPFIPFLM 706
Score = 42.0 bits (97), Expect = 7e-04
Identities = 50/139 (35%), Positives = 58/139 (40%), Gaps = 9/139 (6%)
Frame = +1
Query: 427 LTPLMTSLTVFHFPSF----APLPPPFPAHLLHTSLSHYPSGFSPYFTLLD--PFFFPSY 480
++PL+ TV SF +PL PP P L H S P FSP F P F P +
Sbjct: 388 VSPLIFFHTVCPCFSFNFIISPLLPP-PTLLSHISPFPPPPFFSPPFHHFSSSPHFLPFF 564
Query: 481 PFTPSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLP--PI-TSSSPFPLFPHSHKL 537
F P S F FP PF + P S + PP F P PI T S F P
Sbjct: 565 SFFP--YLSPFHFPHPFLHLFSSYPFVSIFSYPPLLSFFPSNPIHTISYDFIFLPFPVSR 738
Query: 538 LPLFPSPSPAVFFSPSCAV 556
L LF P F PS +V
Sbjct: 739 LVLFVDIFPISLF-PSTSV 792
Score = 42.0 bits (97), Expect = 7e-04
Identities = 44/149 (29%), Positives = 57/149 (37%), Gaps = 30/149 (20%)
Frame = +3
Query: 436 VFHFPSFAP--LPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFF---------FPSYPFTP 484
++ FP P PPP P S +P P F L +F P +P +P
Sbjct: 324 LYQFPCPPPRTAPPPLPVFSFCLSSYFFPHRL-PLFFL*FYYFPITPPPHPSLPYFPVSP 500
Query: 485 SPLY----SSFVFPPPFPMAV----TRSPLPSPAARPPT--------AIFLPPITSSSPF 528
+P+ S F F PPFP P P P + PP+ FLPP P
Sbjct: 501 APILFPPISPFFFLPPFPPLFFFFPVSLPFPFPPSIPPSFFFISLCFHFFLPP-----PL 665
Query: 529 PLF---PHSHKLLPLFPSPSPAVFFSPSC 554
LF PHS+ L + S P + P C
Sbjct: 666 ILFSF*PHSYHFL*FYLSAFPRLPSRPVC 752
Score = 37.7 bits (86), Expect = 0.013
Identities = 35/121 (28%), Positives = 52/121 (42%), Gaps = 2/121 (1%)
Frame = +2
Query: 427 LTPLMTSLTVFHFP--SFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPFFFPSYPFTP 484
++P+ +S+ H P F P PP +P LL + F P+ +L S+ +P
Sbjct: 593 ISPIHSSIFFLHIPLFPFFPTPPSYPFFLL--------TPFIPFLMIL------SFCLSP 730
Query: 485 SPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFPSP 544
SP+ S FP +R PL S +A L P +S SP + +P P P
Sbjct: 731 SPVSSCLSI--SFPSLFSRPPLSSLSAISAPRTLL-PFSSISPL------SRRIPSLPHP 883
Query: 545 S 545
S
Sbjct: 884 S 886
Score = 37.0 bits (84), Expect = 0.021
Identities = 42/132 (31%), Positives = 48/132 (35%), Gaps = 25/132 (18%)
Frame = +1
Query: 446 PPPFPAHLLHTSLSHYPSGFSPYFTLL--DPFFFPSYPFTPSPLYSSFVFPPPFPMAVTR 503
P PFP H +P F P+ + L F F + F S PPPFP +
Sbjct: 238 PNPFPFH--------FPPPF-PFLSFLRRSVFVFAPFSFINSLAPPLVPPPPPFPFFLFV 390
Query: 504 SPL---------------PSPAARPPTAI-----FLPPITSSSPFPLFPHSHKLLP---L 540
SPL SP PPT + F PP S PF F S LP
Sbjct: 391 SPLIFFHTVCPCFSFNFIISPLLPPPTLLSHISPFPPPPFFSPPFHHFSSSPHFLPFFSF 570
Query: 541 FPSPSPAVFFSP 552
FP SP F P
Sbjct: 571 FPYLSPFHFPHP 606
>TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1171
Score = 47.8 bits (112), Expect = 1e-05
Identities = 41/115 (35%), Positives = 53/115 (45%), Gaps = 3/115 (2%)
Frame = +2
Query: 415 GRFIDHAMLSAELTPLMTSLTVFHFPSFAP--LPPPFPAHLLHTSLSHYPSGFSPYFTLL 472
G + +L L+P S + SF+P L PP P T+ S PS FS L
Sbjct: 317 GLHLKQPLLLIYLSPYHFSYCLSSSCSFSPSAL*PPSPT-ASSTASSAVPSNFSLQSPLS 493
Query: 473 DPFFFPSYPFTPSPLYSSFVFPPPFPMAVTRSP-LPSPAARPPTAIFLPPITSSS 526
P FF S P SP SS P P P + + SP L S A PP++ P +SS+
Sbjct: 494 SPPFFLSLPQPSSPTSSSSPSPSPSPSSASPSPSLKSFALSPPSSFLHSPFSSST 658
Score = 29.3 bits (64), Expect = 4.5
Identities = 31/88 (35%), Positives = 38/88 (42%), Gaps = 2/88 (2%)
Frame = +2
Query: 472 LDPFFFPSYPFTPSPLYSSFVFPPPFPMA--VTRSPLPSPAARPPTAIFLPPITSSSPFP 529
L P+ F SY + S +S PP P A S +PS + + + PP S P P
Sbjct: 353 LSPYHF-SYCLSSSCSFSPSAL*PPSPTASSTASSAVPSNFSLQ-SPLSSPPFFLSLPQP 526
Query: 530 LFPHSHKLLPLFPSPSPAVFFSPSCAVP 557
P S PSPSP SPS A P
Sbjct: 527 SSPTSSS----SPSPSP----SPSSASP 586
>AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (10%)
Length = 478
Score = 47.4 bits (111), Expect = 2e-05
Identities = 48/141 (34%), Positives = 62/141 (43%), Gaps = 7/141 (4%)
Frame = +1
Query: 430 LMTSLTVFHFPSFAPLPPPFPAHLLHTSLSHYPSGFSPYFTLLDPF--FFPSYPFT---- 483
+ T+ F+FP F LPPP P+H L+ PS F P PF F P P +
Sbjct: 22 MATNQPNFYFP-FPHLPPP-PSHPLNPP----PSPFHPINPPPSPFHPFNPPPPHSIKSP 183
Query: 484 -PSPLYSSFVFPPPFPMAVTRSPLPSPAARPPTAIFLPPITSSSPFPLFPHSHKLLPLFP 542
P+P +SS PPP P P PS PP PP P P P + + P
Sbjct: 184 PPAPSHSSP--PPPPPPRKXPPPPPSHPFSPP-----PPHNHPPPSPHHPIAPSPPHVRP 342
Query: 543 SPSPAVFFSPSCAVPLVLAVL 563
SP P + SPS P V+ ++
Sbjct: 343 SPPPPLPPSPSPYHPTVIVIV 405
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,526,716
Number of Sequences: 36976
Number of extensions: 486378
Number of successful extensions: 7861
Number of sequences better than 10.0: 469
Number of HSP's better than 10.0 without gapping: 5659
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6695
length of query: 596
length of database: 9,014,727
effective HSP length: 102
effective length of query: 494
effective length of database: 5,243,175
effective search space: 2590128450
effective search space used: 2590128450
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126018.1


