

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
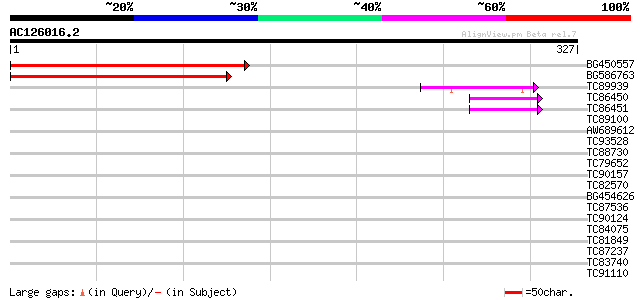
Sequences producing significant alignments: (bits) Value
BG450557 similar to GP|21618279|gb| unknown {Arabidopsis thalian... 270 7e-73
BG586763 weakly similar to GP|21618279|gb| unknown {Arabidopsis ... 241 2e-64
TC89939 similar to GP|15289911|dbj|BAB63606. hypothetical protei... 42 4e-04
TC86450 similar to GP|21537318|gb|AAM61659.1 unknown {Arabidopsi... 41 6e-04
TC86451 similar to GP|21537318|gb|AAM61659.1 unknown {Arabidopsi... 41 6e-04
TC89100 similar to GP|21553849|gb|AAM62942.1 putative RING zinc ... 40 0.001
AW689612 weakly similar to GP|19698963|gb| unknown protein {Arab... 39 0.004
TC93528 similar to GP|20466490|gb|AAM20562.1 unknown protein {Ar... 39 0.004
TC88730 weakly similar to GP|21592748|gb|AAM64697.1 PRT1 {Arabid... 38 0.006
TC79652 weakly similar to GP|19698951|gb|AAL91211.1 unknown prot... 37 0.008
TC90157 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1... 37 0.014
TC82570 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1... 37 0.014
BG454626 similar to GP|16323398|gb unknown protein {Arabidopsis ... 36 0.018
TC87536 similar to GP|21553686|gb|AAM62779.1 unknown {Arabidopsi... 35 0.040
TC90124 similar to GP|17529304|gb|AAL38879.1 putative transcript... 35 0.052
TC84075 similar to GP|3738297|gb|AAC63639.1| unknown protein {Ar... 35 0.052
TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 34 0.068
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 34 0.089
TC83740 similar to GP|6437560|gb|AAF08587.1| hypothetical protei... 34 0.089
TC91110 similar to GP|7576941|gb|AAF64065.1| putative diacylglyc... 34 0.089
>BG450557 similar to GP|21618279|gb| unknown {Arabidopsis thaliana}, partial
(13%)
Length = 664
Score = 270 bits (689), Expect = 7e-73
Identities = 137/138 (99%), Positives = 137/138 (99%)
Frame = +3
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 60
MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA
Sbjct: 249 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 428
Query: 61 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA
Sbjct: 429 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 608
Query: 121 IRERALKQATESLKGKST 138
IRERALKQAT SLKGKST
Sbjct: 609 IRERALKQATXSLKGKST 662
>BG586763 weakly similar to GP|21618279|gb| unknown {Arabidopsis thaliana},
partial (18%)
Length = 678
Score = 241 bits (616), Expect = 2e-64
Identities = 123/128 (96%), Positives = 124/128 (96%)
Frame = +3
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 60
MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA
Sbjct: 288 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 467
Query: 61 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLET FSRDARA
Sbjct: 468 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETGD*FSRDARA 647
Query: 121 IRERALKQ 128
IRERAL +
Sbjct: 648 IRERALNK 671
>TC89939 similar to GP|15289911|dbj|BAB63606. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 F18O14.3, partial
(23%)
Length = 978
Score = 41.6 bits (96), Expect = 4e-04
Identities = 23/73 (31%), Positives = 36/73 (48%), Gaps = 5/73 (6%)
Frame = +2
Query: 238 KMRLATGEDAEEEGAS--LNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALK- 294
K ++ GE S + +++ DA F C IC + DP+ T C H FC C K
Sbjct: 50 KWQVGLGESTSRSPPSPPYSSNNNNSDAGNFECNICFDLAQDPIITLCGHLFCWPCLYKW 229
Query: 295 --HHAKNKKCFVC 305
H+++++C VC
Sbjct: 230 LHFHSQSRECPVC 268
>TC86450 similar to GP|21537318|gb|AAM61659.1 unknown {Arabidopsis
thaliana}, partial (39%)
Length = 1268
Score = 41.2 bits (95), Expect = 6e-04
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = +3
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
F C IC P V+ +ST+C H FC+ C + KC C +
Sbjct: 735 FTCPICMGPMVEEMSTRCGHIFCKSCIKAAISAQAKCPTCRK 860
>TC86451 similar to GP|21537318|gb|AAM61659.1 unknown {Arabidopsis
thaliana}, partial (51%)
Length = 995
Score = 41.2 bits (95), Expect = 6e-04
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = +2
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQ 307
F C IC P V+ +ST+C H FC+ C + KC C +
Sbjct: 647 FTCPICMGPMVEEMSTRCGHIFCKSCIKAAISAQAKCPTCRK 772
>TC89100 similar to GP|21553849|gb|AAM62942.1 putative RING zinc finger
protein {Arabidopsis thaliana}, partial (44%)
Length = 1047
Score = 40.0 bits (92), Expect = 0.001
Identities = 27/84 (32%), Positives = 39/84 (46%), Gaps = 3/84 (3%)
Frame = +1
Query: 225 QMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCK 284
+M + + E+ K+ A+ +S ND D D F C IC + DPV T C
Sbjct: 250 EMSRRFGESTKSAS---ASNPSCSSGNSSNNDPGDVGD---FECNICFDLAQDPVITLCG 411
Query: 285 HYFCEHCA---LKHHAKNKKCFVC 305
H FC C L HH+ +++ VC
Sbjct: 412 HLFCWPCLYRWLHHHSHSQEFPVC 483
>AW689612 weakly similar to GP|19698963|gb| unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 653
Score = 38.5 bits (88), Expect = 0.004
Identities = 16/44 (36%), Positives = 20/44 (45%)
Frame = +1
Query: 263 ALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCN 306
AL C +C N F PV C H FC C + +C +CN
Sbjct: 115 ALELKCPLCLNLFKKPVLLPCNHLFCSSCLADSTSIRSECALCN 246
>TC93528 similar to GP|20466490|gb|AAM20562.1 unknown protein {Arabidopsis
thaliana}, partial (24%)
Length = 755
Score = 38.5 bits (88), Expect = 0.004
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = +3
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVC 305
C IC+ P+ +CKH FCE C + + + C +C
Sbjct: 168 CAICQEKMHSPILLRCKHIFCEDCVSEWFERERTCPLC 281
>TC88730 weakly similar to GP|21592748|gb|AAM64697.1 PRT1 {Arabidopsis
thaliana}, partial (28%)
Length = 802
Score = 37.7 bits (86), Expect = 0.006
Identities = 34/153 (22%), Positives = 60/153 (38%), Gaps = 4/153 (2%)
Frame = +2
Query: 160 RREQTIASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKETGY-CGYGDSC--KFMHD 216
R QT+ EK G + P +FD+ D C+ + G+ C S + +
Sbjct: 347 RTNQTLEEEKKSGYYSP-----------QFDF--DTCESQAKFGHSCSPSSSSTINLVSN 487
Query: 217 RGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALP-FACFICRNPF 275
+ + M++ + + TG EE+ L++ + ++ C +C+
Sbjct: 488 SSNVGTSECMDQPGSTSHEGEPE--ITGTRVEEKALPLDNLTQQKISVADVMCTMCKQLL 661
Query: 276 VDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQP 308
PV C H +CE C K + +C VC P
Sbjct: 662 FHPVVLHCGHVYCETCVYKLADEMLRCQVCQIP 760
Score = 32.7 bits (73), Expect = 0.20
Identities = 16/54 (29%), Positives = 24/54 (43%), Gaps = 4/54 (7%)
Frame = +2
Query: 256 DEDDDEDALP--FACFICRNPFVDPVSTKCKHYFCEHCALKHHA--KNKKCFVC 305
D D+ E+ +P F C +C + P+ C H C C K + + KC C
Sbjct: 101 DHDEHEEEIPDSFCCCVCLDLLYKPIVLSCGHMCCFWCIHKSMSGVRESKCPTC 262
>TC79652 weakly similar to GP|19698951|gb|AAL91211.1 unknown protein
{Arabidopsis thaliana}, partial (20%)
Length = 1590
Score = 37.4 bits (85), Expect = 0.008
Identities = 24/82 (29%), Positives = 38/82 (46%), Gaps = 1/82 (1%)
Frame = +2
Query: 225 QMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCK 284
++EK+ + A + A ED+ E + + D + C ICR+ + V TKC
Sbjct: 632 RVEKDLEVARRNFSHLKAQDEDSSETDKLQQELGEYRDIVK--CSICRDRTKEVVITKCY 805
Query: 285 HYFCEHCALK-HHAKNKKCFVC 305
H FC C K ++ +KC C
Sbjct: 806 HLFCNSCIQKIAGSRQRKCPQC 871
>TC90157 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1a
[imported] - Arabidopsis thaliana (fragment), partial
(44%)
Length = 1486
Score = 36.6 bits (83), Expect = 0.014
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Frame = +1
Query: 260 DEDALPFACFICRNPFV--DPVS-TKCKHYFCEHCALKHHAKNKKCFVCNQ 307
D+D+ +C IC F DP + T CKH + HC L+ ++K+C +C Q
Sbjct: 160 DDDSFEDSCSICLESFSVHDPSTVTCCKHEYHLHCILEWSQRSKECPICWQ 312
>TC82570 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1a
[imported] - Arabidopsis thaliana (fragment), partial
(23%)
Length = 738
Score = 36.6 bits (83), Expect = 0.014
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Frame = +2
Query: 260 DEDALPFACFICRNPFV--DPVS-TKCKHYFCEHCALKHHAKNKKCFVCNQ 307
D+D+ +C IC F DP + T CKH + HC L+ ++K+C +C Q
Sbjct: 188 DDDSFEDSCSICLESFSVHDPSTVTCCKHEYHLHCILEWSQRSKECPICWQ 340
>BG454626 similar to GP|16323398|gb unknown protein {Arabidopsis thaliana},
partial (72%)
Length = 689
Score = 36.2 bits (82), Expect = 0.018
Identities = 18/75 (24%), Positives = 35/75 (46%), Gaps = 7/75 (9%)
Frame = +1
Query: 241 LATGEDAEEEGASLNDEDDDEDALPF--ACFICRNPFVDPVSTKCKHYFCEHCALKHHAK 298
L++ ++ + ++N+E + + P C IC F P C H+FC +C ++
Sbjct: 91 LSSQSQSQSQN*TMNEEIESGEGPPTNDLCSICHGNFQLPCQANCSHWFCANCIIQVWQY 270
Query: 299 NK-----KCFVCNQP 308
+ KC +C +P
Sbjct: 271 SSPLQPCKCPLCRRP 315
>TC87536 similar to GP|21553686|gb|AAM62779.1 unknown {Arabidopsis
thaliana}, partial (94%)
Length = 1358
Score = 35.0 bits (79), Expect = 0.040
Identities = 22/87 (25%), Positives = 42/87 (47%), Gaps = 2/87 (2%)
Frame = +3
Query: 243 TGEDAEEEGASLNDEDDDEDAL-PF-ACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNK 300
T ++ + G E +D++ PF AC +C +DP+S + H FC+ C L
Sbjct: 183 TYDEKRKLGYGTQKERLGKDSIKPFDACSLCLKSLIDPMSCQKGHLFCKECIL------- 341
Query: 301 KCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C + + + VAH ++K +++
Sbjct: 342 ECLLSQKKDIQRRLVAHSAQQKQEKEE 422
Score = 33.5 bits (75), Expect = 0.12
Identities = 14/46 (30%), Positives = 24/46 (51%), Gaps = 4/46 (8%)
Frame = +3
Query: 266 FACFICRNPFVDPVS----TKCKHYFCEHCALKHHAKNKKCFVCNQ 307
+ C C+ + +S + C H FC+ C+ + A +K C VCN+
Sbjct: 771 YICPSCKTTLTNTMSLVALSSCGHVFCKKCSDRFMAVDKVCLVCNK 908
>TC90124 similar to GP|17529304|gb|AAL38879.1 putative transcription factor
{Arabidopsis thaliana}, partial (34%)
Length = 1315
Score = 34.7 bits (78), Expect = 0.052
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = +2
Query: 266 FACFICRNPFVDPVSTKCKHYFCEHC 291
F C ICR P++T C H FC+ C
Sbjct: 548 FGCNICRKGLASPLTTPCAHNFCKAC 625
>TC84075 similar to GP|3738297|gb|AAC63639.1| unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 615
Score = 34.7 bits (78), Expect = 0.052
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +3
Query: 196 CKDYKETGYCGYGDSCKFMHDR 217
C Y TG+CGYG C+F H R
Sbjct: 351 CAYYMRTGFCGYGGRCRFNHPR 416
Score = 33.1 bits (74), Expect = 0.15
Identities = 21/56 (37%), Positives = 28/56 (49%), Gaps = 6/56 (10%)
Frame = +3
Query: 173 SHGPLRASAHIRVSARFDYQPDI----CKDYKETGYCGYGDSCKFMHDR--GDYKS 222
+H RA+ V A DY + C+ Y +TG C +G SCKF H + G Y S
Sbjct: 405 NHPRDRAAVAAAVRATGDYPERLGEPPCQYYLKTGTCKFGASCKFHHPKNGGGYLS 572
>TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (16%)
Length = 1477
Score = 34.3 bits (77), Expect = 0.068
Identities = 23/88 (26%), Positives = 39/88 (44%), Gaps = 3/88 (3%)
Frame = +1
Query: 221 KSGWQMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVS 280
+ G + W + + + + A G + + G + ++ F C IC + DPV
Sbjct: 34 EEGIVVSVNWGKRKSSHLIAKALGMEEIDNG-----KVEESSGNFFDCNICLDIARDPVL 198
Query: 281 TKCKHYFCEHCALK---HHAKNKKCFVC 305
T C H FC C + ++K K+C VC
Sbjct: 199 TCCGHLFCWPCFYQLSYAYSKAKECPVC 282
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 33.9 bits (76), Expect = 0.089
Identities = 35/151 (23%), Positives = 61/151 (40%), Gaps = 1/151 (0%)
Frame = +1
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKAD 61
+D E EN++ + ++ K+ K ENE+N+++ +++ + KKN +
Sbjct: 142 KDEEKSKQENEEIKDGEKIQQENEENKDEEKSQQENEENKDEEKSQQE--NELKKNEGGE 315
Query: 62 NKLFFSTGSSKSSASAKPNEE-SEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
+ TG S + NEE SE S E+ + Q D+K E D +
Sbjct: 316 KE----TGEITEEKSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEE 483
Query: 121 IRERALKQATESLKGKSTSSEDVKLYKGINN 151
+ E K K +S D ++ G N
Sbjct: 484 TNGTEGGEKEEHDKIKEDTSSDNQVQDGEKN 576
Score = 32.3 bits (72), Expect = 0.26
Identities = 50/242 (20%), Positives = 95/242 (38%), Gaps = 8/242 (3%)
Frame = +1
Query: 2 EDSEAKS-TENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 60
E SE KS EN+++ K + K +ENE+N+++ +++ + + K
Sbjct: 46 ESSEEKSHLENEES--------KETGESSEEKSHLENEENKDEEKSKQ-----ENEEIKD 186
Query: 61 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
K+ + + K E+S++ + + ++ Q +++ K E ET + ++
Sbjct: 187 GEKI------QQENEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKETGEITEEKS 348
Query: 121 IRERALKQATESLKGKSTSS----EDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGP 176
+E T S ++ S D K G N+ D K G E G G
Sbjct: 349 KQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQG-------TEETNGTEGGE 507
Query: 177 LRASAHIRVSARFDYQ---PDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEA 233
I+ D Q + + +E Y G S + D +S + E+++D+
Sbjct: 508 KEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASSAVV-DNKSQESSNKTEEQFDKK 684
Query: 234 EK 235
EK
Sbjct: 685 EK 690
Score = 28.9 bits (63), Expect = 2.8
Identities = 26/132 (19%), Positives = 61/132 (45%)
Frame = +1
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 60
++D E EN++ + ++ K+ K ENE +N+ +E+ + ++ +K
Sbjct: 178 IKDGEKIQQENEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKET-GEITEEKSKQ 354
Query: 61 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
+N+ +S++++ K NEES + ++ +++ H+ + E ET+ +
Sbjct: 355 ENE-----ETSETNSKDKENEESNQNG--SDAKEQVGENHEQDSKQGTE-ETNGTEGGEK 510
Query: 121 IRERALKQATES 132
+K+ T S
Sbjct: 511 EEHDKIKEDTSS 546
>TC83740 similar to GP|6437560|gb|AAF08587.1| hypothetical protein
{Arabidopsis thaliana}, partial (10%)
Length = 435
Score = 33.9 bits (76), Expect = 0.089
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +2
Query: 196 CKDYKETGYCGYGDSCKFMHDR 217
C Y TG+CGYG C+F H R
Sbjct: 323 CIYYLRTGFCGYGSRCRFNHPR 388
>TC91110 similar to GP|7576941|gb|AAF64065.1| putative diacylglycerol
acyltransferase {Brassica napus}, partial (19%)
Length = 692
Score = 33.9 bits (76), Expect = 0.089
Identities = 27/108 (25%), Positives = 42/108 (38%), Gaps = 19/108 (17%)
Frame = +2
Query: 107 TLETETDFSRDARAIRERALKQA-------------------TESLKGKSTSSEDVKLYK 147
T+ET+TD R + R A A +SL GK + E+VK K
Sbjct: 116 TIETDTDLKRSSLRRRPSATSTAGGLFDAESAAADAVRDSGSDDSLNGKINNEEEVKDRK 295
Query: 148 GINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSARFDYQPDI 195
TDH G + + K G + + ++ V +F Y+P +
Sbjct: 296 -----TDHAEGIVDDDDDNAVKKNGGNDVINDRENVAVDFKFTYRPSV 424
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,299,320
Number of Sequences: 36976
Number of extensions: 155186
Number of successful extensions: 1049
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 955
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1024
length of query: 327
length of database: 9,014,727
effective HSP length: 96
effective length of query: 231
effective length of database: 5,465,031
effective search space: 1262422161
effective search space used: 1262422161
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126016.2


