

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
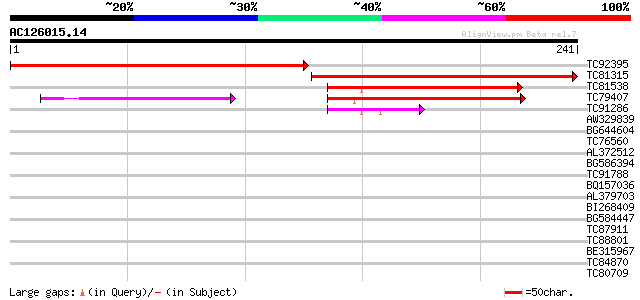
Query= AC126015.14 + phase: 0 /pseudo
(241 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92395 similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_11... 275 1e-74
TC81315 weakly similar to GP|19699320|gb|AAL91270.1 AT3g12090/T2... 248 1e-66
TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associat... 99 1e-21
TC79407 similar to GP|20197174|gb|AAM14957.1 hypothetical protei... 99 2e-21
TC91286 weakly similar to PIR|E84578|E84578 probable senescence-... 47 9e-06
AW329839 similar to PIR|T47500|T475 hypothetical protein F9K21.1... 33 0.099
BG644604 30 0.64
TC76560 homologue to SP|P25804|CYSP_PEA Cysteine proteinase 15A ... 30 1.1
AL372512 similar to GP|21536926|gb unknown {Arabidopsis thaliana... 29 1.4
BG586394 similar to GP|7295463|gb| CG4835 gene product {Drosophi... 29 1.9
TC91788 auxin efflux carrier protein [Medicago truncatula] 29 1.9
BQ157036 similar to GP|21537250|gb putative cell differentiation... 28 2.4
AL379703 homologue to GP|6539600|gb| amino acid transporter a {V... 28 3.2
BI268409 28 3.2
BG584447 28 3.2
TC87911 similar to PIR|E71427|E71427 hypothetical protein - Arab... 28 4.1
TC88801 similar to GP|4337044|gb|AAD18052.1| molybdopterin synth... 28 4.1
BE315967 similar to GP|10177808|dbj cytochrome P450-like protein... 28 4.1
TC84870 similar to GP|21554728|gb|AAM63672.1 putative esterase {... 28 4.1
TC80709 similar to PIR|B84616|B84616 hypothetical protein At2g22... 28 4.1
>TC92395 similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_110
{Arabidopsis thaliana}, partial (54%)
Length = 575
Score = 275 bits (702), Expect = 1e-74
Identities = 127/127 (100%), Positives = 127/127 (100%)
Frame = +3
Query: 1 MDGKEQYNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRF 60
MDGKEQYNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRF
Sbjct: 195 MDGKEQYNLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRF 374
Query: 61 DYIWFWCD**RWWCRSAW*IL**VSSYRLFTLVEEKNTRSSLLEYYKELYFRV*NL*QTC 120
DYIWFWCD**RWWCRSAW*IL**VSSYRLFTLVEEKNTRSSLLEYYKELYFRV*NL*QTC
Sbjct: 375 DYIWFWCD**RWWCRSAW*IL**VSSYRLFTLVEEKNTRSSLLEYYKELYFRV*NL*QTC 554
Query: 121 LLDPFGL 127
LLDPFGL
Sbjct: 555 LLDPFGL 575
>TC81315 weakly similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_110
{Arabidopsis thaliana}, partial (25%)
Length = 629
Score = 248 bits (634), Expect = 1e-66
Identities = 113/113 (100%), Positives = 113/113 (100%)
Frame = +3
Query: 129 AE*YVSNTGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
AE*YVSNTGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN
Sbjct: 9 AE*YVSNTGCCKPPTACNYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDYH 241
WHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDYH
Sbjct: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRPRWDYH 347
>TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associated protein
5-like protein {Arabidopsis thaliana}, partial (92%)
Length = 1073
Score = 99.0 bits (245), Expect = 1e-21
Identities = 41/90 (45%), Positives = 59/90 (65%), Gaps = 7/90 (7%)
Frame = +1
Query: 136 TGCCKPPTACNYN-------MEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRRN 188
+GCCKPPT C Y + + D DC +WSN+ T LCY CDSCKAG+L ++R+
Sbjct: 538 SGCCKPPTQCAYTFVNPTYWISPINNAADMDCLQWSNDQTTLCYNCDSCKAGLLANLRKE 717
Query: 189 WHKLSVLTVTMLILLIGIYSIGCCAFRNAR 218
W + +V+ + +++LI +Y IGC AFRNA+
Sbjct: 718 WKRANVILIITVVVLIVVYLIGCFAFRNAK 807
>TC79407 similar to GP|20197174|gb|AAM14957.1 hypothetical protein
{Arabidopsis thaliana}, partial (87%)
Length = 1053
Score = 98.6 bits (244), Expect = 2e-21
Identities = 43/92 (46%), Positives = 60/92 (64%), Gaps = 8/92 (8%)
Frame = +2
Query: 136 TGCCKPPTAC--------NYNMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 187
+GCCKP C N+ M A + DC W+N+P +LC+ C SCKAG+L++++
Sbjct: 623 SGCCKPSNDCGFTYQNPTNWTMPAGGTYTNPDCDTWTNDPKVLCFNCKSCKAGLLDNLKT 802
Query: 188 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARR 219
NW K++V+ + LI LI +YSIGCCAFRN RR
Sbjct: 803 NWKKVAVVNIIFLIFLIIVYSIGCCAFRNNRR 898
Score = 43.9 bits (102), Expect = 6e-05
Identities = 33/83 (39%), Positives = 43/83 (51%)
Frame = +1
Query: 14 PNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWCD**RWW 73
P S NRF + FHW L SC M ++V+ +V+A+ IRF + F C * R W
Sbjct: 259 PRCFSLNRFS------NGFHWWLLSCYMASLVLSLFHVLAHRCAIRFHHFCFCCH*QRCW 420
Query: 74 CRSAW*IL**VSSYRLFTLVEEK 96
S * L V ++ LF LV EK
Sbjct: 421 *ISFQ*GLQRV*TW*LF*LVAEK 489
>TC91286 weakly similar to PIR|E84578|E84578 probable senescence-associated
protein 5 [imported] - Arabidopsis thaliana, partial
(71%)
Length = 759
Score = 46.6 bits (109), Expect = 9e-06
Identities = 20/48 (41%), Positives = 28/48 (57%), Gaps = 7/48 (14%)
Frame = +2
Query: 136 TGCCKPPTACNYN-MEAVMMTQ------DSDCYKWSNEPTLLCYECDS 176
+GCCKPPT C Y+ + +M T DS C W+N+ LCY C++
Sbjct: 596 SGCCKPPTVCGYSYVSPIMWTNPVNPMADSRCNLWNNDQKQLCYNCNA 739
>AW329839 similar to PIR|T47500|T475 hypothetical protein F9K21.180 -
Arabidopsis thaliana, partial (11%)
Length = 229
Score = 33.1 bits (74), Expect = 0.099
Identities = 13/24 (54%), Positives = 18/24 (74%)
Frame = +2
Query: 213 AFRNARRAETDYPYGENRMTKVRP 236
A+RN ++ + D PYGE RMTK +P
Sbjct: 11 AYRNNKKMDNDEPYGEARMTKSQP 82
>BG644604
Length = 646
Score = 30.4 bits (67), Expect = 0.64
Identities = 11/41 (26%), Positives = 21/41 (50%)
Frame = +2
Query: 22 FYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDY 62
F+ + S+F +C F CC+C+ + G+ V+ + Y
Sbjct: 410 FFILVCLASFFFFCKFGCCVCSPIEKGNTVVDTLEIFQIRY 532
>TC76560 homologue to SP|P25804|CYSP_PEA Cysteine proteinase 15A precursor
(EC 3.4.22.-) (Turgor-responsive protein 15A). [Garden
pea], partial (98%)
Length = 1642
Score = 29.6 bits (65), Expect = 1.1
Identities = 21/67 (31%), Positives = 28/67 (41%)
Frame = +2
Query: 8 NLCKFSPNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWFWC 67
N CK S SC+ + C+ D + WCL +C G + +SS WFW
Sbjct: 944 NCCKSSKEWPSCSCY*CSVD-ADIYEWCLMPIHLC-QSTFGSW--GSSS-------WFWP 1090
Query: 68 D**RWWC 74
RW C
Sbjct: 1091---RWLC 1102
>AL372512 similar to GP|21536926|gb unknown {Arabidopsis thaliana}, partial
(34%)
Length = 462
Score = 29.3 bits (64), Expect = 1.4
Identities = 14/52 (26%), Positives = 23/52 (43%)
Frame = +1
Query: 14 PNTTSCNRFYCTCDIISWFHWCLFSCCMCTMVVLGDYVIANSSTIRFDYIWF 65
P T + Y +I+ WF+ F+C + D + N++ R Y WF
Sbjct: 97 PVTLCTGQTYERNNILKWFNMGHFTCPTTMQELWDDSITPNTTLYRLIYTWF 252
>BG586394 similar to GP|7295463|gb| CG4835 gene product {Drosophila
melanogaster}, partial (2%)
Length = 734
Score = 28.9 bits (63), Expect = 1.9
Identities = 14/33 (42%), Positives = 16/33 (48%), Gaps = 12/33 (36%)
Frame = -3
Query: 20 NRFYCTC-----------DIISWFH-WCLFSCC 40
+RF C C +I WF WCLFSCC
Sbjct: 540 SRFSCCCFIFF*CCSSLWNICFWFFSWCLFSCC 442
>TC91788 auxin efflux carrier protein [Medicago truncatula]
Length = 2076
Score = 28.9 bits (63), Expect = 1.9
Identities = 13/38 (34%), Positives = 18/38 (47%), Gaps = 11/38 (28%)
Frame = +3
Query: 17 TSCNRFY---------CTCDIIS--WFHWCLFSCCMCT 43
TSCN FY C C ++ W+ W +CC C+
Sbjct: 1620 TSCNIFYGCKVLDRSSCDCGNLNSNWYPWSSVTCCNCS 1733
>BQ157036 similar to GP|21537250|gb putative cell differentiation protein
{Arabidopsis thaliana}, partial (50%)
Length = 672
Score = 28.5 bits (62), Expect = 2.4
Identities = 8/30 (26%), Positives = 19/30 (62%)
Frame = +3
Query: 39 CCMCTMVVLGDYVIANSSTIRFDYIWFWCD 68
C C++ V D VI+ + + +++W++C+
Sbjct: 123 CSSCSLQVSKDQVISRTCSFIMEFLWYYCN 212
>AL379703 homologue to GP|6539600|gb| amino acid transporter a {Vicia faba},
partial (60%)
Length = 481
Score = 28.1 bits (61), Expect = 3.2
Identities = 19/54 (35%), Positives = 28/54 (51%)
Frame = +1
Query: 157 DSDCYKWSNEPTLLCYECDSCKAGVLEDIRRNWHKLSVLTVTMLILLIGIYSIG 210
D+ CY SN P ++ + C VL I N+HKLS L++ ++ SIG
Sbjct: 157 DAKCYI-SNNPFMIIFACIQI---VLSQIP-NFHKLSWLSIVAAVMSFAYSSIG 303
>BI268409
Length = 148
Score = 28.1 bits (61), Expect = 3.2
Identities = 15/46 (32%), Positives = 20/46 (42%), Gaps = 3/46 (6%)
Frame = +1
Query: 23 YCTCDIISWFHWCLF---SCCMCTMVVLGDYVIANSSTIRFDYIWF 65
YC C W+H C F S C T+ +L +V D +WF
Sbjct: 34 YCLCHTSFWYHLCKFQANSACDITLCLLMMFV--------KDVVWF 147
>BG584447
Length = 802
Score = 28.1 bits (61), Expect = 3.2
Identities = 10/30 (33%), Positives = 15/30 (49%), Gaps = 3/30 (10%)
Frame = -1
Query: 42 CTMVVLGDYVIANSSTIR---FDYIWFWCD 68
C + G Y + S ++ + Y WFWCD
Sbjct: 358 CFRCIAGSYHFSGRSPLKALWYSYSWFWCD 269
>TC87911 similar to PIR|E71427|E71427 hypothetical protein - Arabidopsis
thaliana, partial (24%)
Length = 639
Score = 27.7 bits (60), Expect = 4.1
Identities = 13/37 (35%), Positives = 18/37 (48%), Gaps = 1/37 (2%)
Frame = +3
Query: 8 NLCKFSPNTTSCNR-FYCTCDIISWFHWCLFSCCMCT 43
N+C + P S N +YCTC + S CC C+
Sbjct: 468 NVCMWMPQEYSKNM*YYCTCHVAS------LKCCPCS 560
>TC88801 similar to GP|4337044|gb|AAD18052.1| molybdopterin synthase
sulphurylase {Nicotiana plumbaginifolia}, partial (96%)
Length = 1547
Score = 27.7 bits (60), Expect = 4.1
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +3
Query: 13 SPNTTSCNRFYCTCDIISWFHWCL 36
SP TT RF+C C II WF + +
Sbjct: 189 SPKTT---RFFCFCCIIQWFFFTI 251
>BE315967 similar to GP|10177808|dbj cytochrome P450-like protein
{Arabidopsis thaliana}, partial (4%)
Length = 437
Score = 27.7 bits (60), Expect = 4.1
Identities = 10/21 (47%), Positives = 12/21 (56%)
Frame = -2
Query: 26 CDIISWFHWCLFSCCMCTMVV 46
C SW HWC C+ TMV+
Sbjct: 337 CVARSWGHWCKECSCLLTMVL 275
>TC84870 similar to GP|21554728|gb|AAM63672.1 putative esterase {Arabidopsis
thaliana}, partial (12%)
Length = 500
Score = 27.7 bits (60), Expect = 4.1
Identities = 23/87 (26%), Positives = 33/87 (37%), Gaps = 26/87 (29%)
Frame = -2
Query: 5 EQYNLCKFSPNTT---------------SCNRF-YCTCDIISWFHWCLFSC--------- 39
E+ NLC F N+ SC +F + + + W W +F C
Sbjct: 376 ERTNLCLFQCNSKVQYEISVMGGTFSLHSCFQFVFFSSSPLRWKGWIVFLCFLLFGWVTG 197
Query: 40 CMCTMVVLGDYVIANSSTI-RFDYIWF 65
C+C V YV+A S + F Y F
Sbjct: 196 CVCGTAVTSCYVLAGSEVVCSFGYYLF 116
>TC80709 similar to PIR|B84616|B84616 hypothetical protein At2g22730
[imported] - Arabidopsis thaliana, partial (40%)
Length = 1250
Score = 27.7 bits (60), Expect = 4.1
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +3
Query: 180 GVLEDIRRNWHKLSVLTVTMLILLIGIYSIG 210
GVL+D +W K S+ ++ L G++ IG
Sbjct: 630 GVLQDHINDWRKTSICLTSIFFLAAGVWFIG 722
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.343 0.149 0.575
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,407,395
Number of Sequences: 36976
Number of extensions: 218789
Number of successful extensions: 3216
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 3109
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3204
length of query: 241
length of database: 9,014,727
effective HSP length: 93
effective length of query: 148
effective length of database: 5,575,959
effective search space: 825241932
effective search space used: 825241932
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126015.14


