

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.12 - phase: 0
(1620 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
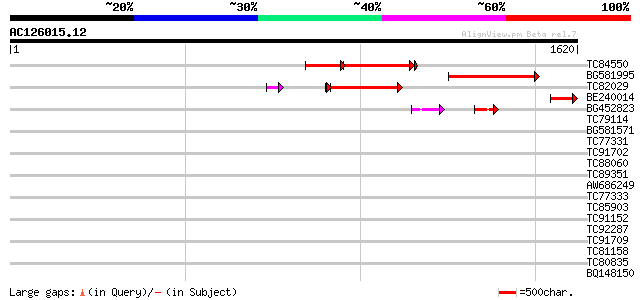
Score E
Sequences producing significant alignments: (bits) Value
TC84550 weakly similar to PIR|T46095|T46095 hypothetical protein... 362 e-164
BG581995 522 e-148
TC82029 weakly similar to PIR|T46095|T46095 hypothetical protein... 181 9e-46
BE240014 153 6e-37
BG452823 similar to PIR|E90181|E901 hypothetical protein SSO0379... 75 3e-13
TC79114 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~un... 42 0.003
BG581571 40 0.007
TC77331 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~un... 39 0.021
TC91702 homologue to GP|21593278|gb|AAM65227.1 contains similari... 37 0.048
TC88060 homologue to GP|21593278|gb|AAM65227.1 contains similari... 35 0.24
TC89351 similar to PIR|G84786|G84786 hypothetical protein At2g36... 34 0.40
AW686249 similar to GP|9294690|dbj gene_id:MHC9.11~unknown prote... 34 0.53
TC77333 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~un... 33 0.69
TC85903 similar to PIR|B84530|B84530 probable RING zinc finger p... 33 0.69
TC91152 similar to GP|9759423|dbj|BAB09910.1 contains similarity... 33 0.90
TC92287 weakly similar to PIR|H84919|H84919 hypothetical protein... 33 0.90
TC91709 homologue to GP|21213868|emb|CAD12767. LHY protein {Phas... 33 1.2
TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidops... 32 2.6
TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF com... 32 2.6
BQ148150 weakly similar to GP|10185719|gb NTS1 protein {Nicotian... 31 3.4
>TC84550 weakly similar to PIR|T46095|T46095 hypothetical protein T25B15.20 -
Arabidopsis thaliana, partial (1%)
Length = 969
Score = 362 bits (929), Expect(3) = e-164
Identities = 182/203 (89%), Positives = 188/203 (91%)
Frame = +1
Query: 955 ETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGLNLANPVPGINGSPLND 1014
+++ F + + + RCVGTKAQEHCKRFFSKTRKCLGLNLANPVPGINGSPLND
Sbjct: 337 DSSLFFRLYHLLVRILRRYHRCVGTKAQEHCKRFFSKTRKCLGLNLANPVPGINGSPLND 516
Query: 1015 DANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNPLEATSLSAKLNE 1074
DANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNPLEATSLSAKLNE
Sbjct: 517 DANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNPLEATSLSAKLNE 696
Query: 1075 SREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSVNGVGLGGTVRESI 1134
SREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSVNGVGLGGTVRESI
Sbjct: 697 SREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSVNGVGLGGTVRESI 876
Query: 1135 SASEIIKPRECGSVALDRTVSEG 1157
SAS IIKPRECGSVALDRTVS G
Sbjct: 877 SASXIIKPRECGSVALDRTVSXG 945
Score = 236 bits (602), Expect(3) = e-164
Identities = 116/116 (100%), Positives = 116/116 (100%)
Frame = +2
Query: 844 RRYLLGYGNVKASRGEDSIIERSNSFDTLGDERETAAAADVLAGICGSFSSEAMSSCITS 903
RRYLLGYGNVKASRGEDSIIERSNSFDTLGDERETAAAADVLAGICGSFSSEAMSSCITS
Sbjct: 2 RRYLLGYGNVKASRGEDSIIERSNSFDTLGDERETAAAADVLAGICGSFSSEAMSSCITS 181
Query: 904 SIDPVDGNKETKFLKANPLFKQPLTPDISQNADDETCSDESCGEATEWTDDETAAF 959
SIDPVDGNKETKFLKANPLFKQPLTPDISQNADDETCSDESCGEATEWTDDETAAF
Sbjct: 182 SIDPVDGNKETKFLKANPLFKQPLTPDISQNADDETCSDESCGEATEWTDDETAAF 349
Score = 21.2 bits (43), Expect(3) = e-164
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = +2
Query: 1157 GSSGGLCL 1164
GSSGGLCL
Sbjct: 944 GSSGGLCL 967
>BG581995
Length = 799
Score = 522 bits (1345), Expect = e-148
Identities = 258/259 (99%), Positives = 259/259 (99%)
Frame = +1
Query: 1254 LLLKAAAAQCEKTVSQDRLSSTCDIQGGRDMRCHSSGSNGDHQLPLSGSHVETVSVLQGY 1313
LLLKAAAAQCEKTVSQDRLSSTCDIQGGRDMRCHSSGSNGDHQLPLSGSHVETVSVLQGY
Sbjct: 22 LLLKAAAAQCEKTVSQDRLSSTCDIQGGRDMRCHSSGSNGDHQLPLSGSHVETVSVLQGY 201
Query: 1314 SMQVPIKKEVDGDVNCSSSAAEFPLLPQKVKQTDGHFKPSFHSSNSEKTSRNGDVKLFGK 1373
SMQVPIKKEVDGDVNCSSSAAEFPLLPQKVKQTDGHFKPSFHSSNSEKTSRNGDVKLFGK
Sbjct: 202 SMQVPIKKEVDGDVNCSSSAAEFPLLPQKVKQTDGHFKPSFHSSNSEKTSRNGDVKLFGK 381
Query: 1374 ILTNPSSTQNPNLTAKRSEENGSHHPKLNNKSSNLNFTGHQNSDENLNFLKFGLENVPVM 1433
ILTNPSSTQNPNLTAKRSEENGSHHPKLNNKSSNLNFTGHQNSDENLNFLKFGLENVPVM
Sbjct: 382 ILTNPSSTQNPNLTAKRSEENGSHHPKLNNKSSNLNFTGHQNSDENLNFLKFGLENVPVM 561
Query: 1434 SYGYWEGNAIQSRQSGLSSLPDSSFLLAKYPAAFSNYPTSSSNLEQQPPLQAFAKNSQRH 1493
SYGYWEGNAIQSRQSGLSSLPDSSFLLAKYPAAFSNYPTSSSNLEQQPPLQAFAKNSQRH
Sbjct: 562 SYGYWEGNAIQSRQSGLSSLPDSSFLLAKYPAAFSNYPTSSSNLEQQPPLQAFAKNSQRH 741
Query: 1494 LTGASTFTARDVNGSNAML 1512
LTGASTFTA+DVNGSNAML
Sbjct: 742 LTGASTFTAKDVNGSNAML 798
>TC82029 weakly similar to PIR|T46095|T46095 hypothetical protein T25B15.20 -
Arabidopsis thaliana, partial (3%)
Length = 776
Score = 181 bits (458), Expect(2) = 9e-46
Identities = 97/206 (47%), Positives = 134/206 (64%), Gaps = 2/206 (0%)
Frame = +1
Query: 918 KANPLFKQPLTPDISQNADDETCSDESCGEA--TEWTDDETAAFLQAVSSFGKDFEKISR 975
K + KQP P ++Q+ D E+CSDESCGE T+WTD E AAFL A+SSFGKDF KI++
Sbjct: 88 KPRAICKQPPMPVMTQSIDPESCSDESCGEMELTDWTDVEKAAFLHALSSFGKDFAKIAQ 267
Query: 976 CVGTKAQEHCKRFFSKTRKCLGLNLANPVPGINGSPLNDDANGGESDTDDACVVEAGSVV 1035
CVGT++Q C+ FF KT K +G++L G+ GSP +DD +GG SD D+AC+ E S
Sbjct: 268 CVGTRSQFQCRAFFCKTHKRIGMDLMGERAGVVGSPGDDDVDGGRSDADNACIPETASAN 447
Query: 1036 DADKSGNKTDEDLPSDALNTFHDESNPLEATSLSAKLNESREISGTEVCLENVDVASVAC 1095
+D SG+KT D P+ N + +ESNPL A +LSA+ +ES EI+G + + V++ S
Sbjct: 448 GSDTSGSKTVVDQPAYDKNLYQEESNPLVARNLSAESDESEEING-KADRKEVNIFSNEY 624
Query: 1096 AINVESKLGSDVSGVGLCTTDKSGSV 1121
ESKLG+D + L + D G+V
Sbjct: 625 VTGSESKLGTDCNAAALYSFDGPGTV 702
Score = 43.5 bits (101), Expect = 7e-04
Identities = 19/47 (40%), Positives = 26/47 (54%)
Frame = +1
Query: 735 INPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ WT EK FL ++FGKDF KIA + ++ C F+ K HK
Sbjct: 184 LTDWTDVEKAAFLHALSSFGKDFAKIAQCVGTRSQFQCRAFFCKTHK 324
Score = 23.1 bits (48), Expect(2) = 9e-46
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +3
Query: 901 ITSSIDPVDGNKETKFLKA 919
+TSS DPV G + K +KA
Sbjct: 36 LTSSADPVKGKRVRKSVKA 92
>BE240014
Length = 278
Score = 153 bits (386), Expect = 6e-37
Identities = 73/75 (97%), Positives = 74/75 (98%)
Frame = +3
Query: 1546 HSFEAISSLQQQGRGMMGMNSVGRPGILVGGSCSGVSDPVAAIKMHYSNSEKYGGQNGSV 1605
HSFEAISSLQQ+GRGMMGMNSVGRPGILVGGSCSGVSDPVAAIKMHYSNSEKYGGQNGSV
Sbjct: 3 HSFEAISSLQQRGRGMMGMNSVGRPGILVGGSCSGVSDPVAAIKMHYSNSEKYGGQNGSV 182
Query: 1606 VRDDESWGGKGDLGR 1620
VRDDESWGGKGDL R
Sbjct: 183 VRDDESWGGKGDLSR 227
>BG452823 similar to PIR|E90181|E901 hypothetical protein SSO0379 [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 654
Score = 74.7 bits (182), Expect = 3e-13
Identities = 42/69 (60%), Positives = 49/69 (70%)
Frame = +3
Query: 1329 CSSSAAEFPLLPQKVKQTDGHFKPSFHSSNSEKTSRNGDVKLFGKILTNPSSTQNPNLTA 1388
CSSSA E PLLPQKV+Q H K NSE T +VK+FGK+LT PSSTQ PN +A
Sbjct: 390 CSSSATELPLLPQKVEQQGPH-KSLQCLPNSEDTP---NVKIFGKVLTIPSSTQKPNSSA 557
Query: 1389 KRSEENGSH 1397
KR+EENG+H
Sbjct: 558 KRNEENGTH 584
Score = 54.7 bits (130), Expect = 3e-07
Identities = 45/96 (46%), Positives = 54/96 (55%), Gaps = 1/96 (1%)
Frame = +3
Query: 1147 SVALDRTVSEGSSGGLCLGSEVERQRVSAPHCVVDKDVEHVADAGVVVELKNCVLESSTA 1206
S A R VS+GSSG G+E+E V C VD D E V D VV+ELK+ V S T+
Sbjct: 93 SAAEGRLVSDGSSG--LQGNELEASTV----CRVDTD-EVVTD--VVIELKDNVHNSRTS 245
Query: 1207 ANVSFSPVVNSCSGLSFGSENK-HVSFGKPHTSALS 1241
N S SPV SCS LS +EN+ + KP S LS
Sbjct: 246 GNTSLSPVEASCSRLSADAENEPQLCLEKPQFSGLS 353
>TC79114 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~unknown
protein {Arabidopsis thaliana}, partial (32%)
Length = 759
Score = 41.6 bits (96), Expect = 0.003
Identities = 19/49 (38%), Positives = 30/49 (60%), Gaps = 4/49 (8%)
Frame = +1
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRK 994
+WTD+E FL+A+ +G+ + KI VGTK + H ++FFSK +
Sbjct: 283 KWTDEEHKKFLEALKLYGRAWRKIEEHVGTKTAVQIRSHAQKFFSKINR 429
Score = 37.0 bits (84), Expect = 0.062
Identities = 25/89 (28%), Positives = 39/89 (43%), Gaps = 3/89 (3%)
Frame = +1
Query: 709 EKMVTKFISSNG---LVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLD 765
+K++ +F N P I K+R WT EE + FLE +G+ +RKI +
Sbjct: 199 QKLLDQFSCGNDHALKARKPYTITKQREK---WTDEEHKKFLEALKLYGRAWRKIEEHVG 369
Query: 766 HKTTADCIEFYYKNHKSECFEKLKRKDIG 794
KT ++H + F K+ R G
Sbjct: 370 TKTAVQ-----IRSHAQKFFSKINRDTDG 441
>BG581571
Length = 439
Score = 40.0 bits (92), Expect = 0.007
Identities = 25/39 (64%), Positives = 25/39 (64%)
Frame = +3
Query: 1429 NVPVMSYGYWEGNAIQSRQSGLSSLPDSSFLLAKYPAAF 1467
N P M G GN IQS G SSLP SS LLAKYPAAF
Sbjct: 90 NAPYM--GCSHGNTIQS---GFSSLPASSILLAKYPAAF 191
>TC77331 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~unknown protein
{Arabidopsis thaliana}, partial (25%)
Length = 1835
Score = 38.5 bits (88), Expect = 0.021
Identities = 19/66 (28%), Positives = 37/66 (55%), Gaps = 4/66 (6%)
Frame = +1
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTKA----QEHCKRFFSKTRKCLGLNLANPVP 1005
+WTD+E FL+A+ +G+ + +I +G+K + H ++FFSK + + +P+
Sbjct: 367 KWTDEEHQKFLEALKLYGRGWRQIEEHIGSKTAIQIRSHAQKFFSKVVREPDGSAESPIQ 546
Query: 1006 GINGSP 1011
I+ P
Sbjct: 547 PIDIPP 564
Score = 37.0 bits (84), Expect = 0.062
Identities = 22/73 (30%), Positives = 34/73 (46%)
Frame = +1
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
V P I K+R WT EE + FLE +G+ +R+I + KT ++H
Sbjct: 331 VRKPYTITKQREK---WTDEEHQKFLEALKLYGRGWRQIEEHIGSKTA-----IQIRSHA 486
Query: 782 SECFEKLKRKDIG 794
+ F K+ R+ G
Sbjct: 487 QKFFSKVVREPDG 525
>TC91702 homologue to GP|21593278|gb|AAM65227.1 contains similarity to
MYB-related DNA-binding protein {Arabidopsis thaliana},
partial (28%)
Length = 676
Score = 37.4 bits (85), Expect = 0.048
Identities = 21/72 (29%), Positives = 35/72 (48%)
Frame = +1
Query: 719 NGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYK 778
N + P I K R WT +E + FLE F +D++KI +F+ KT +
Sbjct: 265 NKKIRKPYTITKSRES---WTDQEHDKFLEALQLFDRDWKKIEAFVGSKTV-----IQIR 420
Query: 779 NHKSECFEKLKR 790
+H + F K+++
Sbjct: 421 SHAQKYFLKVQK 456
>TC88060 homologue to GP|21593278|gb|AAM65227.1 contains similarity to
MYB-related DNA-binding protein {Arabidopsis thaliana},
partial (48%)
Length = 1370
Score = 35.0 bits (79), Expect = 0.24
Identities = 20/69 (28%), Positives = 33/69 (46%)
Frame = +3
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ P I K R WT E + FLE F +D++KI +F+ KT ++H
Sbjct: 180 IRKPYTITKSREN---WTEPEHDKFLEALQLFDRDWKKIEAFVGSKTV-----IQIRSHA 335
Query: 782 SECFEKLKR 790
+ F K+++
Sbjct: 336 QKYFLKVQK 362
>TC89351 similar to PIR|G84786|G84786 hypothetical protein At2g36960
[imported] - Arabidopsis thaliana, partial (21%)
Length = 968
Score = 34.3 bits (77), Expect = 0.40
Identities = 30/132 (22%), Positives = 56/132 (41%)
Frame = +1
Query: 647 TCSSNMKNRSSIRSRFTFPAGNHLSLVPTTEIINFTSKLLSESQAQLQRNTLKMPALILD 706
T SS K +S I SR ++ E + +++L + QL T+ + +
Sbjct: 19 TASSRNKLKSQIASR---KRRI*IARACEEEKMEAEAEILIDLNTQLHSETI----VSVQ 177
Query: 707 EKEKMVTKFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDH 766
++ V+ ++++ ++ A +K WT E+E F GK+F KI S +
Sbjct: 178 NEDPGVSTSLANDVIMPQQPAPKKPTRQWAAWTRNEEESFFTALRQVGKNFEKITSRVQS 357
Query: 767 KTTADCIEFYYK 778
K +YY+
Sbjct: 358 KNKDQVRHYYYR 393
Score = 32.3 bits (72), Expect = 1.5
Identities = 13/39 (33%), Positives = 24/39 (61%)
Frame = +1
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFF 989
WT +E +F A+ GK+FEKI+ V +K ++ + ++
Sbjct: 271 WTRNEEESFFTALRQVGKNFEKITSRVQSKNKDQVRHYY 387
>AW686249 similar to GP|9294690|dbj gene_id:MHC9.11~unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 657
Score = 33.9 bits (76), Expect = 0.53
Identities = 14/49 (28%), Positives = 26/49 (52%)
Frame = +1
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGL 998
+W+ E F +A +GKD++K++ V + E + ++K R L L
Sbjct: 319 QWSKAELERFYEAYREYGKDWKKVALAVRNRTMEMVEALYTKNRAYLSL 465
>TC77333 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~unknown
protein {Arabidopsis thaliana}, partial (19%)
Length = 1058
Score = 33.5 bits (75), Expect = 0.69
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = +1
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCVGT 979
+WTD+E FL+A+ +G+ + +I C GT
Sbjct: 418 KWTDEEHQKFLEALKLYGRGWRQIEGCAGT 507
Score = 30.4 bits (67), Expect = 5.8
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +1
Query: 722 VEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKI 760
V P I K+R WT EE + FLE +G+ +R+I
Sbjct: 382 VRKPYTITKQREK---WTDEEHQKFLEALKLYGRGWRQI 489
>TC85903 similar to PIR|B84530|B84530 probable RING zinc finger protein
[imported] - Arabidopsis thaliana, partial (26%)
Length = 2572
Score = 33.5 bits (75), Expect = 0.69
Identities = 27/119 (22%), Positives = 46/119 (37%), Gaps = 16/119 (13%)
Frame = +2
Query: 58 EASNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVN-----TWEQHNLKDQHAKTGG 112
EASN P++ +++ R+ D T+ HP S +N + HN H G
Sbjct: 1034 EASNAFPSMVSIAENVERPLRNFDRRMTHLHHPESVPLNLTSTGSARHHNYPSPHQIPGS 1213
Query: 113 VN-----------GLGTGPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSM 160
++ G+ +N S+ P W R+ + S S+SS R++
Sbjct: 1214 LSFNESLDLRLAAGVTAANSAVPQNQSPSLHMHPFPWNRAANPRVARSSSSYSSGERAV 1390
>TC91152 similar to GP|9759423|dbj|BAB09910.1 contains similarity to unknown
protein~dbj|BAA90946.1~gene_id:K7J8.7 {Arabidopsis
thaliana}, partial (44%)
Length = 792
Score = 33.1 bits (74), Expect = 0.90
Identities = 43/162 (26%), Positives = 69/162 (42%), Gaps = 4/162 (2%)
Frame = +1
Query: 19 MLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSW--EASNGSPNLSR--RPQDMN 74
M E L R D K S+ D G Q+ ++G W + N +SR R Q +
Sbjct: 322 MSTEKAVKLGERKDVKPTMSTVD--GNAQCQKCYQGGHWTFQCKNERVYISRPSRTQQLK 495
Query: 75 NEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPRCDRENSLSSIDWK 134
N + D S Y +++ N ++K++ AK + + R+ S
Sbjct: 496 NPKLRPDVSVGYDLDDNNN--NNHRDRDVKEEKAKV-------SSKKSKRKYDSVSNSED 648
Query: 135 PLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKN 176
+ T SGS SS +G +SSSS S + ++S E + K K+
Sbjct: 649 SVFETDSGSGSSSATGSDYSSSSGSSSSSESEEERRRKKKKS 774
>TC92287 weakly similar to PIR|H84919|H84919 hypothetical protein At2g47820
[imported] - Arabidopsis thaliana, partial (11%)
Length = 989
Score = 33.1 bits (74), Expect = 0.90
Identities = 22/75 (29%), Positives = 30/75 (39%)
Frame = +2
Query: 738 WTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSECFEKLKRKDIGKLG 797
WT E + FL AFGK+ + F+ K+ D + FYY K K +
Sbjct: 623 WTDTEYDSFLLGLYAFGKNLTFLKRFVGTKSMGDILFFYYSKF-------FKSKGYSRWS 781
Query: 798 KSYAAKTNLMASGKK 812
AKT G+K
Sbjct: 782 GCRKAKTKRCIFGQK 826
Score = 32.0 bits (71), Expect = 2.0
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = +2
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKA 981
WTD E +FL + +FGK+ + R VGTK+
Sbjct: 623 WTDTEYDSFLLGLYAFGKNLTFLKRFVGTKS 715
>TC91709 homologue to GP|21213868|emb|CAD12767. LHY protein {Phaseolus
vulgaris}, partial (8%)
Length = 649
Score = 32.7 bits (73), Expect = 1.2
Identities = 12/30 (40%), Positives = 21/30 (70%)
Frame = +1
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTK 980
WT+DE FL+A+ +G+ +++I +GTK
Sbjct: 547 WTEDEHNRFLEALKLYGRAWQRIEEHIGTK 636
>TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidopsis thaliana},
partial (6%)
Length = 890
Score = 31.6 bits (70), Expect = 2.6
Identities = 16/50 (32%), Positives = 26/50 (52%)
Frame = +1
Query: 1003 PVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDA 1052
P +NG+ DD + GES+++ + S + SG+ +DED DA
Sbjct: 379 PSVEVNGNEDKDDGSEGESESESSESESESSTSSSSTSGSDSDEDDEDDA 528
>TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF complex
subunit SW13 [imported] - Arabidopsis thaliana, partial
(34%)
Length = 830
Score = 31.6 bits (70), Expect = 2.6
Identities = 11/28 (39%), Positives = 21/28 (74%)
Frame = +2
Query: 950 EWTDDETAAFLQAVSSFGKDFEKISRCV 977
EWT+ ET L+A+++FG D+++++ V
Sbjct: 731 EWTEKETLNLLEAITNFGDDWKRVAHQV 814
>BQ148150 weakly similar to GP|10185719|gb NTS1 protein {Nicotiana tabacum},
partial (31%)
Length = 669
Score = 31.2 bits (69), Expect = 3.4
Identities = 16/62 (25%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +2
Query: 1195 ELKNCVLESSTAAN-VSFSPVVNSCSGLSFGSENKHVSFGKPHTSALSMSMSDLQATANS 1253
++KN +++ T+ N + F ++ CS +SF + + P+T + +S SD NS
Sbjct: 293 KIKNAIVQDVTSLNSMQFHFHLHGCSNISFTNLHITAPGNSPNTDGMHISSSDFITVTNS 472
Query: 1254 LL 1255
++
Sbjct: 473 VI 478
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.128 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,354,108
Number of Sequences: 36976
Number of extensions: 691348
Number of successful extensions: 3101
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 3016
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3088
length of query: 1620
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1511
effective length of database: 4,984,343
effective search space: 7531342273
effective search space used: 7531342273
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC126015.12


