

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.9 + phase: 0
(233 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
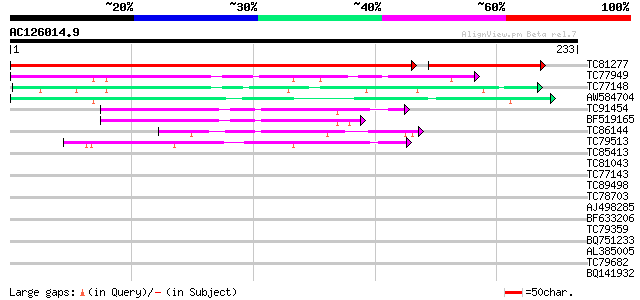
Score E
Sequences producing significant alignments: (bits) Value
TC81277 similar to GP|21536976|gb|AAM61317.1 unknown {Arabidopsi... 353 e-121
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 64 4e-11
TC77148 ENOD20 48 4e-06
AW584704 similar to SP|P05790|FBOH Fibroin heavy chain precursor... 45 2e-05
TC91454 homologue to GP|1071881|dbj|BAA11125.1 orf324 {Beta vulg... 44 7e-05
BF519165 weakly similar to GP|11994144|dbj contains similarity t... 42 3e-04
TC86144 weakly similar to GP|10727325|gb|AAF50995.2 CG15635 gene... 41 3e-04
TC79513 weakly similar to GP|9885806|gb|AAG01535.1| stellacyanin... 40 6e-04
TC85413 similar to PIR|S23737|S23737 proline-rich protein precur... 39 0.001
TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 39 0.001
TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-pro... 39 0.001
TC89498 weakly similar to PIR|A86218|A86218 protein T27G7.18 [im... 39 0.002
TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroele... 38 0.003
AJ498285 weakly similar to GP|20799090|em ADP-ribosylation facto... 38 0.004
BF633206 similar to GP|21553614|gb| blue copper protein putativ... 37 0.005
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 37 0.005
BQ751233 similar to GP|18307448|em probable low-affinity hexose ... 37 0.006
AL385005 weakly similar to GP|21438547|emb Ratten cDNA {Rattus n... 36 0.011
TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 36 0.014
BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 36 0.014
>TC81277 similar to GP|21536976|gb|AAM61317.1 unknown {Arabidopsis
thaliana}, partial (12%)
Length = 740
Score = 353 bits (905), Expect(2) = e-121
Identities = 167/167 (100%), Positives = 167/167 (100%)
Frame = +1
Query: 1 MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQK 60
MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQK
Sbjct: 43 MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWKNPTVSIGDSITFKHKQNYNLYIFKNQK 222
Query: 61 AFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKA 120
AFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKA
Sbjct: 223 AFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTKA 402
Query: 121 SAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEA 167
SAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEA
Sbjct: 403 SAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEA 543
Score = 100 bits (248), Expect(2) = e-121
Identities = 47/48 (97%), Positives = 47/48 (97%)
Frame = +3
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVPI 220
VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQV I
Sbjct: 561 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVMI 704
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 64.3 bits (155), Expect = 4e-11
Identities = 62/217 (28%), Positives = 89/217 (40%), Gaps = 24/217 (11%)
Frame = +3
Query: 1 MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWK-NPTVS-----------IGDSITFKHK 48
M I +F++ +L S + V G W NP+ + I D+I FK+K
Sbjct: 120 MEFKIPLFFVLFTLVVSTSQAFKFFVGGKDGWTLNPSENYNQWAGRNRFQISDTIVFKYK 299
Query: 49 QNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDS 108
+ + + ++ + CN T + T + R G FYF D + C++
Sbjct: 300 KGSDSVLEVKKEDYEKCNKTNPIKKFEDGETEFTL----DRAGPFYFISGKD--QNCENG 461
Query: 109 QKLAIKV----TPTKASAPEAS-----SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSP 159
QKL + V TP + +P A SP PTT PS G SP P P SSP
Sbjct: 462 QKLTLVVISPRTPKSSPSPSAGGLSPPSPSPTTTTPSPSG----SPPSPVAIPP--ASSP 623
Query: 160 GPAPTPEASSPITVPLVPYKG---SGDGMPFINSNPA 193
P P ASSP V P G + P +++ PA
Sbjct: 624 VPTSGPTASSPSPVVSTPPAGGPMASSPSPVVSTPPA 734
Score = 35.8 bits (81), Expect = 0.014
Identities = 25/65 (38%), Positives = 29/65 (44%), Gaps = 9/65 (13%)
Frame = +3
Query: 115 VTPTKASAPEASSPMPTTPGPSSGGDIQSSPS---FPWPFHPHQGSS------PGPAPTP 165
V+ A P ASSP P P +GG + SSPS P P G S P P P
Sbjct: 666 VSTPPAGGPMASSPSPVVSTPPAGGPMASSPSPAGGPPALSPAGGPSTAAGGPPAPGPGG 845
Query: 166 EASSP 170
A+SP
Sbjct: 846 AATSP 860
Score = 30.4 bits (67), Expect = 0.61
Identities = 26/94 (27%), Positives = 34/94 (35%)
Frame = -1
Query: 120 ASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYK 179
A AP + PGP +GG + P G G P E I P
Sbjct: 880 APAPPGPGEVAAPPGPGAGGPPAAVDG------PPAGERAGGPPAGEGDEAIGPPAGGVL 719
Query: 180 GSGDGMPFINSNPAVPLPTGEVDSATIHPLATSG 213
+G+G I L TG+ D A + PL +G
Sbjct: 718 TTGEGDDAIGPPAGGVLTTGDGDDA-VGPLVGTG 620
>TC77148 ENOD20
Length = 1108
Score = 47.8 bits (112), Expect = 4e-06
Identities = 59/246 (23%), Positives = 95/246 (37%), Gaps = 28/246 (11%)
Frame = +3
Query: 2 SLPIFFYFLI-LSLFFKLSHSTTILV-DGSSEWKNPTVS--------------IGDSITF 45
S PI F+ + + S ST LV D + WK P + +GD+ITF
Sbjct: 30 SSPILLMFIFSIWMLISYSESTDYLVGDSENSWKFPLPTRHALTRWASNYQFIVGDTITF 209
Query: 46 KHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKAC 105
++ ++ ++ C ++ T + G +F + + C
Sbjct: 210 QYNNKTESVHEVEEEDYDRCGIRGEHVDHYDGNTMVVL----KKTGIHHFI--SGKKRHC 371
Query: 106 QDSQKLAI--KVTPTKASAPEASSPMPTTPGPSSGGDIQSSP--SFPWPFHPHQGSSPGP 161
+ KLA+ V P +S P P P+ P P S I P S P P P SP P
Sbjct: 372 RLGLKLAVVVMVAPVLSSPP----PPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSP 539
Query: 162 APTPE-ASSPITVPLVPYKGSGDGMPFINSNPA-------VPLPTGEVDSATIHPLATSG 213
+P+P S+PI P S P ++ +P+ P P+ V A++ P ++
Sbjct: 540 SPSPSPRSTPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSPSDSV--ASLAPSSSPS 713
Query: 214 HQGQVP 219
+ P
Sbjct: 714 DESPSP 731
Score = 31.6 bits (70), Expect = 0.27
Identities = 27/85 (31%), Positives = 36/85 (41%), Gaps = 7/85 (8%)
Frame = +3
Query: 117 PTKASAPEASSPMPT---TPGPSSGGDIQSSPSFPW----PFHPHQGSSPGPAPTPEASS 169
P K S P + SP P+ +P PS + SPS P SP PAP+P +S
Sbjct: 579 PRKRS-PASPSPSPSLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSPAPSPSSSG 755
Query: 170 PITVPLVPYKGSGDGMPFINSNPAV 194
KG G G F+ + A+
Sbjct: 756 S--------KGGGAGHGFLEVSIAM 806
>AW584704 similar to SP|P05790|FBOH Fibroin heavy chain precursor (Fib-H)
(H-fibroin). [Silk moth] {Bombyx mori}, partial (0%)
Length = 679
Score = 45.1 bits (105), Expect = 2e-05
Identities = 51/237 (21%), Positives = 83/237 (34%), Gaps = 13/237 (5%)
Frame = +1
Query: 1 MSLPIFFYFLILSLFFKLSHSTTILVDGSSEWK----------NPTVSIGDSITFKHKQN 50
+S +F + LI ++F ++ + +V S W N +GD++TFK+
Sbjct: 58 LSRSLFLFALIATIFSTMAVAKDFVVGDESGWTLGVDYQAWAANKVFRLGDTLTFKYVAW 237
Query: 51 YNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQK 110
+ + N F C+ A + + T + R + + C++ QK
Sbjct: 238 KDNVVRVNGSDFQSCSVPWAAPVLTSGHDKIALTTYGRR------WYISGVANHCENGQK 399
Query: 111 LAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
L I V P + +P P P SP P P PEA+ P
Sbjct: 400 LFINVLP------------------------KQDGWYPAPSSPSASPSPSPVPAPEAAPP 507
Query: 171 ITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSA---TIHPLATSGHQGQVPILFSH 224
P+ S S P PT ++A H L HQ + +L+ H
Sbjct: 508 SN---APWAASVQTSELTWSPVPSPSPTPAPEAAPPSMHHGLLACRHQR*IGLLYPH 669
>TC91454 homologue to GP|1071881|dbj|BAA11125.1 orf324 {Beta vulgaris},
partial (3%)
Length = 667
Score = 43.5 bits (101), Expect = 7e-05
Identities = 39/129 (30%), Positives = 56/129 (43%), Gaps = 2/129 (1%)
Frame = +1
Query: 38 SIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTF 97
++GDS+ F + + + I Q+++ CN L + + + T H G FYFT
Sbjct: 169 TLGDSLLFLYPPSQDSLIQVTQESYKSCNTKDPILYMNNGNSLFNITSH----GDFYFTS 336
Query: 98 SNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTP--GPSSGGDIQSSPSFPWPFHPHQ 155
+ CQ +QK+ I V T EA+SP + P PSS S P P
Sbjct: 337 GENG--HCQKNQKIHISVGGTGNVDAEANSPSSSLPASAPSSQTVFGSIPVAP------- 489
Query: 156 GSSPGPAPT 164
SS P PT
Sbjct: 490 SSSNSPHPT 516
>BF519165 weakly similar to GP|11994144|dbj contains similarity to
nodulin~gene_id:K10D20.11 {Arabidopsis thaliana},
partial (13%)
Length = 561
Score = 41.6 bits (96), Expect = 3e-04
Identities = 35/114 (30%), Positives = 53/114 (45%), Gaps = 5/114 (4%)
Frame = +2
Query: 38 SIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTF 97
++GDS+ F + + + I Q+++ CN L + + + T H G FYFT
Sbjct: 218 TLGDSLLFLYPPSQDSLIQVTQESYKSCNTKDPILYMNNGNSLFNITSH----GDFYFTS 385
Query: 98 SNDSLKACQDSQKLAIKVTPTKASAPEASSPMPTTP--GPSSG---GDIQSSPS 146
+ CQ +QK+ I V T EA+SP + P PSS G I +PS
Sbjct: 386 GENG--HCQKNQKIHISVGGTGNVDAEANSPSSSLPASAPSSQTVFGSIPVAPS 541
>TC86144 weakly similar to GP|10727325|gb|AAF50995.2 CG15635 gene product
{Drosophila melanogaster}, partial (4%)
Length = 922
Score = 41.2 bits (95), Expect = 3e-04
Identities = 36/116 (31%), Positives = 53/116 (45%), Gaps = 7/116 (6%)
Frame = +3
Query: 62 FNLCNFTQANLL--TDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKVTPTK 119
F+ CN Q L+ T P+ + + R G FYF + C QKL++KV+ +
Sbjct: 321 FDNCNINQNGLVITTGPARVTLN------RTGDFYFICTIQG--HCSSGQKLSVKVSAST 476
Query: 120 ASAPEASSPM---PTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGP-APT-PEASSP 170
S P + P PT+ P+SG P P G +P P +PT P+A+ P
Sbjct: 477 PSPPSPTPPTSTPPTSTPPTSG---------TTPTPPTNGGTPSPSSPTGPDATPP 617
>TC79513 weakly similar to GP|9885806|gb|AAG01535.1| stellacyanin-like
protein CASLP1 precursor {Capsicum annuum}, partial
(33%)
Length = 1093
Score = 40.4 bits (93), Expect = 6e-04
Identities = 40/158 (25%), Positives = 65/158 (40%), Gaps = 15/158 (9%)
Frame = +3
Query: 23 TILVDGSS---EW-KNPTVSIGDSITFKHKQNYNLYIFKNQKAFNLCN--FTQANLLTDP 76
TI +G+S W N T ++GD++ F + + + A++ CN T L P
Sbjct: 504 TIPTNGASFYTNWASNKTFTVGDTLVFNYASGQHDVAKVTKTAYDSCNGANTLFTLTNSP 683
Query: 77 STTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLAIKV---------TPTKASAPEASS 127
+T + + T + F C QKL+I V PT +++P ++
Sbjct: 684 ATVTLNETGQQN--------FICAVPGHCSAGQKLSINVVKASASPVSAPTPSASPPKAT 839
Query: 128 PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTP 165
P PT S +++ P P +SP PAP P
Sbjct: 840 PAPTPVPAKSPAPTKAATPAP---APSTTASPTPAPAP 944
>TC85413 similar to PIR|S23737|S23737 proline-rich protein precursor -
kidney bean, partial (56%)
Length = 1139
Score = 39.3 bits (90), Expect = 0.001
Identities = 33/107 (30%), Positives = 41/107 (37%), Gaps = 9/107 (8%)
Frame = +1
Query: 122 APEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEAS-SPITVPL----- 175
A E + PT P ++ + + P H H SP PAPTP S SPI PL
Sbjct: 106 AEELETLTPTYPPHTTPAPLHPPANAPHHHHHHHIHSPTPAPTPTPSPSPIHTPLHPPYH 285
Query: 176 ---VPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP 219
VP K G + +P P P + PL H VP
Sbjct: 286 SAPVPAKPPTHGHHHHHPHPPAPTPVHTPVAPAHPPLHPPVHTPVVP 426
>TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (19%)
Length = 555
Score = 39.3 bits (90), Expect = 0.001
Identities = 29/71 (40%), Positives = 32/71 (44%)
Frame = +1
Query: 128 PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMPF 187
P P P P S G SP P P P GS P PAP P P + P P GS P
Sbjct: 199 PKPPPPAPPSPG----SPPKP-PAPPSPGSPPAPAPAPPPIPP-SPPAPPSPGSPPSPP- 357
Query: 188 INSNPAVPLPT 198
++PA P PT
Sbjct: 358 -PASPAPPKPT 387
Score = 35.0 bits (79), Expect = 0.025
Identities = 26/83 (31%), Positives = 33/83 (39%), Gaps = 2/83 (2%)
Frame = +1
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGS--SPGPAPTPEASSPITVP 174
P + P SP P P P S G + P P P + SPG P+P +SP
Sbjct: 205 PPPPAPPSPGSP-PKPPAPPSPGSPPAPAPAPPPIPPSPPAPPSPGSPPSPPPASPAPPK 381
Query: 175 LVPYKGSGDGMPFINSNPAVPLP 197
P KGS P +P+P
Sbjct: 382 PTPPKGS----------PLLPMP 420
>TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-protein {Pyrus
communis}, partial (30%)
Length = 956
Score = 39.3 bits (90), Expect = 0.001
Identities = 28/83 (33%), Positives = 36/83 (42%)
Frame = +3
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPL 175
TP A+ P A++P P TP P+ +SP P P +S P P +SSP
Sbjct: 276 TPPTATPPPAAAPTPATPAPA------TSPPAPTP------TSDAPTPDSTSSSP----- 404
Query: 176 VPYKGSGDGMPFINSNPAVPLPT 198
P G G P S A P P+
Sbjct: 405 -PAPGPGGPAPGPGSTDAPPSPS 470
>TC89498 weakly similar to PIR|A86218|A86218 protein T27G7.18 [imported] -
Arabidopsis thaliana, partial (57%)
Length = 767
Score = 38.5 bits (88), Expect = 0.002
Identities = 35/127 (27%), Positives = 49/127 (38%), Gaps = 19/127 (14%)
Frame = +1
Query: 38 SIGDSITFKHKQNYNLYIFKNQKAFNLCNFTQAN-------LLTDPSTTSCSYTWHPSRV 90
S+GD + F N++L N + C++ A DPS T HP V
Sbjct: 217 SLGDFLIFNTDNNHSLVQTYNFTTYKQCDYDDAQDKDTIQWSSVDPSNTDI----HPVTV 384
Query: 91 -------GFFYFTFSNDSLKACQDSQKLAIKVT-----PTKASAPEASSPMPTTPGPSSG 138
G YF S+ + C++ Q I VT P P SP P +P
Sbjct: 385 AVPLLKEGATYFFSSDYDGEQCKNGQHFKINVTHGQGLPKSLQKPSEDSPSPASP---IS 555
Query: 139 GDIQSSP 145
GD +S+P
Sbjct: 556 GDDESAP 576
>TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroelement pol
polyprotein {Oryza sativa} [Oryza sativa (japonica
cultivar-group)], partial (2%)
Length = 1235
Score = 38.1 bits (87), Expect = 0.003
Identities = 55/247 (22%), Positives = 86/247 (34%), Gaps = 29/247 (11%)
Frame = +3
Query: 9 FLILSLFFKLSHSTTILVDGSSEWKNPT---------------VSIGDSITFKHKQNYNL 53
F +L L + +V G W P+ +GDS+ F ++ +
Sbjct: 105 FCLLVLLMHKGDAYEFVVGGQKGWSAPSDPNANPYNQWAEKSRFQVGDSLVFNYQSGQDS 284
Query: 54 YI-FKNQKAFNLCNFTQANLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLA 112
I +Q+ + CN ++ + T + G YF N + C ++KL
Sbjct: 285 VIQVTSQQDYENCNTDASSEKSSDGHTVIKLI----KSGPHYFISGNKN--NCLQNEKLL 446
Query: 113 IKVTPTKA--------SAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPT 164
+ V + S P S + +P P S + P P P GSS P+P
Sbjct: 447 VIVLADRTNKNSNQTTSPPSPSPSVAPSPSPLSSHSSDALTPIPPP-SPLNGSSTPPSPV 623
Query: 165 PEASSPITVPLVPYKGSGDGMPFINSNPAVPLPTGEVDSATIHPLATSGHQGQVP----- 219
+ S P P P GS P + + P P G + I P + + P
Sbjct: 624 LDGSFP---PPSPLDGSTLTPPPVQQVGSSPPPLGTDVTNPITPTQSPVSEPPPPNAASS 794
Query: 220 ILFSHSC 226
IL S C
Sbjct: 795 ILVSFGC 815
>AJ498285 weakly similar to GP|20799090|em ADP-ribosylation factor guanine
nucleotide-exchange factor 6 {Homo sapiens}, partial
(2%)
Length = 568
Score = 37.7 bits (86), Expect = 0.004
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPIT 172
TP+ AS+PE+ P + PS + SPS PF SP +P+P ++
Sbjct: 145 TPSPASSPESLPPESPSQSPSMSPSMNMSPSMSPPFPTDASPSPASSPSPSTGESMS 315
Score = 29.3 bits (64), Expect = 1.3
Identities = 15/30 (50%), Positives = 17/30 (56%)
Frame = +1
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPS 146
PT AS ASSP P+T S + SSPS
Sbjct: 253 PTDASPSPASSPSPSTGESMSDNPVASSPS 342
>BF633206 similar to GP|21553614|gb| blue copper protein putative
{Arabidopsis thaliana}, partial (56%)
Length = 478
Score = 37.4 bits (85), Expect = 0.005
Identities = 35/145 (24%), Positives = 56/145 (38%), Gaps = 12/145 (8%)
Frame = +2
Query: 5 IFFYFLILSLFFKLSHSTTILVDGSSEW----------KNPTVSIGDSITFKHKQNYNLY 54
I L+ +L K +T V GS W T +GD + FK+ +++
Sbjct: 56 ILLALLLATLITKEVLATQHNVGGSQGWDPSSDFDSWSSGQTFKVGDQLVFKYTSMHSVV 235
Query: 55 IFKNQKAFNLCNFTQA--NLLTDPSTTSCSYTWHPSRVGFFYFTFSNDSLKACQDSQKLA 112
++ A+ C+ + +L T + G YFT +L C K+
Sbjct: 236 ELSDESAYKKCDISTPLNSLSTGKDVVKLD------KPGTRYFTCG--TLGHCDQGMKVK 391
Query: 113 IKVTPTKASAPEASSPMPTTPGPSS 137
I V S+ ASSP ++ PSS
Sbjct: 392 ITVGNGNGSSSTASSPSSSSSSPSS 466
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 37.4 bits (85), Expect = 0.005
Identities = 34/115 (29%), Positives = 49/115 (42%), Gaps = 11/115 (9%)
Frame = +3
Query: 116 TPTKASAPEASS-----PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSP 170
TP A++P A S P +P S ++ S+P P SSP A SSP
Sbjct: 378 TPPPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPPVSSP-----ASSPPTAAVSPVSSP 542
Query: 171 ITVPLV--PYKGSGDGMPFINSN----PAVPLPTGEVDSATIHPLATSGHQGQVP 219
+ P V P + S G+P ++ PA LP+ + S P ++S Q P
Sbjct: 543 VEAPSVSSPPEASSAGIPSSSATPADAPAATLPSSK--SPGTSPASSSPETSQGP 701
>BQ751233 similar to GP|18307448|em probable low-affinity hexose transporter
HXT3 {Neurospora crassa}, partial (30%)
Length = 659
Score = 37.0 bits (84), Expect = 0.006
Identities = 31/92 (33%), Positives = 43/92 (46%), Gaps = 3/92 (3%)
Frame = +1
Query: 116 TPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPE-ASSPIT-- 172
+PT ++A S P+P+ GP+S +S PS P P SS A P +SSP T
Sbjct: 367 SPTLSAAA*PSPPLPS--GPASVSSSRSRPSPPGTSSPSAVSSTVSASAPSPSSSPCTSP 540
Query: 173 VPLVPYKGSGDGMPFINSNPAVPLPTGEVDSA 204
PL P + P +S+P+ P V A
Sbjct: 541 SPLRPSSAASSSRPTSSSSPSASGPPRWVTGA 636
>AL385005 weakly similar to GP|21438547|emb Ratten cDNA {Rattus norvegicus},
partial (5%)
Length = 469
Score = 36.2 bits (82), Expect = 0.011
Identities = 24/68 (35%), Positives = 31/68 (45%), Gaps = 4/68 (5%)
Frame = +2
Query: 105 CQDSQKLAIKVTPTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGP--- 161
C QKL+I V S + SP ++P PS+G S+ P P P SS P
Sbjct: 11 CFAGQKLSINVGSGSNSPATSPSPYASSPSPSTGATPPSASGSPSPGSPVTPSSQSPGGS 190
Query: 162 -APTPEAS 168
+P PE S
Sbjct: 191 VSPPPENS 214
>TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (17%)
Length = 1031
Score = 35.8 bits (81), Expect = 0.014
Identities = 20/51 (39%), Positives = 24/51 (46%)
Frame = +1
Query: 128 PMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPY 178
P P P PSS PSFP+P G +P P P P S PL+P+
Sbjct: 73 PNPFQPPPSS-----PPPSFPFPPIVIPGLTPSPPPPPPPKSIFPAPLLPF 210
>BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (30%)
Length = 1338
Score = 35.8 bits (81), Expect = 0.014
Identities = 19/52 (36%), Positives = 25/52 (47%)
Frame = -1
Query: 123 PEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVP 174
P P P TP PSS + S P P+ F P + P P+P+P P+ P
Sbjct: 315 PAVPPPPPPTPPPSSPPALVSPPPRPFLFPPPP-NQPSPSPSPPPPPPLPPP 163
Score = 31.2 bits (69), Expect = 0.35
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 5/101 (4%)
Frame = -3
Query: 117 PTKASAPEASSPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLV 176
PT P P+P+ P P+ + S P P S P P A+S + +PL
Sbjct: 523 PTHQPHPCLPPPIPSAPPPTRAFALPPSAPPPSDLPPPPRSPTVIDPRPPAASAVPLPLR 344
Query: 177 PYKGSGDGMPF-----INSNPAVPLPTGEVDSATIHPLATS 212
P D P +P++ P+ + T PL++S
Sbjct: 343 PLYPLRDSPPRRPPPPPPHSPSILPPSSRLSPPTPLPLSSS 221
Score = 27.3 bits (59), Expect = 5.1
Identities = 27/99 (27%), Positives = 38/99 (38%)
Frame = -2
Query: 127 SPMPTTPGPSSGGDIQSSPSFPWPFHPHQGSSPGPAPTPEASSPITVPLVPYKGSGDGMP 186
+P P P P S SP P H H +P AP P +P + +
Sbjct: 1013 APSPGLPAPLSRYYTSLSPP---PLHTHDTLTPPSAPAP-------LPSISSRRLLRPTL 864
Query: 187 FINSNPAVPLPTGEVDSATIHPLATSGHQGQVPILFSHS 225
++S PA LP SA PL + P L++H+
Sbjct: 863 HLSSPPARALPILHRPSAVNPPL-----RDPTPSLYAHA 762
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,687,984
Number of Sequences: 36976
Number of extensions: 175924
Number of successful extensions: 2237
Number of sequences better than 10.0: 259
Number of HSP's better than 10.0 without gapping: 1922
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2115
length of query: 233
length of database: 9,014,727
effective HSP length: 93
effective length of query: 140
effective length of database: 5,575,959
effective search space: 780634260
effective search space used: 780634260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC126014.9


