

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126014.1 + phase: 0
(405 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
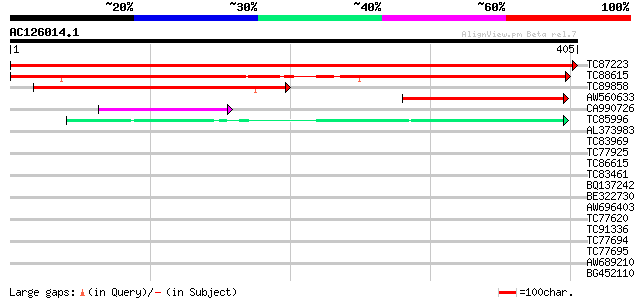
Score E
Sequences producing significant alignments: (bits) Value
TC87223 similar to GP|21536674|gb|AAM61006.1 unknown {Arabidopsi... 815 0.0
TC88615 similar to PIR|D96773|D96773 unknown protein F1M20.13 [i... 363 e-101
TC89858 weakly similar to PIR|D96773|D96773 unknown protein F1M2... 232 2e-61
AW560633 weakly similar to GP|14517472|gb At1g18740/F6A14_15 {Ar... 167 9e-42
CA990726 similar to PIR|T04778|T04 hypothetical protein F10M10.9... 46 3e-05
TC85996 weakly similar to GP|16648871|gb|AAL24287.1 Unknown prot... 45 4e-05
AL373983 similar to PIR|D96664|D966 hypothetical protein T12P18.... 40 0.002
TC83969 weakly similar to GP|16604342|gb|AAL24177.1 At2g46080/T3... 39 0.004
TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1 ... 32 0.34
TC86615 weakly similar to GP|8778409|gb|AAF79417.1| F16A14.5 {Ar... 31 0.75
TC83461 similar to PIR|T04778|T04778 hypothetical protein F10M10... 31 0.75
BQ137242 similar to GP|9800281|gb|A pR69 {rat cytomegalovirus Ma... 31 0.98
BE322730 30 1.3
AW696403 similar to GP|13346157|gb| bZIP protein DPBF4 {Arabidop... 30 1.7
TC77620 similar to GP|10129643|emb|CAC08239. Pspzf zinc finger p... 30 2.2
TC91336 similar to GP|20160854|dbj|BAB89793. hypothetical protei... 28 4.9
TC77694 homologue to PIR|T09704|T09704 probable arginine/serine-... 28 4.9
TC77695 similar to PIR|T09704|T09704 probable arginine/serine-ri... 28 4.9
AW689210 weakly similar to GP|20260444|gb| glycine-rich RNA-bind... 28 6.4
BG452110 28 6.4
>TC87223 similar to GP|21536674|gb|AAM61006.1 unknown {Arabidopsis
thaliana}, partial (69%)
Length = 1917
Score = 815 bits (2106), Expect = 0.0
Identities = 405/405 (100%), Positives = 405/405 (100%)
Frame = +1
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPH 60
MPATDYQGSSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPH
Sbjct: 247 MPATDYQGSSPSSLTNFGRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPH 426
Query: 61 DELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAI 120
DELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAI
Sbjct: 427 DELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAI 606
Query: 121 RDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHR 180
RDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHR
Sbjct: 607 RDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHR 786
Query: 181 NRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQAIGN 240
NRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQAIGN
Sbjct: 787 NRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQAIGN 966
Query: 241 NINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIPRSYTWAIP 300
NINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIPRSYTWAIP
Sbjct: 967 NINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSIPRSYTWAIP 1146
Query: 301 LLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEKEGEVRQKV 360
LLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEKEGEVRQKV
Sbjct: 1147LLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEEKEGEVRQKV 1326
Query: 361 HEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE 405
HEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE
Sbjct: 1327HEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLGRPNNE 1461
>TC88615 similar to PIR|D96773|D96773 unknown protein F1M20.13 [imported] -
Arabidopsis thaliana, partial (51%)
Length = 1693
Score = 363 bits (933), Expect = e-101
Identities = 202/409 (49%), Positives = 270/409 (65%), Gaps = 9/409 (2%)
Frame = +2
Query: 1 MPATDYQGSSPSSLTNFGRSILSFRREQIHSMEGS----TLEIELDSFQQHVTDRFVDLS 56
MP+T+ Q S SS ++FGRS+ R++Q+HS+E S + +EL SFQ+ VTDRF +LS
Sbjct: 317 MPSTENQSPSSSSFSSFGRSLFGMRQDQVHSVEASHESDSCNLELGSFQKRVTDRFNELS 496
Query: 57 SVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDV 116
DELLS+ W+ K+L F+ C +EF+AIL +K Q+ +PP+DRM SE+ ERSVKALD+
Sbjct: 497 GAGDDELLSIDWMQKLLTAFICCHDEFRAILLNNKEQLSKPPMDRMTSEFIERSVKALDI 676
Query: 117 CNAIRDGVEQIRVWQKLLEIVLYAL-DHQRSIGEGQFRRAKKALIDLSISMLDDGKESNA 175
CNA RDG+E IR WQK LEIV AL ++R + EGQFRRA+KAL+DL++SM+D+ KES +
Sbjct: 677 CNAGRDGIENIRGWQKQLEIVSCALGSNKRPLTEGQFRRARKALMDLALSMIDE-KESGS 853
Query: 176 SVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQL 235
+ R+RSFGR N +D H HS RS SWSVSR+WS +
Sbjct: 854 VFSQRHRSFGRHNSSKDS--HSSSTRHS---------------RSHSWSVSRSWSGS--- 973
Query: 236 QAIGNNINPPRAN----DLMATNGLAMSVYTMNSVLLFVMWALVAAIPCQDRGLNVHFSI 291
QA + P + D + +Y FV+ LVAAIPCQDRGLN+HF I
Sbjct: 974 QATTVHCK*PGSTSCK*DCCK*RAFSFCLYNEL*FSWFVLGTLVAAIPCQDRGLNIHFPI 1153
Query: 292 PRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDLVDSAHFPLTEE 351
P+ +TW+ + L++RI++ESKK++ +N+ GLL+EI QIE R ++DLVDSA FPLTEE
Sbjct: 1154PKQFTWSTAVASLYDRILDESKKKEHRNSSGLLKEIYQIETATRHLTDLVDSAQFPLTEE 1333
Query: 352 KEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSLG 400
EV Q + E+ +A + GLDPLER VREVF +IV S+TEGLDSLG
Sbjct: 1334HIMEVEQDLKELKLASEAFRVGLDPLERHVREVFRKIVDSRTEGLDSLG 1480
>TC89858 weakly similar to PIR|D96773|D96773 unknown protein F1M20.13
[imported] - Arabidopsis thaliana, partial (28%)
Length = 635
Score = 232 bits (592), Expect = 2e-61
Identities = 117/186 (62%), Positives = 147/186 (78%), Gaps = 3/186 (1%)
Frame = +3
Query: 18 GRSILSFRREQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFL 77
G SIL+ R Q+HSM+ S+L+ ELD FQ+HV+DRF +LSS+ +D LSL WV K+LD FL
Sbjct: 75 GSSILNLRATQVHSMQYSSLDKELDLFQKHVSDRFNELSSISNDNFLSLSWVRKLLDTFL 254
Query: 78 ICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIV 137
CQEEFK ILH ++ V +PPLDR+V++++ERSVKALDVCNAIRDGVEQIR W+K LEIV
Sbjct: 255 CCQEEFKMILHNNRSMVSKPPLDRLVNDFYERSVKALDVCNAIRDGVEQIRQWKKSLEIV 434
Query: 138 LYALDHQRSIGEGQFRRAKKALIDLSISMLD-DGKESN--ASVAHRNRSFGRSNGGRDRD 194
L ALDH+R IGEGQFRRAKKAL+DL I M+D K+SN +S +RNRSF R+N +D+D
Sbjct: 435 LCALDHKRIIGEGQFRRAKKALVDLEIGMIDASSKDSNNVSSFGNRNRSFSRNNVSKDKD 614
Query: 195 HHQHGN 200
Q G+
Sbjct: 615 SSQFGH 632
>AW560633 weakly similar to GP|14517472|gb At1g18740/F6A14_15 {Arabidopsis
thaliana}, partial (30%)
Length = 514
Score = 167 bits (422), Expect = 9e-42
Identities = 81/119 (68%), Positives = 99/119 (83%)
Frame = -1
Query: 281 QDRGLNVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDL 340
QDRGL+++FS+ R WA P++ LHERI+EESKKR+RKN CGLLREIQ+IEKC R M++L
Sbjct: 514 QDRGLHLNFSVLRQLPWASPVMSLHERILEESKKRERKNCCGLLREIQKIEKCARGMNEL 335
Query: 341 VDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
DS FPL+EEKE EVR KV +V VC++LKDGLDPL+ QVREVFHRIVR + EGLDS+
Sbjct: 334 ADSLEFPLSEEKEEEVRIKVEDVVIVCESLKDGLDPLKIQVREVFHRIVRGRMEGLDSV 158
>CA990726 similar to PIR|T04778|T04 hypothetical protein F10M10.90 -
Arabidopsis thaliana, partial (56%)
Length = 761
Score = 45.8 bits (107), Expect = 3e-05
Identities = 29/97 (29%), Positives = 49/97 (49%), Gaps = 1/97 (1%)
Frame = +2
Query: 64 LSLKWVGKMLDCFL-ICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRD 122
LS + ++ +C L + QE K IL K L +V EYF+ S+K LD CNA+
Sbjct: 194 LSFDSLKQVTECLLEMNQEVVKVILDCKKDIWKSQELFELVEEYFDNSLKTLDFCNALEK 373
Query: 123 GVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKAL 159
+++ R Q L+++ L D + G+ + + + L
Sbjct: 374 CLKRARDSQLLIDVALQKFDEETVSGDNCYVKTLQEL 484
>TC85996 weakly similar to GP|16648871|gb|AAL24287.1 Unknown protein
{Arabidopsis thaliana}, partial (53%)
Length = 1446
Score = 45.4 bits (106), Expect = 4e-05
Identities = 66/359 (18%), Positives = 125/359 (34%)
Frame = +2
Query: 41 LDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLD 100
L F+ + +R L DE+LSL W+ + + + ++ T V D
Sbjct: 248 LHGFEATLAERLRKLMPKSKDEILSLAWMTLAMKSLCETHNDIRTLI-TDLELPVSVWDD 424
Query: 101 RMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALI 160
+ V YF+ S K LD+CN + ++ L+ L+ L S F RA
Sbjct: 425 KWVDVYFDISTKLLDICNIFSSELSRLNQGNLPLKCALHNLGPASS---KSFVRA----- 580
Query: 161 DLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRS 220
S+LDD + R
Sbjct: 581 ---CSLLDDWR-----------------------------------------------RH 610
Query: 221 LSWSVSRTWSAARQLQAIGNNINPPRANDLMATNGLAMSVYTMNSVLLFVMWALVAAIPC 280
++ R + L + +++ P+ + L ++Y + FV AA
Sbjct: 611 INAKNPRIEKCSTILDGLVGSLDLPKVKNSAKGKALMQAMYGVKVETAFVCSVFSAAFSG 790
Query: 281 QDRGLNVHFSIPRSYTWAIPLLLLHERIMEESKKRDRKNACGLLREIQQIEKCVRVMSDL 340
+ L + +P ++WA + L + EE + R +L E++ ++ V+ +
Sbjct: 791 SSKKL-LDLDVPDMHSWAPAFISLQNLVNEEIRVRLSGGKFSVLIELEAVDAVVKELYPT 967
Query: 341 VDSAHFPLTEEKEGEVRQKVHEVSKVCDALKDGLDPLERQVREVFHRIVRSKTEGLDSL 399
+ + ++ + V E+ + L G+D L + V F ++ S+ L SL
Sbjct: 968 IQGGVNTEDKVEQESHLKTVEELGVAAEKLSQGMDLLAKGVDGFFQAVLTSRDTLLSSL 1144
>AL373983 similar to PIR|D96664|D966 hypothetical protein T12P18.5 [imported]
- Arabidopsis thaliana, partial (9%)
Length = 373
Score = 40.0 bits (92), Expect = 0.002
Identities = 17/28 (60%), Positives = 24/28 (85%)
Frame = +1
Query: 372 DGLDPLERQVREVFHRIVRSKTEGLDSL 399
+GL+PL+ Q+REVFHR+VR++TE L L
Sbjct: 1 EGLEPLQLQIREVFHRLVRTRTEFLHVL 84
>TC83969 weakly similar to GP|16604342|gb|AAL24177.1 At2g46080/T3F17.27
{Arabidopsis thaliana}, partial (38%)
Length = 634
Score = 38.9 bits (89), Expect = 0.004
Identities = 28/119 (23%), Positives = 54/119 (44%)
Frame = +2
Query: 41 LDSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLD 100
L F+ + +R +L DE++SL W+ + + K+++ T+ V +
Sbjct: 233 LSDFEATLEERLKNLIPKSKDEVVSLSWMTSAMQALCESHNDIKSLM-TNLELPVTDWDE 409
Query: 101 RMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVLYALDHQRSIGEGQFRRAKKAL 159
+ + Y + SVK LD+C A + ++ + LL+ L+ H S Q +A +L
Sbjct: 410 KWIDVYLDISVKLLDICIAFSSELSRLNQGRLLLQCTLH---HFGSFSSDQLFQAYSSL 577
>TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1
{Arabidopsis thaliana}, partial (47%)
Length = 1538
Score = 32.3 bits (72), Expect = 0.34
Identities = 16/50 (32%), Positives = 25/50 (50%)
Frame = +3
Query: 169 DGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQF 218
D + + HR RS N R+ HH +GNS+ N+ + HR+ G +
Sbjct: 156 DSYDDRHTDRHRRRSISPEN--RNPRHHDNGNSNGNHLPKKFGHRNNGSY 299
>TC86615 weakly similar to GP|8778409|gb|AAF79417.1| F16A14.5 {Arabidopsis
thaliana}, partial (31%)
Length = 1278
Score = 31.2 bits (69), Expect = 0.75
Identities = 16/52 (30%), Positives = 31/52 (58%)
Frame = +1
Query: 179 HRNRSFGRSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWS 230
HR+R +G + +R H+HG H+N N + ++R+ ++R+ +W + WS
Sbjct: 670 HRHR-YGHGHSNWNRHGHRHG--HTNWNRHRNRYRNWNRYRNWNW-YNHRWS 813
>TC83461 similar to PIR|T04778|T04778 hypothetical protein F10M10.90 -
Arabidopsis thaliana, partial (25%)
Length = 663
Score = 31.2 bits (69), Expect = 0.75
Identities = 22/98 (22%), Positives = 46/98 (46%), Gaps = 1/98 (1%)
Frame = +3
Query: 42 DSFQQHVTDRFVDLSSVPHDELLSLKWVGKMLDCFLICQEEF-KAILHTHKGQVVRPPLD 100
+S ++H L++ LS + ++ D L E K IL + L
Sbjct: 267 ESIKEHTNRVISSLATGVEVRTLSFNSLREVTDSLLEMNHEVVKVILDCKRDIWGNKDLF 446
Query: 101 RMVSEYFERSVKALDVCNAIRDGVEQIRVWQKLLEIVL 138
+V++YF+ S++ L+ CN++ + + R Q +++ V+
Sbjct: 447 ALVNDYFDNSLQTLEFCNSLEKCLRRARENQVMVKSVI 560
>BQ137242 similar to GP|9800281|gb|A pR69 {rat cytomegalovirus Maastricht},
partial (2%)
Length = 1023
Score = 30.8 bits (68), Expect = 0.98
Identities = 13/51 (25%), Positives = 23/51 (44%)
Frame = +2
Query: 186 RSNGGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLSWSVSRTWSAARQLQ 236
RS G H +H ++H Y G R SW ++TW+ +++ +
Sbjct: 410 RSTGATATSHRRHSDTH------VYSENGGGGRREASWDATQTWTKSQETE 544
>BE322730
Length = 414
Score = 30.4 bits (67), Expect = 1.3
Identities = 21/72 (29%), Positives = 38/72 (52%), Gaps = 1/72 (1%)
Frame = +3
Query: 69 VGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRM-VSEYFERSVKALDVCNAIRDGVEQI 127
+ ++LD + Q K +L++ R DR V +Y E +V+ LDVCN + +E I
Sbjct: 183 LAQLLDASITTQ---KIVLNSLVNVSYRDDSDRGDVDKYLEDNVEILDVCNYFVEKIESI 353
Query: 128 RVWQKLLEIVLY 139
+ L++V++
Sbjct: 354 NNYVDTLKMVVH 389
>AW696403 similar to GP|13346157|gb| bZIP protein DPBF4 {Arabidopsis
thaliana}, partial (28%)
Length = 587
Score = 30.0 bits (66), Expect = 1.7
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Frame = +2
Query: 23 SFRREQIHSM--EGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLKW 68
S +R Q+HS+ + S + LD Q + D LSS+ DELL W
Sbjct: 224 SGKRSQLHSLVRQNSVYSLTLDEVQNQLGDLGKPLSSMNLDELLKNVW 367
>TC77620 similar to GP|10129643|emb|CAC08239. Pspzf zinc finger protein-like
{Arabidopsis thaliana}, partial (26%)
Length = 1958
Score = 29.6 bits (65), Expect = 2.2
Identities = 15/41 (36%), Positives = 26/41 (62%)
Frame = +3
Query: 27 EQIHSMEGSTLEIELDSFQQHVTDRFVDLSSVPHDELLSLK 67
EQIH +E + L+ F QH D +D+ ++ ++ELL+L+
Sbjct: 1362 EQIHLLESNLFLTGLNFFDQH-RDMRLDIDNMSYEELLALE 1481
>TC91336 similar to GP|20160854|dbj|BAB89793. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (12%)
Length = 677
Score = 28.5 bits (62), Expect = 4.9
Identities = 36/131 (27%), Positives = 49/131 (36%), Gaps = 22/131 (16%)
Frame = +1
Query: 138 LYALDHQRSIGEGQFRRAKKALIDLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDH-- 195
L + HQR GE K+ D DDG SNAS N+ F NG R+
Sbjct: 55 LLTVAHQRGSGEFTGYTVKREGDDHEGGS-DDGGSSNASQLDFNQMFSGYNGDRELSEMV 231
Query: 196 ----HQHGNSHSNNNINTYQHR-------SMGQFRSLSWSVSRTWSA---ARQLQAIGNN 241
H + + + N + R S G S S S +WS R ++ G+N
Sbjct: 232 SALTHVASSGSNQMSSNEWIQRSDFPLISSFGNASSSSSYASSSWSGQKRGRDEESSGSN 411
Query: 242 ------INPPR 246
+ PPR
Sbjct: 412 QYITQAVVPPR 444
>TC77694 homologue to PIR|T09704|T09704 probable arginine/serine-rich
splicing factor - alfalfa, partial (95%)
Length = 1494
Score = 28.5 bits (62), Expect = 4.9
Identities = 16/40 (40%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Frame = +3
Query: 168 DDGKESNASVAHRNRSFGRS----NGGRDRDHHQHGNSHS 203
DD KE + R+RS+ R + GRDRDH + S S
Sbjct: 423 DDYKEKDYRRRSRSRSYDRHERDRHRGRDRDHRRRSRSRS 542
>TC77695 similar to PIR|T09704|T09704 probable arginine/serine-rich splicing
factor - alfalfa, partial (83%)
Length = 1657
Score = 28.5 bits (62), Expect = 4.9
Identities = 21/68 (30%), Positives = 32/68 (46%), Gaps = 5/68 (7%)
Frame = +1
Query: 168 DDGKESNASVAHRNRSFGRSN-----GGRDRDHHQHGNSHSNNNINTYQHRSMGQFRSLS 222
DD ++ R+RS+ R GGRDRD+ + S S + Y+ R G++
Sbjct: 448 DDYRDKGYRRRSRSRSYDRYERDRYRGGRDRDYRRRSRSRSAS--PDYKRRGRGRYDDER 621
Query: 223 WSVSRTWS 230
S SR+ S
Sbjct: 622 RSRSRSRS 645
>AW689210 weakly similar to GP|20260444|gb| glycine-rich RNA-binding protein
putative {Arabidopsis thaliana}, partial (25%)
Length = 633
Score = 28.1 bits (61), Expect = 6.4
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = +1
Query: 153 RRAKKALIDLSISMLDDGKESNASVAHRNRSFGRSNGGRDRDHHQHGNSHSNNN 206
R A A+ D++ ++ V R GRSN GR+R+ + H N N +
Sbjct: 235 RSAIDAINDMNGRTINGRVVKVNGVKSRGGGSGRSNFGRERERYYHHNDERNGD 396
>BG452110
Length = 490
Score = 28.1 bits (61), Expect = 6.4
Identities = 28/104 (26%), Positives = 43/104 (40%)
Frame = +1
Query: 67 KWVGKMLDCFLICQEEFKAILHTHKGQVVRPPLDRMVSEYFERSVKALDVCNAIRDGVEQ 126
K + +LD +L+ EE A +VVR LDR S Y + + + Q
Sbjct: 124 KNISHLLDNYLVI-EELSATKEEVANEVVREALDRAPSGYIKWRPDPI---------IGQ 273
Query: 127 IRVWQKLLEIVLYALDHQRSIGEGQFRRAKKALIDLSISMLDDG 170
+ V +LL L + S+ E + R KK S +D+G
Sbjct: 274 LIVNNRLLGEANKGLSARMSLLESETREMKKLYGSAQSSYVDEG 405
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,928,678
Number of Sequences: 36976
Number of extensions: 184267
Number of successful extensions: 1097
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1058
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1080
length of query: 405
length of database: 9,014,727
effective HSP length: 98
effective length of query: 307
effective length of database: 5,391,079
effective search space: 1655061253
effective search space used: 1655061253
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126014.1


