

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126010.6 + phase: 0
(518 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
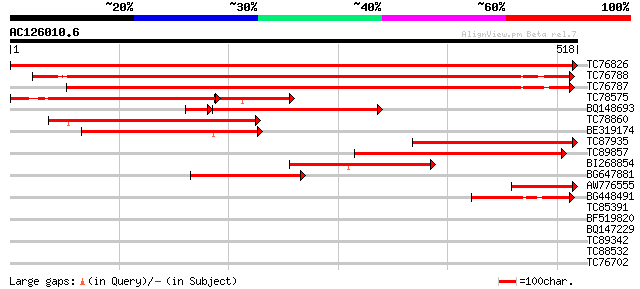
Score E
Sequences producing significant alignments: (bits) Value
TC76826 homologue to SP|P34899|GLYM_PEA Serine hydroxymethyltran... 1026 0.0
TC76788 similar to PIR|B71400|B71400 glycine hydroxymethyltransf... 535 e-152
TC76787 homologue to PIR|B71400|B71400 glycine hydroxymethyltran... 531 e-151
TC78575 homologue to GP|14334888|gb|AAK59622.1 putative glycine ... 284 e-100
BQ148693 homologue to SP|P34899|GLYM Serine hydroxymethyltransfe... 305 1e-93
TC78860 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220 ... 258 4e-69
BE319174 homologue to SP|P34899|GLYM Serine hydroxymethyltransfe... 250 1e-66
TC87935 similar to GP|15809972|gb|AAL06913.1 AT4g37930/F20D10_50... 249 1e-66
TC89857 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220 ... 204 7e-53
BI268854 similar to PIR|F86484|F86 probable hydroxymethyltransfe... 159 2e-39
BG647881 similar to GP|19386847|db putative hydroxymethyltransfe... 122 4e-28
AW776555 similar to SP|P34899|GLYM_ Serine hydroxymethyltransfer... 96 3e-20
BG448491 weakly similar to GP|8388673|emb| (mitochondrial?) seri... 79 6e-15
TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chape... 33 0.27
BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {A... 31 1.0
BQ147229 similar to GP|17064942|gb Unknown protein {Arabidopsis ... 30 2.9
TC89342 similar to GP|10178056|dbj|BAB11420. contains similarity... 28 8.6
TC88532 similar to PIR|S45033|S45033 probable imbibition protein... 28 8.6
TC76702 similar to SP|Q9LZG0|ADK2_ARATH Adenosine kinase 2 (EC 2... 28 8.6
>TC76826 homologue to SP|P34899|GLYM_PEA Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
complete
Length = 1942
Score = 1026 bits (2652), Expect = 0.0
Identities = 518/518 (100%), Positives = 518/518 (100%)
Frame = +2
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID
Sbjct: 101 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 280
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID
Sbjct: 281 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 460
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS
Sbjct: 461 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 640
Query: 181 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 240
HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD
Sbjct: 641 HGYQTDTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYD 820
Query: 241 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 300
YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR
Sbjct: 821 YARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFR 1000
Query: 301 KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 360
KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV
Sbjct: 1001KGLKEVNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQV 1180
Query: 361 LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 420
LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG
Sbjct: 1181LSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPG 1360
Query: 421 DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 480
DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL
Sbjct: 1361DVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETL 1540
Query: 481 QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK
Sbjct: 1541QSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 1654
>TC76788 similar to PIR|B71400|B71400 glycine hydroxymethyltransferase (EC
2.1.2.1) - Arabidopsis thaliana, complete
Length = 1871
Score = 535 bits (1377), Expect = e-152
Identities = 277/497 (55%), Positives = 350/497 (69%), Gaps = 2/497 (0%)
Frame = +2
Query: 22 FSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEIDPEIADIIELEKARQWKGLELI 81
F S ++ S P + D + W N+ L +DPEI D+IE EK RQ +G+ELI
Sbjct: 92 FFLSFFFHSDSFPTFPIMDPVSE---WG---NTPLVTVDPEIHDLIEKEKRRQCRGIELI 253
Query: 82 PSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKW 141
SENFTS +V++A+GS +TNKYSEG PG RYYGGNE+ID E LC+ RAL+AF +DP W
Sbjct: 254 ASENFTSFAVIEALGSALTNKYSEGMPGNRYYGGNEFIDQIENLCRSRALQAFHIDPQSW 433
Query: 142 GVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTD-TKKISAVSIFFETM 200
GVNVQP SGSP+NF YTA+L PHDRIM LDLP GGHL+HGY T KKISA SI+FE++
Sbjct: 434 GVNVQPYSGSPANFAAYTAVLNPHDRIMGLDLPSGGHLTHGYYTSGGKKISATSIYFESL 613
Query: 201 PYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIRKVCDKQKAVMLADMA 260
PY+++ +TG+IDYD+LE+ A FRP+LI+ G SAY R +DY R R V DK A++L DMA
Sbjct: 614 PYKVNSTTGFIDYDRLEEKALDFRPRLIICGGSAYPRDWDYKRFRDVADKCGALLLCDMA 793
Query: 261 HISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNK-QGKEVFYDYED 319
H SGLVAA + +PF+Y D+VTTTTHKSLRGPR MIF+RKG K K Q + YD+ED
Sbjct: 794 HFSGLVAAQEVNNPFEYCDIVTTTTHKSLRGPRAGMIFYRKGPKPPKKGQPENAVYDFED 973
Query: 320 KINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKGYELV 379
KIN AVFP LQGGPHNH I LAVALKQA +P +KAY +QV +N L KGY LV
Sbjct: 974 KINFAVFPSLQGGPHNHQIGALAVALKQAMSPGFKAYAKQVKANAVAIGNYLMSKGYSLV 1153
Query: 380 SGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTS 439
+GGTENHLVL +L+ G+ G++VEK+ + +I NKN V GD SA+ PGG+R+G PA+TS
Sbjct: 1154TGGTENHLVLWDLRPLGLTGNKVEKLCDLCNITVNKNAVFGDSSALAPGGVRVGAPAMTS 1333
Query: 440 RGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEE 499
RG VE+DF K+ E+ +V L L+I+ E G LKDF + L + I++L+ DVE+
Sbjct: 1334RGLVEKDFEKIGEFLHRAVTLTLEIQKE-HGKLLKDFNKGLVDN----KAIAELKADVEK 1498
Query: 500 FAKQFPTIGFEKSSMKY 516
F+ F GF S MKY
Sbjct: 1499FSSLFGMPGFLVSEMKY 1549
>TC76787 homologue to PIR|B71400|B71400 glycine hydroxymethyltransferase (EC
2.1.2.1) - Arabidopsis thaliana, complete
Length = 1846
Score = 531 bits (1369), Expect = e-151
Identities = 272/466 (58%), Positives = 340/466 (72%), Gaps = 2/466 (0%)
Frame = +2
Query: 53 NSSLEEIDPEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARY 112
N+ L +DPEI D+IE EK RQ +G+ELI SENFTS +V++A+GS +TNKYSEG PG RY
Sbjct: 137 NTPLVTVDPEIHDLIEKEKRRQCRGIELIASENFTSFAVIEALGSALTNKYSEGMPGNRY 316
Query: 113 YGGNEYIDMAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALD 172
YGGNE+ID E +C+ RAL AF LD A WGVNVQP SGSP+NF YTA+L PHDRIM LD
Sbjct: 317 YGGNEFIDQIENICRSRALTAFHLDAATWGVNVQPYSGSPANFAAYTAVLNPHDRIMGLD 496
Query: 173 LPHGGHLSHGYQTD-TKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAG 231
LP GGHL+HGY T KKISA SI+FE++PY+++ +TG+IDYD+LE+ A FRPKLI+ G
Sbjct: 497 LPSGGHLTHGYYTSGGKKISATSIYFESLPYKVNSTTGFIDYDRLEEKALDFRPKLIICG 676
Query: 232 ASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRG 291
SAY R +DY R R+V DK A++L DMAHISGLVAA PF + D+VTTTTHKSLRG
Sbjct: 677 GSAYPRDWDYGRFRQVADKCGALLLCDMAHISGLVAAQEANDPFAFCDIVTTTTHKSLRG 856
Query: 292 PRGAMIFFRKGLKEVNK-QGKEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATT 350
PR MIF+RKG K K Q + YD+EDKIN AVFP LQGGPHNH I LAVALKQATT
Sbjct: 857 PRAGMIFYRKGPKPPKKGQPENAVYDFEDKINFAVFPSLQGGPHNHQIGALAVALKQATT 1036
Query: 351 PEYKAYQEQVLSNCAKFAQALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVH 410
P +KAY +QV +N L KGY LV+GGTENHLVL +L+ G+ G++VEK+ + +
Sbjct: 1037PGFKAYAKQVKANAVALGNYLISKGYSLVTGGTENHLVLWDLRPLGLTGNKVEKLCDLCN 1216
Query: 411 IAANKNTVPGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKG 470
I NKN V GD SA+ PGG+R+G PA+TSRG VE+DF ++AE+ +V L L+I+ E G
Sbjct: 1217ITVNKNAVFGDSSALAPGGVRIGAPAMTSRGLVEKDFEQIAEFLHRAVTLTLEIQKE-YG 1393
Query: 471 TKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
LKDF + L ++ ++ L+ DVE+F+ F GF S +KY
Sbjct: 1394KLLKDFNKGLVNNKALED----LKADVEKFSASFDMPGFLVSELKY 1519
>TC78575 homologue to GP|14334888|gb|AAK59622.1 putative glycine
hydroxymethyltransferase {Arabidopsis thaliana}, partial
(43%)
Length = 821
Score = 284 bits (727), Expect(2) = e-100
Identities = 146/192 (76%), Positives = 157/192 (81%)
Frame = +1
Query: 1 MAMAMALRRLSSSINKSSRPLFSASSVYYKSSLPDEAVYDKENSRVSWPKQLNSSLEEID 60
+ MAMALRRL+S+INK + S+Y SS + S W KQLN L +D
Sbjct: 58 ITMAMALRRLTSTINKPT-------SLYRLSS--SLSAQHTHKSHPDWIKQLNDPLGVVD 210
Query: 61 PEIADIIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYID 120
PEI DIIELEKARQWKGLELIPSENFTSLSVMQAVGS+MTNKYSEGYPGARYYGGNEYID
Sbjct: 211 PEIEDIIELEKARQWKGLELIPSENFTSLSVMQAVGSVMTNKYSEGYPGARYYGGNEYID 390
Query: 121 MAETLCQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLS 180
MAETLCQKRALE F LDP +WGVNVQ LSGSPSNF VYTALLKPH+RIMALDLPHGGHLS
Sbjct: 391 MAETLCQKRALETFGLDPTQWGVNVQSLSGSPSNFQVYTALLKPHERIMALDLPHGGHLS 570
Query: 181 HGYQTDTKKISA 192
HGYQTDTKKIS+
Sbjct: 571 HGYQTDTKKISS 606
Score = 99.8 bits (247), Expect(2) = e-100
Identities = 54/76 (71%), Positives = 59/76 (77%), Gaps = 3/76 (3%)
Frame = +2
Query: 188 KKISAVSIFFETMPYRLDESTGYI---DYDQLEKSATLFRPKLIVAGASAYARLYDYARI 244
++ AVSIFFETMPY++ Q+EKSA LFRPKLIVAGASAYARLYDYARI
Sbjct: 593 RRYQAVSIFFETMPYKVG*EHWLY*L*PGRQMEKSAALFRPKLIVAGASAYARLYDYARI 772
Query: 245 RKVCDKQKAVMLADMA 260
RKVCDKQKAVM ADMA
Sbjct: 773 RKVCDKQKAVMFADMA 820
>BQ148693 homologue to SP|P34899|GLYM Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (34%)
Length = 545
Score = 305 bits (781), Expect(2) = 1e-93
Identities = 149/155 (96%), Positives = 153/155 (98%)
Frame = +1
Query: 186 DTKKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFRPKLIVAGASAYARLYDYARIR 245
DTKKISAVSIFFETMPYRLDESTGYIDYDQ+EKSA LFRPKLIVAGASAYARLYDYARIR
Sbjct: 79 DTKKISAVSIFFETMPYRLDESTGYIDYDQMEKSAALFRPKLIVAGASAYARLYDYARIR 258
Query: 246 KVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKE 305
KVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKE
Sbjct: 259 KVCDKQKAVMLADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKE 438
Query: 306 VNKQGKEVFYDYEDKINQAVFPGLQGGPHNHTITG 340
+NK+G+EV YDYEDKINQAVFPGLQGGPHNHTITG
Sbjct: 439 INKKGQEVLYDYEDKINQAVFPGLQGGPHNHTITG 543
Score = 57.0 bits (136), Expect(2) = 1e-93
Identities = 24/25 (96%), Positives = 25/25 (100%)
Frame = +3
Query: 161 LLKPHDRIMALDLPHGGHLSHGYQT 185
LLKPH+RIMALDLPHGGHLSHGYQT
Sbjct: 3 LLKPHERIMALDLPHGGHLSHGYQT 77
>TC78860 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220
{Arabidopsis thaliana}, partial (33%)
Length = 915
Score = 258 bits (659), Expect = 4e-69
Identities = 129/206 (62%), Positives = 152/206 (73%), Gaps = 12/206 (5%)
Frame = +3
Query: 36 EAVYDKENSRVSWPKQL------------NSSLEEIDPEIADIIELEKARQWKGLELIPS 83
EA + K +S S P L + L E DP++ II EK RQ++ LELI S
Sbjct: 294 EATFGKSSSSSSSPSSLPKKIGGDGSSFLDYGLSEADPDVHAIINKEKDRQFRSLELIAS 473
Query: 84 ENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETLCQKRALEAFRLDPAKWGV 143
ENFTS +VM+AVGS +TNKYSEG PG RYYGGNE+ID E LCQ+RAL AF LD KWGV
Sbjct: 474 ENFTSKAVMEAVGSCLTNKYSEGLPGKRYYGGNEHIDELEILCQQRALAAFHLDGDKWGV 653
Query: 144 NVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQTDTKKISAVSIFFETMPYR 203
NVQPLSGSP+NF VYTA+LKPHDRIM LDLPHGGHLSHG+ T +++S SI+FE+MPYR
Sbjct: 654 NVQPLSGSPANFAVYTAILKPHDRIMGLDLPHGGHLSHGFMTAKRRVSGTSIYFESMPYR 833
Query: 204 LDESTGYIDYDQLEKSATLFRPKLIV 229
LDESTG IDYD LEK+A LFRP ++
Sbjct: 834 LDESTGVIDYDMLEKTAALFRPNSLL 911
>BE319174 homologue to SP|P34899|GLYM Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (29%)
Length = 537
Score = 250 bits (638), Expect = 1e-66
Identities = 130/174 (74%), Positives = 137/174 (78%), Gaps = 8/174 (4%)
Frame = +1
Query: 66 IIELEKARQWKGLELIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETL 125
++ L K +G LIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETL
Sbjct: 16 LLSLRKLDNGRGWNLIPSENFTSLSVMQAVGSIMTNKYSEGYPGARYYGGNEYIDMAETL 195
Query: 126 CQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQ- 184
CQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQ
Sbjct: 196 CQKRALEAFRLDPAKWGVNVQPLSGSPSNFHVYTALLKPHDRIMALDLPHGGHLSHGYQL 375
Query: 185 ------TDTKKISAVSIFFETMPYRLDESTG-YIDYDQLEKSATLFRPKLIVAG 231
K+ FFETMPYRLDE ++ SAT F+ KLIVAG
Sbjct: 376 LSSLCRPTPKRYLQSLYFFETMPYRLDEKHRIHLIMTS*RNSATPFKAKLIVAG 537
Score = 39.7 bits (91), Expect = 0.003
Identities = 18/18 (100%), Positives = 18/18 (100%)
Frame = +3
Query: 62 EIADIIELEKARQWKGLE 79
EIADIIELEKARQWKGLE
Sbjct: 3 EIADIIELEKARQWKGLE 56
>TC87935 similar to GP|15809972|gb|AAL06913.1 AT4g37930/F20D10_50
{Arabidopsis thaliana}, partial (29%)
Length = 884
Score = 249 bits (637), Expect = 1e-66
Identities = 124/150 (82%), Positives = 138/150 (91%)
Frame = +2
Query: 369 QALSEKGYELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPG 428
Q+L EKGY+LVSGGTENHLVLVNL+NKGIDGSRVEKVLE+VHIAANKNTVPGDVSAMVPG
Sbjct: 68 QSLLEKGYDLVSGGTENHLVLVNLRNKGIDGSRVEKVLESVHIAANKNTVPGDVSAMVPG 247
Query: 429 GIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSYVQS 488
GIRMGTPALTSRGFVE+DF KVAEYFDA+V +AL+IK SKGTKLKDFVE ++S S VQS
Sbjct: 248 GIRMGTPALTSRGFVEDDFKKVAEYFDAAVKIALQIKENSKGTKLKDFVEAMESDSQVQS 427
Query: 489 EISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
+I+ LRHDVE +AKQFPTIGFE +MKYNK
Sbjct: 428 QIADLRHDVEGYAKQFPTIGFEIETMKYNK 517
>TC89857 similar to GP|14030719|gb|AAK53034.1 AT4g32520/F8B4_220
{Arabidopsis thaliana}, partial (34%)
Length = 952
Score = 204 bits (519), Expect = 7e-53
Identities = 101/194 (52%), Positives = 138/194 (71%), Gaps = 1/194 (0%)
Frame = +1
Query: 316 DYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQALSEKG 375
D E IN AVFPGLQGGPHNHTI GLAV LK A +P++K YQ QV++NC A L E
Sbjct: 40 DLESAINNAVFPGLQGGPHNHTIGGLAVCLKYAQSPDFKNYQNQVVANCRALANRLVEHE 219
Query: 376 YELVSGGTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTP 435
Y+LVSGG++NHLVLV+L+ GIDG+RVEK+L+ I NKN+VPGD SA+VPGGIR+G+P
Sbjct: 220 YKLVSGGSDNHLVLVDLRPSGIDGARVEKILDMASITLNKNSVPGDKSALVPGGIRIGSP 399
Query: 436 ALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSSSY-VQSEISKLR 494
A+T+RG E++F +A+ V ++L+ K+ GTK++DF+ + + + + ++S LR
Sbjct: 400 AMTTRGLGEKEFELIADLIHEGVRISLEAKSLVSGTKVQDFLNFVLAPEFPLGDKVSNLR 579
Query: 495 HDVEEFAKQFPTIG 508
VE A Q+P G
Sbjct: 580 RKVEALATQYPIPG 621
>BI268854 similar to PIR|F86484|F86 probable hydroxymethyltransferase
49598-47322 [imported] - Arabidopsis thaliana, partial
(22%)
Length = 429
Score = 159 bits (403), Expect = 2e-39
Identities = 83/140 (59%), Positives = 99/140 (70%), Gaps = 6/140 (4%)
Frame = +3
Query: 256 LADMAHISGLVAAGVIPSPFDYADVVTTTTHKSLRGPRGAMIFFRKGLKEVNK-----QG 310
L DMA ISG++AA +PFDY DVVT+TTHKSLRGPRG +IF+RKG K + QG
Sbjct: 3 LCDMAQISGIIAAKECVNPFDYCDVVTSTTHKSLRGPRGGIIFYRKGTKPRKRGILLTQG 182
Query: 311 KEVF-YDYEDKINQAVFPGLQGGPHNHTITGLAVALKQATTPEYKAYQEQVLSNCAKFAQ 369
E YD+E+KIN AVFP LQGGPHN+ I LA+ALKQ TPEYKAY +QV N A
Sbjct: 183 HESDQYDFEEKINFAVFPSLQGGPHNNHIAALAIALKQVATPEYKAYMQQVKXNAQALAS 362
Query: 370 ALSEKGYELVSGGTENHLVL 389
AL + LV+GGT+NHL+L
Sbjct: 363 ALLRRKCPLVTGGTDNHLIL 422
>BG647881 similar to GP|19386847|db putative hydroxymethyltransferase {Oryza
sativa (japonica cultivar-group)}, partial (17%)
Length = 793
Score = 122 bits (305), Expect = 4e-28
Identities = 59/106 (55%), Positives = 78/106 (72%), Gaps = 1/106 (0%)
Frame = +1
Query: 166 DRIMALDLPHGGHLSHGYQTDT-KKISAVSIFFETMPYRLDESTGYIDYDQLEKSATLFR 224
DRIM LD P GG+ SHGY T KK+S SIFFE++ Y+++ +G+IDYD+LE+ A FR
Sbjct: 4 DRIMGLDTPSGGNTSHGYYTPNGKKVSGASIFFESLAYKINPQSGFIDYDKLEERALDFR 183
Query: 225 PKLIVAGASAYARLYDYARIRKVCDKQKAVMLADMAHISGLVAAGV 270
PK+++ G S+Y R +DYAR R V DK AV+L DMA ISG++AA V
Sbjct: 184 PKILICGGSSYPREWDYARFRHVADKCGAVLLCDMAQISGIIAAKV 321
>AW776555 similar to SP|P34899|GLYM_ Serine hydroxymethyltransferase
mitochondrial precursor (EC 2.1.2.1) (Serine methylase),
partial (10%)
Length = 394
Score = 95.9 bits (237), Expect = 3e-20
Identities = 52/60 (86%), Positives = 53/60 (87%)
Frame = -1
Query: 459 NLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKYNK 518
NLALKIKAESKGTKLK FV+TLQSSS QSEISKL HDVEEFAKQFP IGFEKSSMK K
Sbjct: 388 NLALKIKAESKGTKLKGFVKTLQSSS*GQSEISKLCHDVEEFAKQFPPIGFEKSSMKAPK 209
>BG448491 weakly similar to GP|8388673|emb| (mitochondrial?) serine
hydroxymethyltransferase {Leishmania major}, partial
(15%)
Length = 467
Score = 78.6 bits (192), Expect = 6e-15
Identities = 42/94 (44%), Positives = 60/94 (63%)
Frame = +2
Query: 423 SAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQS 482
SA+ PGG+R+GTPALTSRGFVE DF VAE+ D ++ LA+ I+ K L +F + ++
Sbjct: 86 SALTPGGVRLGTPALTSRGFVESDFETVAEFLDRTLKLAIVIQQTHK--TLAEFKKAVED 259
Query: 483 SSYVQSEISKLRHDVEEFAKQFPTIGFEKSSMKY 516
+++ LR DVE FA +F G + S +KY
Sbjct: 260 ----HADVKALRRDVEAFATKFYMPGIDVSKLKY 349
>TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chaperone BiP-A
- soybean, partial (52%)
Length = 1231
Score = 33.1 bits (74), Expect = 0.27
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Frame = +1
Query: 454 FDASVNLALKIKAESKGT-KLKDFVETLQSSSYVQSEISKLRHDVEEFAKQ 503
F+ N L +KAE KGT K + T + Q EI ++ + EEFA++
Sbjct: 568 FEVDANGILNVKAEDKGTGKSEKITITNEKGRLSQEEIERMVREAEEFAEE 720
>BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {Arabidopsis
thaliana}, partial (34%)
Length = 574
Score = 31.2 bits (69), Expect = 1.0
Identities = 14/35 (40%), Positives = 19/35 (54%)
Frame = +1
Query: 311 KEVFYDYEDKINQAVFPGLQGGPHNHTITGLAVAL 345
K V YD+ + V+P L GGP N+ G +V L
Sbjct: 361 KSVMYDFNENRVVRVYPELTGGPRNYPSAGSSVML 465
>BQ147229 similar to GP|17064942|gb Unknown protein {Arabidopsis thaliana},
partial (38%)
Length = 649
Score = 29.6 bits (65), Expect = 2.9
Identities = 26/102 (25%), Positives = 43/102 (41%)
Frame = +2
Query: 382 GTENHLVLVNLKNKGIDGSRVEKVLEAVHIAANKNTVPGDVSAMVPGGIRMGTPALTSRG 441
GT +LV +++ G D ++ L HI N V + VP P++ + G
Sbjct: 299 GTVVAYLLVPMRSLGQDSWKIAAALMGRHIGGAVNYVAISDALGVP-------PSILAAG 457
Query: 442 FVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVETLQSS 483
++ + A YF LA K+ ES + D + T+ S
Sbjct: 458 LAADNVI-CAVYFSTLFLLASKVPPESSASVNDDTMTTMSGS 580
>TC89342 similar to GP|10178056|dbj|BAB11420. contains similarity to NAM (no
apical meristem)~gene_id:MUB3.5 {Arabidopsis thaliana},
partial (73%)
Length = 831
Score = 28.1 bits (61), Expect = 8.6
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = -3
Query: 163 KPHDRIMALDLPHGGHLSHGYQTDTKKISAVSIFFETM 200
K H +AL H GHL H ++ D + V IFF M
Sbjct: 580 KLHPNKIALSHLHLGHLHHAHKLDISPMILVVIFFWMM 467
>TC88532 similar to PIR|S45033|S45033 probable imbibition protein - wild
cabbage, partial (33%)
Length = 1273
Score = 28.1 bits (61), Expect = 8.6
Identities = 22/50 (44%), Positives = 24/50 (48%)
Frame = +1
Query: 459 NLALKIKAESKGTKLKDFVETLQSSSYVQSEISKLRHDVEEFAKQFPTIG 508
N L K SKG KL+ +ETLQ Y S I HDV QF IG
Sbjct: 748 NAGLLSKLPSKG-KLEVSLETLQCEVYTVSPIRVFGHDV-----QFAPIG 879
>TC76702 similar to SP|Q9LZG0|ADK2_ARATH Adenosine kinase 2 (EC 2.7.1.20)
(AK 2) (Adenosine 5'- phosphotransferase 2). [Mouse-ear
cress], partial (97%)
Length = 1592
Score = 28.1 bits (61), Expect = 8.6
Identities = 25/87 (28%), Positives = 38/87 (42%)
Frame = +1
Query: 419 PGDVSAMVPGGIRMGTPALTSRGFVEEDFVKVAEYFDASVNLALKIKAESKGTKLKDFVE 478
P V A+ + MG P L V+EDF+K FD +N A I AE K + D +
Sbjct: 127 P*SVMALEGVLLGMGNPLLDISAVVDEDFLK---KFDIQLNNA--ILAEDKHKSMYDEMA 291
Query: 479 TLQSSSYVQSEISKLRHDVEEFAKQFP 505
+ Y+ ++ V ++ Q P
Sbjct: 292 AKYNVEYIAGGATQNSIRVAQWMLQVP 372
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,098,613
Number of Sequences: 36976
Number of extensions: 158685
Number of successful extensions: 702
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 687
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 689
length of query: 518
length of database: 9,014,727
effective HSP length: 100
effective length of query: 418
effective length of database: 5,317,127
effective search space: 2222559086
effective search space used: 2222559086
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126010.6


