

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126009.11 - phase: 0
(176 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
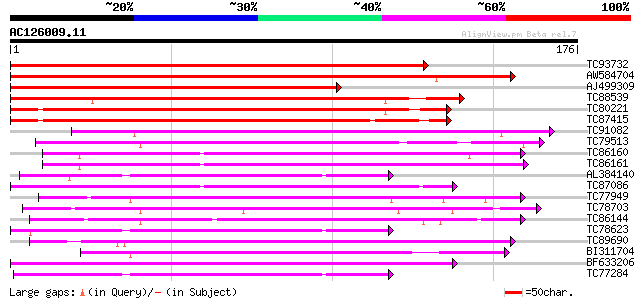
Score E
Sequences producing significant alignments: (bits) Value
TC93732 homologue to OMNI|NT01MC2635 Integral membrane protein o... 269 5e-73
AW584704 similar to SP|P05790|FBOH Fibroin heavy chain precursor... 248 9e-67
AJ499309 weakly similar to GP|7670832|gb|A dicyanin {Lycopersico... 179 7e-46
TC88539 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-... 122 6e-29
TC80221 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-... 122 6e-29
TC87415 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-... 113 4e-26
TC91082 similar to PIR|T01852|T01852 probable blue copper-bindin... 103 3e-23
TC79513 weakly similar to GP|9885806|gb|AAG01535.1| stellacyanin... 93 6e-20
TC86160 weakly similar to PIR|T01605|T01605 phytocyanin At2g4479... 87 4e-18
TC86161 weakly similar to PIR|T01605|T01605 phytocyanin At2g4479... 86 1e-17
AL384140 weakly similar to GP|2094888|pdb|2 Cucumber Basic Prote... 85 1e-17
TC87086 similar to SP|Q41001|BCP_PEA Blue copper protein precurs... 85 1e-17
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 84 4e-17
TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroele... 84 4e-17
TC86144 weakly similar to GP|10727325|gb|AAF50995.2 CG15635 gene... 80 3e-16
TC78623 similar to GP|3860333|emb|CAA10134.1 basic blue copper p... 79 7e-16
TC89690 weakly similar to PIR|G84643|G84643 similar to early nod... 78 2e-15
BI311704 weakly similar to PIR|T04605|T046 hypothetical protein ... 77 4e-15
BF633206 similar to GP|21553614|gb| blue copper protein putativ... 75 1e-14
TC77284 weakly similar to GP|6688810|emb|CAB65280.1 basic blue p... 73 5e-14
>TC93732 homologue to OMNI|NT01MC2635 Integral membrane protein of unknown
function {Magnetococcus sp. MC-1}, partial (2%)
Length = 443
Score = 269 bits (687), Expect = 5e-73
Identities = 130/130 (100%), Positives = 130/130 (100%)
Frame = +3
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV
Sbjct: 54 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 233
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT
Sbjct: 234 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 413
Query: 121 VQPKQDGWSP 130
VQPKQDGWSP
Sbjct: 414 VQPKQDGWSP 443
>AW584704 similar to SP|P05790|FBOH Fibroin heavy chain precursor (Fib-H)
(H-fibroin). [Silk moth] {Bombyx mori}, partial (0%)
Length = 679
Score = 248 bits (633), Expect = 9e-67
Identities = 124/160 (77%), Positives = 129/160 (80%), Gaps = 3/160 (1%)
Frame = +1
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MALSR+LFLFALIA+IFSTMAVAKDFVVGDE GWT DYQ W ANKVFRLGDTLTF YV
Sbjct: 52 MALSRSLFLFALIATIFSTMAVAKDFVVGDESGWTLGVDYQAWAANKVFRLGDTLTFKYV 231
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
KDNVVRVNGSDF+SCSVP APVLTSG DKI +TTYGRRWYIS V +HCENGQKLFI
Sbjct: 232 AWKDNVVRVNGSDFQSCSVPWAAPVLTSGHDKIALTTYGRRWYISGVANHCENGQKLFIN 411
Query: 121 VQPKQDGWSPV---PSPSPSPSLDLVTPEAPPSNAPWPAS 157
V PKQDGW P PS SPSPS APPSNAPW AS
Sbjct: 412 VLPKQDGWYPAPSSPSASPSPSPVPAPEAAPPSNAPWAAS 531
Score = 27.7 bits (60), Expect = 2.5
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 120 TVQPKQDGWSPVPSPSP 136
+VQ + WSPVPSP+P
Sbjct: 627 SVQASEINWSPVPSPTP 677
>AJ499309 weakly similar to GP|7670832|gb|A dicyanin {Lycopersicon
esculentum}, partial (12%)
Length = 342
Score = 179 bits (453), Expect = 7e-46
Identities = 85/103 (82%), Positives = 92/103 (88%)
Frame = -3
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MALSR LFLFALIA+IFSTMAVAKDFVVGDE+GW DYQ W ANKVFR+GDTLTFNYV
Sbjct: 310 MALSRVLFLFALIATIFSTMAVAKDFVVGDERGWKLGVDYQYWAANKVFRVGDTLTFNYV 131
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWY 103
GGKDNVVRVNGSDF+SCS+P APVLTSG D I++TTYGRRWY
Sbjct: 130 GGKDNVVRVNGSDFQSCSIPWRAPVLTSGHDTILLTTYGRRWY 2
>TC88539 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-binding
protein-like {Arabidopsis thaliana}, partial (37%)
Length = 835
Score = 122 bits (307), Expect = 6e-29
Identities = 64/143 (44%), Positives = 89/143 (61%), Gaps = 2/143 (1%)
Frame = +3
Query: 1 MALSRALFLFALIASIFSTMAVAK-DFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNY 59
MA SR + + ++ + S++A+A D +VGD+KGWT FDY W +KVFR+GD L FNY
Sbjct: 105 MASSRVVLILSISMVLLSSVAIAATDHIVGDDKGWTVDFDYTQWAQDKVFRVGDNLVFNY 284
Query: 60 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQ-KLF 118
+ NV +VNG+ F+SC+ P L++G+D I + T GR+WY+ V DHC Q KL
Sbjct: 285 DPARHNVFKVNGTLFQSCTFPPKNEALSTGKDIIQLKTEGRKWYVCGVADHCSARQMKLV 464
Query: 119 ITVQPKQDGWSPVPSPSPSPSLD 141
ITV + P+PSP PS D
Sbjct: 465 ITVLAE-----GAPAPSPPPSSD 518
>TC80221 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-binding
protein-like {Arabidopsis thaliana}, partial (40%)
Length = 813
Score = 122 bits (307), Expect = 6e-29
Identities = 64/138 (46%), Positives = 87/138 (62%), Gaps = 1/138 (0%)
Frame = +1
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
MAL+R L A+ + S++A+A D +VGD+KGWT F+Y WT +KVFR+GD L FNY
Sbjct: 67 MALNRVAIL-AISMVLLSSVAMAADHIVGDDKGWTVDFNYTQWTQDKVFRVGDNLVFNYD 243
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQ-KLFI 119
K N+ +VNG+ FK C+ P L++G+D I + T GR+WY+ V DHC Q K I
Sbjct: 244 NTKHNIFKVNGTLFKDCTFPPKNEALSTGKDIIQLKTEGRKWYVCGVADHCSAHQMKFVI 423
Query: 120 TVQPKQDGWSPVPSPSPS 137
TV + +P PSP PS
Sbjct: 424 TVLAE---GAPAPSPPPS 468
>TC87415 weakly similar to GP|9294157|dbj|BAB02059.1 blue copper-binding
protein-like {Arabidopsis thaliana}, partial (31%)
Length = 710
Score = 113 bits (282), Expect = 4e-26
Identities = 58/137 (42%), Positives = 86/137 (62%)
Frame = +1
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
M SRA++L A+ + S++A+A D VVGDEKGWT F+Y W +KVFR+GD L FNY
Sbjct: 67 MTYSRAIYL-AISMVLLSSVAMAADHVVGDEKGWTVDFNYTQWAQDKVFRVGDNLVFNYD 243
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
K NV +V+G F+SC+ P L++G+D I + T GR+WY+ +HC +++ +
Sbjct: 244 NTKHNVFKVDGKLFQSCTFPSENEALSTGKDVIQLKTEGRKWYVCGKANHCA-ARQMKLV 420
Query: 121 VQPKQDGWSPVPSPSPS 137
+ ++G PSPS S
Sbjct: 421 INVLEEG---APSPSSS 462
>TC91082 similar to PIR|T01852|T01852 probable blue copper-binding protein
F9D12.16 - Arabidopsis thaliana, partial (55%)
Length = 684
Score = 103 bits (258), Expect = 3e-23
Identities = 58/154 (37%), Positives = 81/154 (51%), Gaps = 4/154 (2%)
Frame = +2
Query: 20 MAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSC 77
++ A + VGD GWTTL DY+ W A K F+LGDT+ F Y NV+RV + +KSC
Sbjct: 11 ISYAAVYKVGDSAGWTTLGNIDYKKWAATKNFQLGDTIIFEYSAKFHNVMRVTHAMYKSC 190
Query: 78 SVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPS 137
+ T+G D I IT +G ++ V HC+ GQK+ I V SP PS SPS
Sbjct: 191 NASSPIATFTTGNDTIKITNHGHHFFFCGVPGHCQAGQKVDINVLKVSVAASPAPSSSPS 370
Query: 138 PSLDLVTPEAPPSN--APWPASSVPRRSLLPKKL 169
P SN AP P+++ P++ + K +
Sbjct: 371 ALASPAEATVPASNVPAPSPSNAAPQKFIALKMM 472
>TC79513 weakly similar to GP|9885806|gb|AAG01535.1| stellacyanin-like
protein CASLP1 precursor {Capsicum annuum}, partial
(33%)
Length = 1093
Score = 92.8 bits (229), Expect = 6e-20
Identities = 58/163 (35%), Positives = 84/163 (50%), Gaps = 5/163 (3%)
Frame = +3
Query: 9 LFALIASIFSTMAVAKDFVVGDEKGWTTLFD----YQTWTANKVFRLGDTLTFNYVGGKD 64
L A+ A++F VVGD GWT + Y W +NK F +GDTL FNY G+
Sbjct: 426 LLAVAANLFHGSFAQTRHVVGDTTGWTIPTNGASFYTNWASNKTFTVGDTLVFNYASGQH 605
Query: 65 NVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPK 124
+V +V + + SC+ T LT+ + + G++ +I +V HC GQKL I V
Sbjct: 606 DVAKVTKTAYDSCNGANTLFTLTNSPATVTLNETGQQNFICAVPGHCSAGQKLSINV--V 779
Query: 125 QDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASS-VPRRSLLP 166
+ SPV +P+PS S P+A P+ P PA S P ++ P
Sbjct: 780 KASASPVSAPTPSAS----PPKATPAPTPVPAKSPAPTKAATP 896
Score = 28.9 bits (63), Expect = 1.1
Identities = 14/38 (36%), Positives = 20/38 (51%), Gaps = 4/38 (10%)
Frame = +3
Query: 26 FVVGDEKGWTTLFD----YQTWTANKVFRLGDTLTFNY 59
+ VGD GW + Y T + K F++GD L FN+
Sbjct: 963 YTVGDTIGWIIPSNGTAAYTTXASGKSFKVGDILVFNF 1076
>TC86160 weakly similar to PIR|T01605|T01605 phytocyanin At2g44790
[imported] - Arabidopsis thaliana, partial (34%)
Length = 918
Score = 86.7 bits (213), Expect = 4e-18
Identities = 51/156 (32%), Positives = 71/156 (44%), Gaps = 6/156 (3%)
Frame = +3
Query: 11 ALIASIFSTM-----AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDN 65
++IAS F + A A DF VGD GW DY W + K F++GD L F Y G
Sbjct: 129 SMIASFFVLLLAFPYAFATDFTVGDANGWNLGVDYTKWASGKTFKVGDNLVFKY-GSSHQ 305
Query: 66 VVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQ 125
V V+ SD+KSC+ G K+ +T G+ ++I HC + + + V
Sbjct: 306 VDEVDESDYKSCTSSNAIKNYAGGNSKVPLTKAGKIYFICPTLGHCTSTGGMKLEVNVVA 485
Query: 126 DGWSPVPSPSPSPSLD-LVTPEAPPSNAPWPASSVP 160
+P PS +P P+ TP PS P S P
Sbjct: 486 ASTTPTPSGTPPPTKSPSTTPSTTPSTTPSTTPSAP 593
>TC86161 weakly similar to PIR|T01605|T01605 phytocyanin At2g44790
[imported] - Arabidopsis thaliana, partial (17%)
Length = 736
Score = 85.5 bits (210), Expect = 1e-17
Identities = 51/156 (32%), Positives = 72/156 (45%), Gaps = 5/156 (3%)
Frame = +1
Query: 11 ALIASIFSTM-----AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDN 65
++IAS F + A A DF VGD GWT DY W + K F++GD L F Y G
Sbjct: 109 SMIASFFVLLLAFPYAFATDFTVGDANGWTQGVDYTKWASGKTFKVGDNLVFKY-GSFHQ 285
Query: 66 VVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQ 125
V V+ S +KSCS T G K+ +T G+ ++I HC + + + V
Sbjct: 286 VNEVDESGYKSCSTSNTIKSYDDGDSKVPLTKAGKIYFICPTPGHCTSTGGMKLEVNVVA 465
Query: 126 DGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPR 161
+P PS +P P+ T + PS S P+
Sbjct: 466 ASTTPTPSGTPPPTKSPSTTPSAPSETNSTTPSPPK 573
>AL384140 weakly similar to GP|2094888|pdb|2 Cucumber Basic Protein A Blue
Copper Protein, partial (92%)
Length = 441
Score = 85.1 bits (209), Expect = 1e-17
Identities = 46/118 (38%), Positives = 66/118 (54%), Gaps = 2/118 (1%)
Frame = +2
Query: 4 SRALFLFALIASIF--STMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 61
S ++ L +L+ + S M+ A++ +VGD KGW+ F Q W A K F+ GDTL FNY
Sbjct: 59 STSIILMSLLCMLVFHSNMSFAEEHIVGDGKGWS--FGVQNWPAGKTFKAGDTLVFNYSP 232
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 119
NVV VN S + SC P + V TSG D+I + G +++ HC GQK+ +
Sbjct: 233 TSHNVVVVNKSGYDSCVAPKGSKVYTSGADRITLAK-GGNYFLCGFPGHCNLGQKIAV 403
>TC87086 similar to SP|Q41001|BCP_PEA Blue copper protein precursor. [Garden
pea] {Pisum sativum}, partial (87%)
Length = 1219
Score = 85.1 bits (209), Expect = 1e-17
Identities = 49/139 (35%), Positives = 68/139 (48%)
Frame = +1
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
M S L L + + +A VGD+ GW DY TW ++K F +GD+L FNY
Sbjct: 433 MTFSNPLILGFFLVINVAVPTLATVHTVGDKSGWAIGSDYNTWASDKTFAVGDSLVFNY- 609
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
G V V SD+KSC+ + +SG I + G ++I +V HC G KL +
Sbjct: 610 GAGHTVDEVKESDYKSCTTGNSISTDSSGPTTIPLKKAGTHYFICAVPGHCTGGMKLSVK 789
Query: 121 VQPKQDGWSPVPSPSPSPS 139
V+ S PS +PSPS
Sbjct: 790 VKASSSA-SSAPSATPSPS 843
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 83.6 bits (205), Expect = 4e-17
Identities = 52/161 (32%), Positives = 76/161 (46%), Gaps = 10/161 (6%)
Frame = +3
Query: 10 FALIASIFSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNYVGGKDNVV 67
F L + ST K F VG + GWT +Y W F++ DT+ F Y G D+V+
Sbjct: 144 FVLFTLVVSTSQAFK-FFVGGKDGWTLNPSENYNQWAGRNRFQISDTIVFKYKKGSDSVL 320
Query: 68 RVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKL-FITVQPKQD 126
V D++ C+ G+ + + G ++IS +CENGQKL + + P+
Sbjct: 321 EVKKEDYEKCNKTNPIKKFEDGETEFTLDRAGPFYFISGKDQNCENGQKLTLVVISPRTP 500
Query: 127 GWSPVPS------PSPSPSLDLVTPE-APPSNAPWPASSVP 160
SP PS PSPSP+ +P +PPS P +S P
Sbjct: 501 KSSPSPSAGGLSPPSPSPTTTTPSPSGSPPSPVAIPPASSP 623
>TC78703 homologue to GP|13129456|gb|AAK13114.1 Putative retroelement pol
polyprotein {Oryza sativa} [Oryza sativa (japonica
cultivar-group)], partial (2%)
Length = 1235
Score = 83.6 bits (205), Expect = 4e-17
Identities = 58/182 (31%), Positives = 87/182 (46%), Gaps = 21/182 (11%)
Frame = +3
Query: 5 RALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFD-----YQTWTANKVFRLGDTLTFNY 59
+AL LF L+ + A +FVVG +KGW+ D Y W F++GD+L FNY
Sbjct: 90 QALVLFCLLVLLMHK-GDAYEFVVGGQKGWSAPSDPNANPYNQWAEKSRFQVGDSLVFNY 266
Query: 60 VGGKDNVVRVNG-SDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 118
G+D+V++V D+++C+ ++ + G I + G ++IS ++C +KL
Sbjct: 267 QSGQDSVIQVTSQQDYENCNTDASSEKSSDGHTVIKLIKSGPHYFISGNKNNCLQNEKLL 446
Query: 119 I-------------TVQPKQDGWSPVPSPSP--SPSLDLVTPEAPPSNAPWPASSVPRRS 163
+ T P S PSPSP S S D +TP PPS P SS P
Sbjct: 447 VIVLADRTNKNSNQTTSPPSPSPSVAPSPSPLSSHSSDALTPIPPPS--PLNGSSTPPSP 620
Query: 164 LL 165
+L
Sbjct: 621 VL 626
>TC86144 weakly similar to GP|10727325|gb|AAF50995.2 CG15635 gene product
{Drosophila melanogaster}, partial (4%)
Length = 922
Score = 80.5 bits (197), Expect = 3e-16
Identities = 51/162 (31%), Positives = 78/162 (47%), Gaps = 8/162 (4%)
Frame = +3
Query: 7 LFLFALIASIFSTMAVAKDFVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGK 63
LF+ A A+I + A D VG GW+ Y W A+ F+ D L FN+ GG
Sbjct: 117 LFVVAFAAAILESTEAA-DHTVGGTTGWSVPSGASFYSDWAASNTFKQNDVLVFNFAGGH 293
Query: 64 DNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP 123
V V+ +DF +C++ V+T+G ++ + G ++I ++ HC +GQKL + V
Sbjct: 294 -TVAEVSKADFDNCNINQNGLVITTGPARVTLNRTGDFYFICTIQGHCSSGQKLSVKVSA 470
Query: 124 KQDG-WSPVP----SPSPSPSLDLVTPEAPPSNAPWPASSVP 160
SP P P+ +P TP PP+N P+ S P
Sbjct: 471 STPSPPSPTPPTSTPPTSTPPTSGTTP-TPPTNGGTPSPSSP 593
>TC78623 similar to GP|3860333|emb|CAA10134.1 basic blue copper protein
{Cicer arietinum}, partial (98%)
Length = 743
Score = 79.3 bits (194), Expect = 7e-16
Identities = 49/121 (40%), Positives = 63/121 (51%), Gaps = 2/121 (1%)
Frame = +3
Query: 1 MALSR--ALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 58
MAL R AL L + S +A A + VG GWT F+ W K FR GDTL FN
Sbjct: 33 MALGRGSALVLLVCFFVLNSELAHAATYTVGGPGGWT--FNTVGWPNGKRFRAGDTLVFN 206
Query: 59 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 118
Y NVV VN + SC P A V SG+D+I + G+ ++I + HCE+G K+
Sbjct: 207 YSPSAHNVVAVNKGGYDSCKTPRGAKVYRSGKDQIRLAR-GQNYFICNFVGHCESGMKIA 383
Query: 119 I 119
I
Sbjct: 384 I 386
>TC89690 weakly similar to PIR|G84643|G84643 similar to early nodulins
[imported] - Arabidopsis thaliana, partial (56%)
Length = 732
Score = 77.8 bits (190), Expect = 2e-15
Identities = 51/156 (32%), Positives = 74/156 (46%), Gaps = 5/156 (3%)
Frame = +3
Query: 7 LFLFALIASIFSTMAVAKDFVVGDEK-GW----TTLFDYQTWTANKVFRLGDTLTFNYVG 61
L LF L F+ AKD ++G + W + W ++ F++GD L Y
Sbjct: 30 LVLFVLFGCAFA----AKDILLGGKTDAWKVPSSESDSLNKWASSVRFQVGDHLILKYEA 197
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV 121
GKD+V++V+ D+ SC++ G K+ G +YIS HCE GQKL + V
Sbjct: 198 GKDSVLQVSKEDYDSCNISKPIKHYNDGNTKVRFDHSGPYYYISGEKGHCEKGQKLTVVV 377
Query: 122 QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPAS 157
+ G P+ + SPSPS V A + AP P S
Sbjct: 378 MSLKGGSRPIVAFSPSPSPAEVEGPAASAVAPAPTS 485
>BI311704 weakly similar to PIR|T04605|T046 hypothetical protein F20O9.30 -
Arabidopsis thaliana, partial (11%)
Length = 504
Score = 77.0 bits (188), Expect = 4e-15
Identities = 43/135 (31%), Positives = 68/135 (49%), Gaps = 2/135 (1%)
Frame = +2
Query: 23 AKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVP 80
AK+F VG + GW DY W FR+ DTL F YV G D+V+ V D+ SC+
Sbjct: 20 AKEFHVGGKDGWVVNPSEDYNQWARTHRFRVNDTLHFKYVKGNDSVLVVKKEDYDSCNTN 199
Query: 81 LTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSL 140
L +G K ++ G ++IS D+C++ +K+ + V + P+ +P++
Sbjct: 200 NPKQKLDNGNSKFKLSDSGFYYFISGNADNCKHDEKMIVQVMAVR--------PNVTPNV 355
Query: 141 DLVTPEAPPSNAPWP 155
V P PP++A P
Sbjct: 356 TAVPPSQPPASASPP 400
>BF633206 similar to GP|21553614|gb| blue copper protein putative
{Arabidopsis thaliana}, partial (56%)
Length = 478
Score = 75.5 bits (184), Expect = 1e-14
Identities = 39/139 (28%), Positives = 66/139 (47%)
Frame = +2
Query: 1 MALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 60
M + + L L+A++ + +A VG +GW D+ +W++ + F++GD L F Y
Sbjct: 38 MGCKKMILLALLLATLITKEVLATQHNVGGSQGWDPSSDFDSWSSGQTFKVGDQLVFKYT 217
Query: 61 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 120
V + S +K C + L++G+D + + G R++ HC+ G K+ IT
Sbjct: 218 SMHSVVELSDESAYKKCDISTPLNSLSTGKDVVKLDKPGTRYFTCGTLGHCDQGMKVKIT 397
Query: 121 VQPKQDGWSPVPSPSPSPS 139
V S SPS S S
Sbjct: 398 VGNGNGSSSTASSPSSSSS 454
>TC77284 weakly similar to GP|6688810|emb|CAB65280.1 basic blue protein
{Medicago sativa subsp. x varia}, complete
Length = 814
Score = 73.2 bits (178), Expect = 5e-14
Identities = 39/118 (33%), Positives = 63/118 (53%)
Frame = +2
Query: 2 ALSRALFLFALIASIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 61
+++ + L + + A A + VG GWT ++ TW K F+ GD L+FNY
Sbjct: 116 SMNMVTLISLLCLLVLAESANAASYTVGGTGGWT--YNTDTWPNGKKFKAGDVLSFNYDS 289
Query: 62 GKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 119
NVV V+ S + +C P A V +SG D+I ++ G+ ++I S HC++G K+ I
Sbjct: 290 TTHNVVAVDKSGYNNCKTPGGAKVFSSGSDQIRLSR-GQNYFICSYPGHCQSGMKVSI 460
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,917,887
Number of Sequences: 36976
Number of extensions: 95670
Number of successful extensions: 2161
Number of sequences better than 10.0: 233
Number of HSP's better than 10.0 without gapping: 1296
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1899
length of query: 176
length of database: 9,014,727
effective HSP length: 90
effective length of query: 86
effective length of database: 5,686,887
effective search space: 489072282
effective search space used: 489072282
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC126009.11


