

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.6 + phase: 0
(530 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
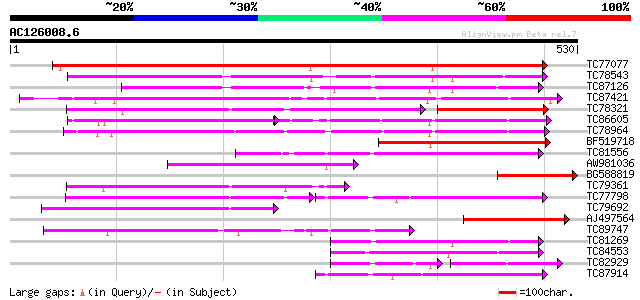
Score E
Sequences producing significant alignments: (bits) Value
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 464 e-131
TC78543 similar to PIR|T14545|T14545 probable sugar transporter ... 220 1e-57
TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {... 214 7e-56
TC87421 sugar transporter 192 3e-49
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 184 1e-46
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 110 4e-46
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 181 5e-46
BF519718 weakly similar to PIR|G84564|G84 probable sugar transpo... 161 7e-40
TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar tran... 159 3e-39
AW981036 weakly similar to PIR|G84564|G84 probable sugar transpo... 145 4e-35
BG588819 similar to PIR|C84593|C845 probable sugar transporter [... 145 5e-35
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 145 5e-35
TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 136 2e-32
TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 134 9e-32
AJ497564 weakly similar to PIR|C84593|C845 probable sugar transp... 124 1e-28
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 119 4e-27
TC81269 similar to GP|23504385|emb|CAC00697. putative sugar tran... 119 4e-27
TC84553 similar to GP|17381265|gb|AAL36051.1 AT5g18840/F17K4_90 ... 116 2e-26
TC82929 similar to PIR|D96589|D96589 hypothetical protein T22H22... 77 9e-26
TC87914 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 110 1e-24
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 464 bits (1193), Expect = e-131
Identities = 237/480 (49%), Positives = 331/480 (68%), Gaps = 18/480 (3%)
Frame = +1
Query: 41 QQQLED----KRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEF 96
Q+ L+D K+ KY ACA+ AS+ ++LLGYD+GVMSGA I+IK DLK+++ ++E
Sbjct: 55 QKNLQDFDPQKKPKRNKYAFACAMLASMTSILLGYDIGVMSGAAIYIKRDLKVSDGKIEV 234
Query: 97 LIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAG 156
L+GI++I SL+GS GRTSD IGR++T+ A +F +G + M +P+Y LM GR +AG
Sbjct: 235 LLGIINIYSLIGSCLAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYNFLMFGRFVAG 414
Query: 157 IGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVM 216
+GIG+ +MI+P+Y AE+SP +RG LT+FPE+FIN GI+LGY+SNYAFS LS+ + WR+M
Sbjct: 415 VGIGYALMIAPVYTAEVSPASSRGFLTSFPEVFINSGILLGYISNYAFSKLSLKLGWRMM 594
Query: 217 LAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAG 276
L VG +PSV + + +PESPRWLVM+ R+ +A VL KT++ ++E + RLAEI+QAAG
Sbjct: 595 LGVGAIPSVILAVGVLAMPESPRWLVMRGRLGDAIKVLNKTSDSKEEAQLRLAEIKQAAG 774
Query: 277 FAN---------SGKYEDKPVWREL-LSPPPALRRMLITGLGIQCFQQISGIDATVYYSP 326
K + VW+EL L P PA+R ++I LGI FQQ SG+DA V YSP
Sbjct: 775 IPEDCNDDVVEVKVKNTGEGVWKELFLYPTPAVRHIVIAALGIHFFQQASGVDAVVLYSP 954
Query: 327 EILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMG 386
I AGI + LL AT+AVG KT+FILVA ++D+ GR+PLL+TS GM L +
Sbjct: 955 TIFKKAGINGDTHLLIATIAVGFVKTLFILVATFMLDRYGRRPLLLTSVGGMVVSLLTLA 1134
Query: 387 VTLSLFE----KGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGA 442
V+L++ + K I L I V VA FS+G GP+ WV +SEIFPLR+RAQ +A G
Sbjct: 1135VSLTIIDHSNTKLNWAIGLSIATVLSYVATFSIGAGPITWVYSSEIFPLRLRAQGAACGV 1314
Query: 443 VANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
V NRV SG+++M+FLS+S I+ GG FFLF I+ + +F + ++PET+GK+LE++E F
Sbjct: 1315VVNRVTSGVISMTFLSLSKGITIGGAFFLFGGIATIGWIFFYIMLPETQGKTLEEMEASF 1494
>TC78543 similar to PIR|T14545|T14545 probable sugar transporter protein -
beet, partial (93%)
Length = 1653
Score = 220 bits (560), Expect = 1e-57
Identities = 141/462 (30%), Positives = 232/462 (49%), Gaps = 14/462 (3%)
Frame = +1
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGR 114
VIAC + +L + G+ G S I DL ++ + + ++ +++G++ G+
Sbjct: 148 VIACVLIVALGPIQFGFTAGYTSPTQSAIITDLGLSVSEFSLFGSLSNVGAMVGAIASGQ 327
Query: 115 TSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEIS 174
++ +GRK ++ +A++ +G + ++ A L +GRLL G G+G P+YIAEIS
Sbjct: 328 IAEYMGRKGSLMIASIPNIIGWLMISFANDSSFLYMGRLLEGFGVGIISYTVPVYIAEIS 507
Query: 175 PNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFII 234
P RGSL + ++ + +GIML Y+ L + + WR + +GI+P + LF I
Sbjct: 508 PQNLRGSLVSVNQLSVTLGIMLAYL-------LGLFVEWRFLAILGIIPCTLLIPGLFFI 666
Query: 235 PESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANS------GKYEDKPV 288
PESPRWL EE + L E ++ + EI+ A AN + + +
Sbjct: 667 PESPRWLAKMGMTEEFENSLQVLRGFETDISVEVNEIKTAVASANRRTTVRFSELKQRRY 846
Query: 289 WRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVG 348
W L+ G+G+ QQ+SGI+ ++YS I AGI AT VG
Sbjct: 847 WLPLM-----------IGIGLLVLQQLSGINGVLFYSSTIFQNAGISSSD---VATFGVG 984
Query: 349 ITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE-----KGPLVIALGI 403
+ + + + L DK GR+ LLI S+ MT L + ++ L + L L +
Sbjct: 985 AVQVLATTLTLWLADKSGRRLLLIVSSSAMTLSLLVVSISFYLKDYYISADSSLYGILSL 1164
Query: 404 LFVCGNVAF---FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVS 460
L V G V FS+G+G + W++ SEI P+ ++ A + +AN S L+ ++ +
Sbjct: 1165LSVAGVVVMVIAFSLGMGAMPWIIMSEILPINIKGLAGSFATLANWFFSWLITLTANLLL 1344
Query: 461 DAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
D S GGTF +++ + A + FV VPETKGK+LE+I+ F
Sbjct: 1345D-WSSGGTFTIYTVVCAFTVGFVAIWVPETKGKTLEEIQQFF 1467
>TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {Arabidopsis
thaliana}, partial (79%)
Length = 1524
Score = 214 bits (545), Expect = 7e-56
Identities = 136/406 (33%), Positives = 225/406 (54%), Gaps = 11/406 (2%)
Frame = +3
Query: 105 SLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVM 164
+++G++ G+ ++ +GRK ++ +A++ +G + ++ A L +GRLL G G+G
Sbjct: 9 AMVGAIASGQIAEYVGRKGSLMIASIPNIIGWLAISFAKDSSFLFMGRLLEGFGVGIISY 188
Query: 165 ISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPS 224
+ P+YIAEI+P RGSL + ++ + +GIML Y+ L + +WRV+ +GILP
Sbjct: 189 VVPVYIAEIAPENMRGSLGSVNQLSVTIGIMLAYL-------LGLFANWRVLAILGILPC 347
Query: 225 VFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYE 284
+ LF IPESPRWL +EE + L + ++ + EI++A A++GK
Sbjct: 348 TVLIPGLFFIPESPRWLAKMGMMEEFETSLQVLRGFDTDISVEVHEIKKAV--ASNGK-- 515
Query: 285 DKPVWRELLSPPPALRRM----LITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKL 340
R + R+ L G+G+ QQ+SGI+ ++YS I AGI +
Sbjct: 516 -----RATIRFADLQRKRYWFPLSVGIGLLVLQQLSGINGVLFYSTSIFANAGISSSN-- 674
Query: 341 LAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL---FEKGPL 397
AATV +G + + VA L+DK GR+ LLI S+ MTA L + + L EK
Sbjct: 675 -AATVGLGAIQVIATGVATWLVDKSGRRVLLIISSSLMTASLLVVSIAFYLEGVVEKDSQ 851
Query: 398 VIA-LGILFVCGNVAF---FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVA 453
+ LGI+ V G V FS+GLGP+ W++ SEI P+ ++ A + +AN + + ++
Sbjct: 852 YFSILGIISVVGLVVMVIGFSLGLGPIPWLIMSEILPVNIKGLAGSTATMANWLVAWIIT 1031
Query: 454 MSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
M+ ++ S GGTF +++ ++A +VF VPETKG++LE+I+
Sbjct: 1032MT-ANLLLTWSSGGTFLIYTVVAAFTVVFTSLWVPETKGRTLEEIQ 1166
>TC87421 sugar transporter
Length = 1753
Score = 192 bits (488), Expect = 3e-49
Identities = 150/547 (27%), Positives = 258/547 (46%), Gaps = 40/547 (7%)
Frame = +2
Query: 10 EMGLSGIPLGTKNKYKRMNSHLVDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLL 69
+M GIP+G NK E N T I C I A++ ++
Sbjct: 26 KMAGGGIPIGGGNK---------------------EYPGNLTPFVTITC-IVAAMGGLIF 139
Query: 70 GYDVGVMSG-------------AVIFIKEDLKITEVQVEF-------LIGILSIVSLLGS 109
GYD+G+ G AV K K T ++ L + +LL S
Sbjct: 140 GYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSS 319
Query: 110 LGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIY 169
L + GRK +M ++F +G + A +L++GR+L G GIGF P+Y
Sbjct: 320 LVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQPVPLY 499
Query: 170 IAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGF 229
++E++P RG+L ++ I +GI++ V NY F+ + WR+ L ++P++ I
Sbjct: 500 LSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITI 679
Query: 230 ALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVW 289
++P++P ++ + + A++ L + E +V+E ++ A+ + + W
Sbjct: 680 GSLVLPDTPNSMIERGDRDGAKAQLKRIRGIE-DVDEEFNDLVA----ASEASMQVENPW 844
Query: 290 RELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGI 349
R LL R L + I FQQ +GI+ ++Y+P + + G +D + L++A V G+
Sbjct: 845 RNLLQ--RKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSA-VITGV 1015
Query: 350 TKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTL-----------SLFEKGPLV 398
V V+I +DK GR+ L + M C + + +L E +V
Sbjct: 1016VNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIV 1195
Query: 399 IALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLS 458
+ +LF+C VA F+ GP+ W++ SEIFPL +R+ A ++ N + + LVA FL
Sbjct: 1196V---VLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLI 1366
Query: 459 VSDAISFGGTFFLFSAISALAI-VFVFTLVPETKGKSLEQIEMMFEN--------EHGSQ 509
+ + FG FLF A L + ++VF L+PETKG +E+++ ++++ EHG
Sbjct: 1367MLCHMKFG--LFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFVEHGDH 1540
Query: 510 GKEMELG 516
G +E+G
Sbjct: 1541GNGVEMG 1561
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 184 bits (466), Expect = 1e-46
Identities = 113/338 (33%), Positives = 175/338 (51%), Gaps = 3/338 (0%)
Frame = +2
Query: 54 YVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIV---SLLGSL 110
YV+ A A + +L GYD GV+SGA+++I+++ E + I+S +++G+
Sbjct: 218 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDEFPAVEKKTWLQEAIVSTAIAGAIIGAA 397
Query: 111 GGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYI 170
GG +D GRK ++ +A +F +G I + AP+ L++GR+ G+G+G M SP+YI
Sbjct: 398 IGGWINDRFGRKVSIIVADTLFLLGSIILAAAPNPATLIVGRVFVGLGVGMASMASPLYI 577
Query: 171 AEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFA 230
+E SP RG+L + I G L Y+ N AF+ +WR ML V P+V
Sbjct: 578 SEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT--KAPGTWRWMLGVAAAPAVIQIVL 751
Query: 231 LFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWR 290
+ +PESPRWL + + EEA+ +L K EVE+ EIQ E + +
Sbjct: 752 MLSLPESPRWLYRKGKEEEAKVILKKI----YEVEDYDNEIQALKESVEMELKETEKISI 919
Query: 291 ELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGIT 350
L ++RR L G+G+ FQQ +GI+ +YYSP I+ AG K L ++
Sbjct: 920 MQLVKTTSVRRGLYAGVGLAFFQQFTGINTVMYYSPSIVQLAGFASKRTALLLSLITSGL 1099
Query: 351 KTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVT 388
+++I IDK GRK L + S G+ L + VT
Sbjct: 1100NAFGSILSIYFIDKTGRKKLALISLTGVVLSLTLLTVT 1213
Score = 81.3 bits (199), Expect = 9e-16
Identities = 43/103 (41%), Positives = 63/103 (60%)
Frame = +2
Query: 401 LGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVS 460
+ IL + + FFS G+G V WV+ SEI+PLR R + + V + +V+ SFLS++
Sbjct: 1490 IAILALALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIASTTVWVSNLVVSQSFLSLT 1669
Query: 461 DAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFE 503
AI TF +F+ I+ +AI FV VPETKG +E++E M E
Sbjct: 1670 VAIGPAWTFMIFAIIAIVAIFFVIIFVPETKGVPMEEVESMLE 1798
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 110 bits (274), Expect(2) = 4e-46
Identities = 75/268 (27%), Positives = 133/268 (48%), Gaps = 8/268 (2%)
Frame = +3
Query: 247 IEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITG 306
+++ ++VL K + +E E+ +A+ A K+ +R LL R L+
Sbjct: 1992 LDKGKAVLRKIRGTDN-IEPEFLELVEASRVAKEVKHP----FRNLLKRNN--RPQLVIS 2150
Query: 307 LGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVG 366
+ + FQQ +GI+A ++Y+P + G ++ + L +A + G + +V+I +DK+G
Sbjct: 2151 IALMIFQQFTGINAIMFYAPVLFNTLGFKNDAALYSAVIT-GAINVISTIVSIYSVDKLG 2327
Query: 367 RKPLLITSTIGMTACLFCMGVTLSL--------FEKGPLVIALGILFVCGNVAFFSVGLG 418
R+ LL+ + + M + + L + KG AL ++ VC V+ F+ G
Sbjct: 2328 RRKLLLEAGVQMLLSQMVIAIVLGIKVKDHSEELSKG--YAALVVVMVCIFVSAFAWSWG 2501
Query: 419 PVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISAL 478
P+ W++ SEIFPL R+ ++ N + + ++A +FLS+ F G FF FS
Sbjct: 2502 PLAWLIPSEIFPLETRSAGQSVTVCVNFLFTAVIAQAFLSMLCYFKF-GIFFFFSGWILF 2678
Query: 479 AIVFVFTLVPETKGKSLEQIEMMFENEH 506
FVF LVPETK +E++ +H
Sbjct: 2679 MSTFVFFLVPETKNVPIEEMTQRVWKQH 2762
Score = 93.6 bits (231), Expect(2) = 4e-46
Identities = 56/219 (25%), Positives = 107/219 (48%), Gaps = 21/219 (9%)
Frame = +2
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVI---FIKEDL------------------KITEVQ 93
+I+C I A+ ++ GYDVGV G F+K+ K
Sbjct: 1355 IISC-IMAATGGLMFGYDVGVSGGVASMPPFLKKFFPTVLRQTTESDGSESNYCKYDNQG 1531
Query: 94 VEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRL 153
++ L + L + T+ ++GR+ TM +A F G A + +L++GR+
Sbjct: 1532 LQLFTSSLYLAGLTVTFFASYTTRVLGRRLTMLIAGFFFIAGVSLNASAQNLLMLIVGRV 1711
Query: 154 LAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISW 213
L G GIGF P++++EI+P+ RG+L ++ I +GI+ + NYA + + H W
Sbjct: 1712 LLGCGIGFANQAVPVFLSEIAPSRIRGALNILFQLDITLGILYANLVNYATNKIKGHWGW 1891
Query: 214 RVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARS 252
R+ L +G +P++ + +++ ++P L+ + + + +S
Sbjct: 1892 RISLGLGGIPALLLTLGAYLVVDTPNSLIERGHLGQRKS 2008
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 181 bits (460), Expect = 5e-46
Identities = 129/481 (26%), Positives = 228/481 (46%), Gaps = 27/481 (5%)
Frame = +1
Query: 51 TRKYVIACAIFASLNNVLLGYDVGVMSGAV------------IFIKEDLKITEVQ----- 93
T + C + A L L GYD+GV G ++ K+ + E
Sbjct: 97 TAYFAFTCVVGA-LGGSLFGYDLGVSGGVTSMDDFLEKFFPDVYRKKHAHLKETDYCKYD 273
Query: 94 ---VEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMI 150
+ L +L+ + + GRK T+ + A+ F +G I A + L+I
Sbjct: 274 NQVLTLFTSSLYFSALVMTFFASYLTRNKGRKATIIVGALSFLIGAILNAAAQNIPTLII 453
Query: 151 GRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVH 210
GR+ G GIGFG P+Y++E++P +RG++ + GI++ + NY + H
Sbjct: 454 GRVFLGGGIGFGNQAVPLYLSEMAPASSRGAVNQLFQFTTCAGILIANLVNYFTDKIHPH 633
Query: 211 ISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAE 270
WR+ L + +P+V + E+P LV Q R++EAR VL K K V+ +
Sbjct: 634 -GWRISLGLAGIPAVLMLLGGIFCAETPNSLVEQGRLDEARKVLEKV-RGTKNVDAEFED 807
Query: 271 IQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILM 330
++ A+ A + K K + + P +++I LG FQQ++G ++ ++Y+P I
Sbjct: 808 LKDASELAQAVKSPFKVLLKRKYRP-----QLIIGALGTPAFQQLTGNNSILFYAPVIFQ 972
Query: 331 AAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLS 390
++G + L ++ + G V ++++ L+DK G + + + M C+ V L+
Sbjct: 973 SSGFGSNAALFSSFITNG-ALLVATVISMFLVDKFGTRKFFLEAGFEMICCMIITAVVLA 1149
Query: 391 L-------FEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAV 443
+ KG + A ++ + V + GP+ W++ SE+FPL +R+ A ++
Sbjct: 1150VEFGHGKELSKG--ISAFLVIMIFWFVLAYGRSWGPLGWLVPSELFPLEIRSAAQSIAVC 1323
Query: 444 ANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFE 503
N + + LVA FL + + G F LF + + VFVF L+PETK +E+I ++FE
Sbjct: 1324VNMIFTALVAQLFLLSLCHLKY-GIFLLFGGLIVVMSVFVFFLLPETKQVPIEEIYLLFE 1500
Query: 504 N 504
N
Sbjct: 1501N 1503
>BF519718 weakly similar to PIR|G84564|G84 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (20%)
Length = 659
Score = 161 bits (407), Expect = 7e-40
Identities = 78/165 (47%), Positives = 116/165 (70%), Gaps = 4/165 (2%)
Frame = +3
Query: 345 VAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLF----EKGPLVIA 400
+ +GI KT F+L + +++D+ GR+P+L+ + GM LF +G+ +L EK IA
Sbjct: 3 IIMGIAKTCFVLFSALVLDRFGRRPMLLLGSSGMAVSLFGLGMGCTLLHNSDEKPMWAIA 182
Query: 401 LGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVS 460
L ++ VC V+FFS+GLGP WV +SEIFP+R+RAQ ++L NR+ SG+V+MSFLS+S
Sbjct: 183 LCVVAVCAAVSFFSIGLGPTTWVYSSEIFPMRLRAQGTSLAISVNRLISGVVSMSFLSIS 362
Query: 461 DAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENE 505
+ I+FGG FF+ + + LA +F + +PETKGKSLE+IE +FE E
Sbjct: 363 EEITFGGMFFVLAGVMVLATLFFYYFLPETKGKSLEEIEALFEEE 497
>TC81556 similar to GP|2688830|gb|AAB88879.1| putative sugar transporter
{Prunus armeniaca}, partial (59%)
Length = 1074
Score = 159 bits (402), Expect = 3e-39
Identities = 99/288 (34%), Positives = 155/288 (53%)
Frame = +1
Query: 212 SWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEI 271
+WR M + ++PS+ + + I PESPRWL Q +I EA + KT + ER+A +
Sbjct: 16 AWRTMFGIAVVPSILLALGMAISPESPRWLFQQGKIAEAEKAI-KTLYGK----ERVATV 180
Query: 272 QQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMA 331
A+ G E + W +L S +++ G + FQQ++GI+A VYYS + +
Sbjct: 181 MYDLRTASQGSSEPEAGWFDLFSS--RYWKVVSVGAALFLFQQLAGINAVVYYSTSVFRS 354
Query: 332 AGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSL 391
AGI +AA+ VG + + +A L+DK GRK LLITS GM A + + ++ +
Sbjct: 355 AGIASD---VAASALVGASNVIGTAIASSLMDKQGRKSLLITSFSGMAASMLLLSLSFTW 525
Query: 392 FEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGL 451
P L +L V FS+G GPV +L EIF R+RA+A +L + + + +
Sbjct: 526 KVLAPYSGTLAVLGTVLYVLSFSLGAGPVPALLLPEIFASRIRAKAVSLSLGTHWISNFV 705
Query: 452 VAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
+ + FLSV + + FSA+ LA++++ V ETKG+SLE+IE
Sbjct: 706 IGLYFLSVVNKFGISSVYLGFSAVCVLAVLYIAGNVVETKGRSLEEIE 849
>AW981036 weakly similar to PIR|G84564|G84 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (25%)
Length = 575
Score = 145 bits (366), Expect = 4e-35
Identities = 75/190 (39%), Positives = 113/190 (59%), Gaps = 11/190 (5%)
Frame = +2
Query: 148 LMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGL 207
LMIGR +AG G+GF ++I +Y AEIS RG LT+ P++ IN+G +LGY+SNY L
Sbjct: 2 LMIGRCIAGFGVGFALIIVSVYSAEISSPSYRGFLTSLPDLSINIGFLLGYLSNYFLGKL 181
Query: 208 SVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEER 267
S+ + WR+MLA+ +PS+ + + + ESPRWLVMQ R+ +A+ VLL + ++E E+R
Sbjct: 182 SLRLGWRIMLAIPSIPSIGLVILMLQLVESPRWLVMQGRLGDAKKVLLLISNSKQEAEQR 361
Query: 268 LAEIQQAAGFANSGKYEDKPVWRELLS-----------PPPALRRMLITGLGIQCFQQIS 316
+ EI+ A G + V ++ S P P + R+LI +G+ FQ I
Sbjct: 362 MKEIKNAVGIDENCTQNIVHVSKKTRSGGGALKEMFYKPSPHVYRILIAAIGVHIFQNIC 541
Query: 317 GIDATVYYSP 326
G++ YSP
Sbjct: 542 GVEGIFLYSP 571
>BG588819 similar to PIR|C84593|C845 probable sugar transporter [imported] -
Arabidopsis thaliana, partial (8%)
Length = 651
Score = 145 bits (365), Expect = 5e-35
Identities = 74/74 (100%), Positives = 74/74 (100%)
Frame = +2
Query: 457 LSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELG 516
LSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELG
Sbjct: 5 LSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELG 184
Query: 517 DVEQLVQNKTGLTN 530
DVEQLVQNKTGLTN
Sbjct: 185 DVEQLVQNKTGLTN 226
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 145 bits (365), Expect = 5e-35
Identities = 90/270 (33%), Positives = 146/270 (53%), Gaps = 6/270 (2%)
Frame = +1
Query: 54 YVIACAIFASLNNVLLGYDVGVMSGAVIFIKED---LKITEVQVEFLIGILSIVSLLGSL 110
Y++ A A + +L GYD GV+SGA+++IK+D ++ + E ++ + +++G+
Sbjct: 166 YILGLAAVAGIGGLLFGYDTGVISGALLYIKDDFPQVRNSNFLQETIVSMAIAGAIVGAA 345
Query: 111 GGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYI 170
GG +D GRK LA V+F +G I M AP VL+ GRLL G+G+G + +P+YI
Sbjct: 346 FGGWLNDAYGRKKATLLADVIFILGAILMAAAPDPYVLIAGRLLVGLGVGIASVTAPVYI 525
Query: 171 AEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFA 230
AE++P+ RGSL + + I G + Y+ N F+ V +WR ML V +P++
Sbjct: 526 AEVAPSEIRGSLVSTNVLMITGGQFVSYLVNLVFT--QVPGTWRWMLGVSGVPALIQFIC 699
Query: 231 LFIIPESPRWLVMQNRIEEARSVLLK---TNEDEKEVEERLAEIQQAAGFANSGKYEDKP 287
+ +PESPRWL ++NR EA V+ K + E E++ A+ +Q ++ K+
Sbjct: 700 MLFLPESPRWLFIKNRKNEAVDVISKIYDLSRLEDEIDFLTAQSEQERQRRSTIKF---- 867
Query: 288 VWRELLSPPPALRRMLITGLGIQCFQQISG 317
W S R + G G+ FQQ +G
Sbjct: 868 -WHVFRS--KETRLAFLDGGGLLAFQQFTG 948
>TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (88%)
Length = 2467
Score = 136 bits (343), Expect = 2e-32
Identities = 78/234 (33%), Positives = 132/234 (56%), Gaps = 1/234 (0%)
Frame = +3
Query: 53 KYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGG 112
K + AI AS+ N L G+D ++G++++IK+DL + ++ + I + + +
Sbjct: 114 KGAVLVAIAASIGNFLQGWDNATIAGSILYIKKDLALQTTMEGLVVAMSLIGATVITTCS 293
Query: 113 GRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAE 172
G SD +GR+ M +++V++ +G + M +P+ VL + RLL G GIG V + P+YI+E
Sbjct: 294 GPISDWLGRRPMMIISSVLYFLGSLVMLWSPNVYVLCLARLLDGFGIGLAVTLVPVYISE 473
Query: 173 ISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVF-IGFAL 231
+P+ RGSL T P+ + G+ L Y + S LS SWR+ML V +PS+F +
Sbjct: 474 TAPSDIRGSLNTLPQFSGSGGMFLSYCMVFVMS-LSPSPSWRIMLGVLSIPSLFYFLLTV 650
Query: 232 FIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYED 285
F +PESPRWLV + ++ EA+ VL + + +V +A + + G E+
Sbjct: 651 FFLPESPRWLVSKGKMLEAKKVLQRL-RGQDDVSGEMALLVEGLGIGGDASIEE 809
Score = 108 bits (270), Expect = 5e-24
Identities = 75/230 (32%), Positives = 118/230 (50%), Gaps = 14/230 (6%)
Frame = +3
Query: 287 PVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVA 346
P+W LL P ++ LI G+GIQ QQ SGI+ +YY+P+IL AG+ +L A +
Sbjct: 1593 PIWEALLEP--GVKHALIVGIGIQLLQQFSGINGVLYYTPQILEEAGVA----VLLADLG 1754
Query: 347 VGITKTVFILVAIV-------------LIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE 393
+ T + F++ A+ L+D GR+ LL+ TI + + V S+ +
Sbjct: 1755 LSSTSSSFLISAVTTLLMLPSIGLAMRLMDVTGRRQLLLV-TIPVLIVSLVILVLGSVID 1931
Query: 394 KGPLV-IALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLV 452
G +V A+ + V FF +G GP+ +L SEIFP RVR A+ A+ + +V
Sbjct: 1932 FGSVVHAAISTVCVVVYFCFFVMGYGPIPNILCSEIFPTRVRGLCIAICALVFWIGDIIV 2111
Query: 453 AMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
S + ++ G F +++ + ++ VFV+ VPETKG LE I F
Sbjct: 2112 TYSLPVMLSSLGLAGVFGVYAIVCCISWVFVYLKVPETKGMPLEVITEFF 2261
>TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (49%)
Length = 1353
Score = 134 bits (337), Expect = 9e-32
Identities = 72/225 (32%), Positives = 134/225 (59%), Gaps = 3/225 (1%)
Frame = +2
Query: 30 HLVDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKI 89
HL+ + D L + ++ + T + A+ A++ N+L G+D ++G++++IK + ++
Sbjct: 86 HLIQLHADRLIDRSIDQAKKETMSGAVIVAVAAAIGNLLQGWDNATIAGSILYIKREFQL 265
Query: 90 -TEVQVEFLIGILSIV-SLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQV 147
+E VE LI +S++ + + + G SD+ GR+ + ++++++ + + M +P+ +
Sbjct: 266 QSEPTVEGLIVAMSLIGATVVTTCSGALSDLFGRRPMLIISSLLYFLSSLVMFWSPNVYI 445
Query: 148 LMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGL 207
L+ RLL G+GIG V + P+YI+EI+P RGSL T P+ + G+ Y + S L
Sbjct: 446 LLFARLLDGLGIGLAVTLVPLYISEIAPPEIRGSLNTLPQFAGSAGMFFSYCMVFGMS-L 622
Query: 208 SVHISWRVMLAVGILPS-VFIGFALFIIPESPRWLVMQNRIEEAR 251
+ SWR+ML V +PS ++ L ++PESPRWLV + R+ EA+
Sbjct: 623 TKAPSWRLMLGVLSIPSLIYFALTLLVLPESPRWLVSKGRMLEAK 757
>AJ497564 weakly similar to PIR|C84593|C845 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (17%)
Length = 610
Score = 124 bits (310), Expect = 1e-28
Identities = 63/99 (63%), Positives = 79/99 (79%)
Frame = +2
Query: 425 TSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVF 484
+SEIFPLR+RAQASALGAV +RV SG ++MSFLSV+ AI+ GTFF+F IS A+ FV
Sbjct: 2 SSEIFPLRLRAQASALGAVGSRVSSGAISMSFLSVTKAITVAGTFFVFGVISCSAVAFVH 181
Query: 485 TLVPETKGKSLEQIEMMFENEHGSQGKEMELGDVEQLVQ 523
VPETKGKSLE+IE++F+N SQ E+E+GDVE L+Q
Sbjct: 182 YCVPETKGKSLEEIEVLFQNVGESQESEVEMGDVEHLMQ 298
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 119 bits (297), Expect = 4e-27
Identities = 99/359 (27%), Positives = 164/359 (45%), Gaps = 12/359 (3%)
Frame = +3
Query: 32 VDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKIT- 90
V++N D+L ++ N + K + + A++ + L GY +GV++ + I DL
Sbjct: 303 VEENLDLLDNFIDKETTNPSWKLSLPHVLVATITSFLFGYHLGVVNEPLESISVDLGFNG 482
Query: 91 -EVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLM 149
+ ++ I +L G L G +D +GR+ L A+ +G + ++
Sbjct: 483 NTLAEGLVVSICLGGALFGCLLSGWIADAVGRRRAFQLCALPMIIGAAMSAATNNLFGML 662
Query: 150 IGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSV 209
+GRL G G+G G ++ +Y+ E+SP RG+ +I GI+ F G+ V
Sbjct: 663 VGRLFVGTGLGLGPPVAALYVTEVSPAFVRGTYGALIQIATCFGIL-----GSLFIGIPV 827
Query: 210 -HIS--WRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEE 266
IS WRV V +P+ + A+ ESP WL Q R EA + E
Sbjct: 828 KEISGWWRVCFWVSTIPAAILALAMDFCAESPHWLYKQGRTAEAEAEF-----------E 974
Query: 267 RLAEIQQAAGFANS-------GKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGID 319
RL + +A FA S G+ D + ELL + +++ G + QQ+SGI+
Sbjct: 975 RLLGVSEAK-FAMSQLSKVDRGEDTDTVKFSELLHGHHS--KVVFIGSTLFALQQLSGIN 1145
Query: 320 ATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGM 378
A Y+S + +AG+ A V +G+ +++ L+DK+GRK LL S GM
Sbjct: 1146AVFYFSSTVFKSAGVPSD----FANVCIGVANLTGSIISTGLMDKLGRKVLLFWSFFGM 1310
>TC81269 similar to GP|23504385|emb|CAC00697. putative sugar transporter
{Lycopersicon esculentum}, partial (42%)
Length = 966
Score = 119 bits (297), Expect = 4e-27
Identities = 73/202 (36%), Positives = 112/202 (55%), Gaps = 3/202 (1%)
Frame = +2
Query: 301 RMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIV 360
R L G+G+ QQ+ GI+ +Y+ I AG + ++ I + V V
Sbjct: 35 RSLTIGVGLMVCQQLGGINGVCFYTSSIFDLAGFPSAT----GSIIYAILQIVITGVGAA 202
Query: 361 LIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALGILFVCGNVAF---FSVGL 417
LID+ GRKPLL+ S G+ A V L V A+ L V G + + FS+G+
Sbjct: 203 LIDRAGRKPLLLVSGSGLVAGCIFTAVAFYLKVHDVAVGAVPALAVTGILVYIGSFSIGM 382
Query: 418 GPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISA 477
G + WV+ SEIFP+ ++ QA ++ + N + L + +F + S+G TF L++AI+A
Sbjct: 383 GAIPWVVMSEIFPVNIKGQAGSIATLVNWFGAWLCSYTFNFLMSWSSYG-TFVLYAAINA 559
Query: 478 LAIVFVFTLVPETKGKSLEQIE 499
LAI+F+ +VPETKGKSLEQ++
Sbjct: 560 LAILFIAVVVPETKGKSLEQLQ 625
>TC84553 similar to GP|17381265|gb|AAL36051.1 AT5g18840/F17K4_90
{Arabidopsis thaliana}, partial (41%)
Length = 827
Score = 116 bits (290), Expect = 2e-26
Identities = 74/202 (36%), Positives = 114/202 (55%), Gaps = 3/202 (1%)
Frame = +2
Query: 301 RMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIV 360
R ++ G+G+ QQ GI +Y+ E +AA D T+A + F ++ +
Sbjct: 2 RSVVLGVGLMVCQQSVGIMGIGFYTSETFVAA---DFLSGKIGTIAYACMQVPFTILGAI 172
Query: 361 LIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALGILFVCG---NVAFFSVGL 417
L+DK GR+PL+ S G F G+ L ++ L+ + IL V G VA FS+G+
Sbjct: 173 LMDKSGRRPLITASASGTFLGCFMTGIAFFLKDQNLLLELVPILAVAGILIYVAAFSIGM 352
Query: 418 GPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISA 477
GPV WV+ SEIFP+ V+ A +L + N + + +V+ +F + + S GT FL++ S
Sbjct: 353 GPVPWVIMSEIFPIHVKGTAGSLVVLINWLGAWVVSYTF-NFFLSWSSPGTLFLYAGCSL 529
Query: 478 LAIVFVFTLVPETKGKSLEQIE 499
L I+FV LVPETKGK+LE+I+
Sbjct: 530 LTILFVAKLVPETKGKTLEEIQ 595
>TC82929 similar to PIR|D96589|D96589 hypothetical protein T22H22.15
[imported] - Arabidopsis thaliana, partial (30%)
Length = 779
Score = 77.4 bits (189), Expect(2) = 9e-26
Identities = 41/104 (39%), Positives = 62/104 (59%)
Frame = +1
Query: 413 FSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLF 472
FS+G+G + WV+ SEIFP+ V+ A + + +CS +V+ +F + S GTFF+F
Sbjct: 469 FSLGMGGIPWVIMSEIFPINVKGSAGSFVTFVHWLCSWIVSYAFNFLMSWNS-AGTFFIF 645
Query: 473 SAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELG 516
S I L I+FV LVPETKG++LE+++ S + E G
Sbjct: 646 STICGLTILFVAKLVPETKGRTLEEVQASLNPYQASIQQRDEFG 777
Score = 57.8 bits (138), Expect(2) = 9e-26
Identities = 33/110 (30%), Positives = 58/110 (52%), Gaps = 6/110 (5%)
Frame = +2
Query: 301 RMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIV 360
+ L G+G+ QQ G++A +Y+ I ++AG T+A+ + + + ++
Sbjct: 152 KSLTVGVGLIILQQFGGVNAIAFYASSIFVSAGFSRS----IGTIAMVVVQIPMTALGVI 319
Query: 361 LIDKVGRKPLLITSTIGMTACLFCMGVTLSLF------EKGPLVIALGIL 404
L+DK GR+PLL+ S G CL C V+LS + E P++ +G+L
Sbjct: 320 LMDKSGRRPLLLISASG--TCLGCFLVSLSFYLQDLHKEFSPILALVGVL 463
>TC87914 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (39%)
Length = 1129
Score = 110 bits (276), Expect = 1e-24
Identities = 77/230 (33%), Positives = 118/230 (50%), Gaps = 14/230 (6%)
Frame = +3
Query: 287 PVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVA 346
P W +L P ++ L G+G+Q QQ SGI+ +YY+P+IL AG+ L + +
Sbjct: 246 PSWNDLFEP--GVKHALFVGVGLQILQQFSGINGVLYYTPQILEQAGVG----YLLSNLG 407
Query: 347 VGITKTVFIL-------------VAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFE 393
+ T + F++ VA+ L+D GR+ LL+T+ + LF + V SL +
Sbjct: 408 LSSTSSSFLISAVTTLLMLPCIAVAMRLMDISGRRTLLLTTIPVLIVSLFIL-VLGSLVD 584
Query: 394 KGPLVIA-LGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLV 452
G A + + V F +G GPV +L +EIFP RVR A+ A+ +C +V
Sbjct: 585 LGDTANASISTISVVVYFCSFVMGFGPVPNILCAEIFPTRVRGLCIAICALTFWICDIIV 764
Query: 453 AMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
S + +++ GG F L++ + +A VFVF VPETKG LE I F
Sbjct: 765 TYSLPVMLNSVGLGGVFGLYAVVCCIAWVFVFLKVPETKGMPLEVIIEFF 914
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.140 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,623,767
Number of Sequences: 36976
Number of extensions: 196305
Number of successful extensions: 1108
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 1013
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1026
length of query: 530
length of database: 9,014,727
effective HSP length: 101
effective length of query: 429
effective length of database: 5,280,151
effective search space: 2265184779
effective search space used: 2265184779
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126008.6


