

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.14 + phase: 0 /pseudo/partial
(731 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
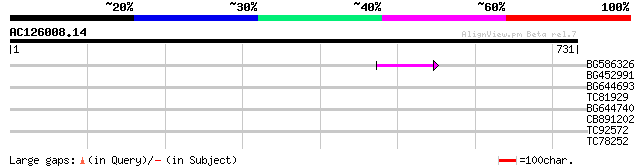
Sequences producing significant alignments: (bits) Value
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 45 8e-05
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 32 0.008
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 39 0.009
TC81929 similar to PIR|C86271|C86271 hypothetical protein AAF813... 32 1.1
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 31 1.9
CB891202 homologue to GP|23497696|gb hypothetical protein {Plasm... 30 4.3
TC92572 similar to SP|Q9ZRB0|TBB3_WHEAT Tubulin beta-3 chain (Be... 29 7.4
TC78252 similar to GP|15524450|emb|CAC69348. unnamed protein pro... 28 9.6
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 45.4 bits (106), Expect = 8e-05
Identities = 37/81 (45%), Positives = 44/81 (53%)
Frame = +3
Query: 473 RGS*KCMRRIIRLMIWNWQPLCIH*RFGGIICMVLSLRCSVTTRV*SICLIRKS*I*GRG 532
+GS*+ MR MI W *RFG CMV R T +V*SI L S* *GRG
Sbjct: 126 QGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMVPRFRYIRTIKV*SIFLPSLS*T*GRG 305
Query: 533 GG*NILRTLIFN*VIIQERLM 553
GG N T I +II+E+L+
Sbjct: 306 GGWNS*LTTI*TSLIIREKLI 368
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 31.6 bits (70), Expect(2) = 0.008
Identities = 18/38 (47%), Positives = 21/38 (54%)
Frame = +1
Query: 577 LSSFET*VWLAG*HLMV*GWVC*S*QAIFWKRSRRVRR 614
L+S ET*VW V* W C* FW S+R+RR
Sbjct: 70 LNSSET*VWFVKCRHRV*NWGC*RSTTNFWIVSKRLRR 183
Score = 26.2 bits (56), Expect(2) = 0.008
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = +3
Query: 566 CLL*WLKSWN*LSSF 580
CLL*WL+SW+ L F
Sbjct: 36 CLL*WLESWSCLEQF 80
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 38.5 bits (88), Expect = 0.009
Identities = 55/166 (33%), Positives = 75/166 (45%)
Frame = +3
Query: 126 FQRILPSYRWRGKLSLQLIWYQARVLFRLRRIRCLHQSWVN*RISWKSCLRSSLLDQVFH 185
F + LP+ GKL+L LI YQ +LF+ I +H + * + K + + VF
Sbjct: 9 FYKFLPN----GKLTLVLICYQI*ILFKFLLIG*IH*N*KC*SSNSKIY*KRVSFNLVFI 176
Query: 186 RGVLQCCWLRRRMEA*GCVWIIGS*TR*RLRIGIHFQGLMI*WIS*LVLRCLVRLI*DQD 245
VL C+ RR+M + I +* R I L I VL RL * D
Sbjct: 177 HRVLWSCF*RRKMGSLE*ALTIPN*ITST*R*SILSLLLTNYLIISKVLNGFSRLT*GWD 356
Query: 246 IIRLE*RRKIYPKLRSERGMDTMSIL*CHLEYLMRQVSLWNI*IEY 291
+ * K++ KL SE M TM *C L + + LWN *IE+
Sbjct: 357 NTNIG**VKMFLKLLSELDMVTMKS**CLLGKQIPRWHLWN**IEF 494
>TC81929 similar to PIR|C86271|C86271 hypothetical protein AAF81304.1
[imported] - Arabidopsis thaliana, partial (66%)
Length = 1651
Score = 31.6 bits (70), Expect = 1.1
Identities = 23/66 (34%), Positives = 36/66 (53%), Gaps = 7/66 (10%)
Frame = +2
Query: 458 LVVCLCRIVRL*LMLRGS*KCMRRIIRL-MIWNWQ------PLCIH*RFGGIICMVLSLR 510
L + L RI+ +LR K ++++ RL + W+W+ PL H*+FG + C+ L
Sbjct: 590 LNILLFRIMLPIFLLR---KEVKKLWRL*LAWHWECLLLALPLVTH*QFGFLFCL*LWFI 760
Query: 511 CSVTTR 516
C TTR
Sbjct: 761 CMQTTR 778
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 30.8 bits (68), Expect = 1.9
Identities = 19/55 (34%), Positives = 25/55 (44%)
Frame = -2
Query: 172 KSCLRSSLLDQVFHRGVLQCCWLRRRMEA*GCVWIIGS*TR*RLRIGIHFQGLMI 226
K L +QVF VL+CC +RM GC I + R + R GL+I
Sbjct: 288 KDSLEKGFNNQVFLHRVLRCCLCEKRMGTLGCALITDNLIRSQQRTSTLSLGLII 124
>CB891202 homologue to GP|23497696|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 646
Score = 29.6 bits (65), Expect = 4.3
Identities = 17/48 (35%), Positives = 24/48 (49%), Gaps = 5/48 (10%)
Frame = +1
Query: 446 HLLCIVMLLRWVLVVC-----LCRIVRL*LMLRGS*KCMRRIIRLMIW 488
H LCI +L W++V+C C R * L KC+ + L+IW
Sbjct: 52 HYLCIFLL--WIVVICKLRLRFCDSRRC*WPLHCGAKCLNTALPLVIW 189
>TC92572 similar to SP|Q9ZRB0|TBB3_WHEAT Tubulin beta-3 chain (Beta-3
tubulin). [Wheat] {Triticum aestivum}, partial (25%)
Length = 720
Score = 28.9 bits (63), Expect = 7.4
Identities = 27/74 (36%), Positives = 38/74 (50%), Gaps = 11/74 (14%)
Frame = +2
Query: 133 YRWRGKLSL----QLIWYQARVLFRLRRIRCLHQSWVN*RISWKSCL---RSSLL----D 181
+R R +L+L Q + Q V+FR + RCL W R +K CL RSSLL +
Sbjct: 5 FRTRTRLTLWNGFQTM*NQVFVIFRQQGCRCLRHLW-GIRHLFKKCLDVFRSSLLLCLRE 181
Query: 182 QVFHRGVLQCCWLR 195
+ F G+L W+R
Sbjct: 182 RHFCIGILLKEWMR 223
>TC78252 similar to GP|15524450|emb|CAC69348. unnamed protein product {Glycine
max}, partial (66%)
Length = 1395
Score = 28.5 bits (62), Expect = 9.6
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -3
Query: 447 LLCIVMLLRWVLVVCLC 463
LLCI++ W+L VCLC
Sbjct: 1246 LLCIIISSSWILKVCLC 1196
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.376 0.169 0.715
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,775,086
Number of Sequences: 36976
Number of extensions: 573777
Number of successful extensions: 11414
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2074
Number of HSP's successfully gapped in prelim test: 411
Number of HSP's that attempted gapping in prelim test: 8990
Number of HSP's gapped (non-prelim): 3138
length of query: 731
length of database: 9,014,727
effective HSP length: 103
effective length of query: 628
effective length of database: 5,206,199
effective search space: 3269492972
effective search space used: 3269492972
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126008.14


