

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
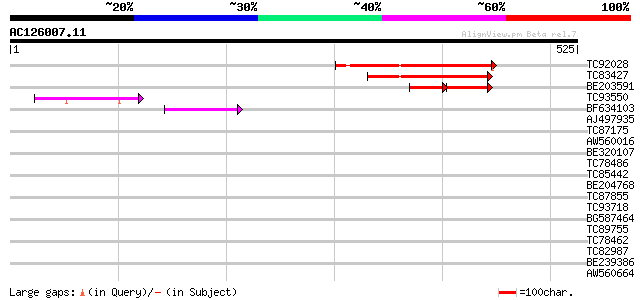
Query= AC126007.11 + phase: 0
(525 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92028 weakly similar to GP|20260434|gb|AAM13115.1 putative PHD... 132 4e-31
TC83427 weakly similar to GP|3236235|gb|AAC23623.1| unknown prot... 124 9e-29
BE203591 similar to PIR|C84679|C84 hypothetical protein At2g2798... 65 6e-21
TC93550 weakly similar to GP|20260434|gb|AAM13115.1 putative PHD... 61 1e-09
BF634103 weakly similar to GP|20260434|gb putative PHD-type zinc... 47 2e-05
AJ497935 similar to GP|3236235|gb|A unknown protein {Arabidopsis... 41 0.001
TC87175 similar to GP|21450874|gb|AAK59489.2 unknown protein {Ar... 38 0.011
AW560016 similar to GP|8778713|gb T25N20.3 {Arabidopsis thaliana... 35 0.054
BE320107 weakly similar to PIR|T47571|T475 hypothetical protein ... 33 0.21
TC78486 similar to GP|21592780|gb|AAM64729.1 nucleic acid bindin... 32 0.60
TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 32 0.60
BE204768 similar to PIR|A41139|A411 protein kinase 1 (EC 2.7.1.-... 32 0.78
TC87855 similar to GP|15081801|gb|AAK82555.1 AT5g41350/MYC6_6 {A... 32 0.78
TC93718 weakly similar to GP|10178129|dbj|BAB11541. gene_id:MOP1... 32 0.78
BG587464 31 1.0
TC89755 similar to GP|10177798|dbj|BAB11289. emb|CAB87778.1~gene... 31 1.3
TC78462 similar to GP|21592780|gb|AAM64729.1 nucleic acid bindin... 30 1.7
TC82987 similar to GP|15991740|gb|AAL13047.1 WRKY transcription ... 30 1.7
BE239386 similar to GP|1747310|dbj| Myb-like DNA binding protein... 30 1.7
AW560664 similar to GP|23495077|gb hypothetical protein {Plasmod... 30 2.3
>TC92028 weakly similar to GP|20260434|gb|AAM13115.1 putative PHD-type zinc
finger protein {Arabidopsis thaliana}, partial (14%)
Length = 794
Score = 132 bits (331), Expect = 4e-31
Identities = 74/151 (49%), Positives = 107/151 (70%), Gaps = 2/151 (1%)
Frame = +2
Query: 302 LDIKWKVFNRQLIVSKIITSSLLSDVVTIFHEQFDSIVVTGTKIDLIPAMVKGRKIKDKY 361
+DI+ ++ N + V+ T LL + ++IF+E FD IV T+ DLIP+MV GR ++ +
Sbjct: 2 IDIRRRLANGK--VASPETRPLLLEALSIFNECFDPIVDAATERDLIPSMVYGRNLQTQD 175
Query: 362 YFGGMYCAVLIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQGFFKCLLSCIENVLKE 421
FGG+YCA+L+VN VVSAG+ R+FG ++AEL LIAT+ + + +G+F+ L SCIE ++
Sbjct: 176 -FGGVYCALLMVNSSVVSAGMLRIFGTDIAELPLIATRHKNRGKGYFQTLFSCIERLVAF 352
Query: 422 LKVERLVLPAAHEAESMWIDKFGF--TEPNQ 450
L V+ LVLPAA EAES+W KFGF T+P Q
Sbjct: 353 LNVKNLVLPAAEEAESIWKYKFGFSRTKPEQ 445
>TC83427 weakly similar to GP|3236235|gb|AAC23623.1| unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 798
Score = 124 bits (311), Expect = 9e-29
Identities = 63/116 (54%), Positives = 84/116 (72%)
Frame = +1
Query: 332 HEQFDSIVVTGTKIDLIPAMVKGRKIKDKYYFGGMYCAVLIVNQVVVSAGIFRVFGKEVA 391
HE FD IV + DLI AMV G+ ++ + FGGMYCA+LIVN VVSAG+ R+FG ++A
Sbjct: 1 HECFDPIVDAVSGRDLIRAMVYGKSVRGQE-FGGMYCALLIVNSSVVSAGMLRIFGTDIA 177
Query: 392 ELSLIATKAEYQKQGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTE 447
EL L+AT +G+F+ L SCIE +L +KV+ LVLPAA EA+S+W DKFGF++
Sbjct: 178 ELPLVATSNSQHGKGYFQALFSCIERLLAFMKVKNLVLPAAEEAQSIWTDKFGFSK 345
>BE203591 similar to PIR|C84679|C84 hypothetical protein At2g27980 [imported]
- Arabidopsis thaliana, partial (7%)
Length = 617
Score = 64.7 bits (156), Expect(2) = 6e-21
Identities = 28/43 (65%), Positives = 36/43 (83%)
Frame = +3
Query: 405 QGFFKCLLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTE 447
QG+F+CL SCIE +L+ L V+ LVLPAA EAES+W +KFGFT+
Sbjct: 228 QGYFQCLFSCIERLLQSLSVKNLVLPAADEAESIWTNKFGFTK 356
Score = 54.3 bits (129), Expect(2) = 6e-21
Identities = 27/35 (77%), Positives = 30/35 (85%)
Frame = +1
Query: 371 LIVNQVVVSAGIFRVFGKEVAELSLIATKAEYQKQ 405
L VNQVVVSAG+FRVFG EVAEL L+AT +EYQ Q
Sbjct: 1 LTVNQVVVSAGVFRVFGPEVAELPLVATVSEYQGQ 105
>TC93550 weakly similar to GP|20260434|gb|AAM13115.1 putative PHD-type zinc
finger protein {Arabidopsis thaliana}, partial (10%)
Length = 911
Score = 60.8 bits (146), Expect = 1e-09
Identities = 41/148 (27%), Positives = 63/148 (41%), Gaps = 47/148 (31%)
Frame = +1
Query: 24 VEKKMKNKKKKKNKNKKESNLRSNESKN---------------------------LQPSP 56
V + K ++KK+ K+ K ++ + K+ + P
Sbjct: 250 VSSQTKKQQKKRTKSSKRLSMSKSSKKSASVAILTHKKSLCSMETKSSKVSVKLKIAPIT 429
Query: 57 KHTTQTSNSHASPTTINYRDQCLHKLVFQENVLEDGAAVGYFVY---------------- 100
++ +S + S IN + Q LHKL+F E+ L DGA V Y+
Sbjct: 430 SNSQCSSPPNKSQLRINKKHQRLHKLIFDEDGLPDGAEVAYYARGQKRLEGIKKKSGILC 609
Query: 101 ----EEVSPSKFEAHAGWASRRKPWPLI 124
E+SP++FE HAGWASRRKP+ I
Sbjct: 610 RCCNTEISPAQFEVHAGWASRRKPYAYI 693
>BF634103 weakly similar to GP|20260434|gb putative PHD-type zinc finger
protein {Arabidopsis thaliana}, partial (3%)
Length = 369
Score = 46.6 bits (109), Expect = 2e-05
Identities = 26/72 (36%), Positives = 34/72 (47%)
Frame = +3
Query: 144 VESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIAKICTLSVKHKEVEH 203
V S P+ YC C++ + +V + V +I DP EQIAK C VK E E
Sbjct: 150 VSSTPRRGRYCPICQHMFLGEGSVALNPDAVAAGRIEGVDPIEQIAKRCIRIVKDFEAET 329
Query: 204 SSCALCSERHFN 215
CALC F+
Sbjct: 330 GGCALCRGSDFS 365
>AJ497935 similar to GP|3236235|gb|A unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 106
Score = 40.8 bits (94), Expect = 0.001
Identities = 18/32 (56%), Positives = 24/32 (74%)
Frame = +1
Query: 401 EYQKQGFFKCLLSCIENVLKELKVERLVLPAA 432
E+Q +G+ + L SCIE +L L VE+LVLPAA
Sbjct: 4 EHQGKGYIQALFSCIERLLSSLNVEKLVLPAA 99
>TC87175 similar to GP|21450874|gb|AAK59489.2 unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 1612
Score = 37.7 bits (86), Expect = 0.011
Identities = 24/60 (40%), Positives = 31/60 (51%)
Frame = +3
Query: 411 LLSCIENVLKELKVERLVLPAAHEAESMWIDKFGFTEPNQGLGRRYYRRSWSFHLNKGVE 470
L S IE L LKVE+LV+PA E W FGFT + L R RS + + G++
Sbjct: 9 LFSAIELALCSLKVEKLVIPAISELVHTWTTVFGFTHLEEPL--RQEMRSLNMLVFPGID 182
>AW560016 similar to GP|8778713|gb T25N20.3 {Arabidopsis thaliana}, partial
(13%)
Length = 657
Score = 35.4 bits (80), Expect = 0.054
Identities = 18/63 (28%), Positives = 24/63 (37%)
Frame = +1
Query: 206 CALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLKDHNMANLKKVPKHYWFCGVDCYDIH 265
C L S + + + CD CEK YH C KD + FCG C ++
Sbjct: 469 CGLASATTDKEDDATVNALRTCDLCEKKYHDRCTKDMGALLANSNMSEHSFCGKSCKELF 648
Query: 266 MKL 268
L
Sbjct: 649 ENL 657
>BE320107 weakly similar to PIR|T47571|T475 hypothetical protein F24B22.80 -
Arabidopsis thaliana, partial (42%)
Length = 372
Score = 33.5 bits (75), Expect = 0.21
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = -3
Query: 33 KKKNKNKKESNLRSNESKNLQPSPKH 58
KKKN NKKE N+R+ K++ S KH
Sbjct: 274 KKKNNNKKEKNIRNKRQKSITRSLKH 197
>TC78486 similar to GP|21592780|gb|AAM64729.1 nucleic acid binding
protein-like {Arabidopsis thaliana}, partial (89%)
Length = 1089
Score = 32.0 bits (71), Expect = 0.60
Identities = 18/46 (39%), Positives = 25/46 (54%), Gaps = 1/46 (2%)
Frame = +3
Query: 196 VKHKEVEHSSCALCSERHFN-NGEFSPWTVMICDQCEKDYHVGCLK 240
V +E + CA C E + + + EF W + CD CEK YH C+K
Sbjct: 624 VDDEEEDQGECAACGESYVSASDEF--W--ICCDICEKWYHGKCVK 749
>TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (62%)
Length = 585
Score = 32.0 bits (71), Expect = 0.60
Identities = 17/51 (33%), Positives = 27/51 (52%)
Frame = +3
Query: 6 EELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESNLRSNESKNLQPSP 56
EE K+ E K+G+ E+ K ++KK + KKE + E K +P+P
Sbjct: 108 EEKKEEAKKEEEKKEGEKKEEPKKEEEKK--EEKKEEGEKKEEEKKKEPAP 254
>BE204768 similar to PIR|A41139|A411 protein kinase 1 (EC 2.7.1.-) - garden
pea (fragment), partial (19%)
Length = 565
Score = 31.6 bits (70), Expect = 0.78
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = -3
Query: 2 SCSHEELKDNIVVEAVAKDGDDVEKKMKNKKKKKNKNKKESN 43
+CS + K V+ + D +EK+ K + K KNKNK SN
Sbjct: 227 ACSIAKKKTQKVIWL*REVSDPIEKRTKGQNKNKNKNKNISN 102
>TC87855 similar to GP|15081801|gb|AAK82555.1 AT5g41350/MYC6_6 {Arabidopsis
thaliana}, partial (48%)
Length = 1194
Score = 31.6 bits (70), Expect = 0.78
Identities = 25/106 (23%), Positives = 43/106 (39%), Gaps = 1/106 (0%)
Frame = +1
Query: 135 TAVSANKEPVESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIA-KICT 193
+++S N V+ P + N K+ E KE+ Q IE D ++ ++
Sbjct: 553 SSLSTNSNSVQEPSGD--------NHGTSPKSEEPKESECKGQTDIEQDTAKDSEIELSK 708
Query: 194 LSVKHKEVEHSSCALCSERHFNNGEFSPWTVMICDQCEKDYHVGCL 239
L VE +C +C E E+ + QC D+H+ C+
Sbjct: 709 LGEPINLVEEDTCPICLE------EYDAENPKLTTQCGHDFHLACI 828
>TC93718 weakly similar to GP|10178129|dbj|BAB11541. gene_id:MOP10.6~unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 687
Score = 31.6 bits (70), Expect = 0.78
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = +2
Query: 29 KNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSN 64
K KKKKK KN ++ N R N + +PK T N
Sbjct: 110 KTKKKKKKKNTEDEN*RRNRRRTKSTNPKIKTTAIN 217
>BG587464
Length = 663
Score = 31.2 bits (69), Expect = 1.0
Identities = 20/58 (34%), Positives = 26/58 (44%), Gaps = 1/58 (1%)
Frame = +3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKH-TTQTSNSHASPTTINYRDQCLHK 81
+K + K KN K +N N L P P H TT T+N PT N + Q + K
Sbjct: 33 DKNVTPKPTDKNVTPKPTN--KNVKPKLMPKPNHTTTTTTNVMPKPTDKNVKPQLMPK 200
>TC89755 similar to GP|10177798|dbj|BAB11289.
emb|CAB87778.1~gene_id:MXA21.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (59%)
Length = 864
Score = 30.8 bits (68), Expect = 1.3
Identities = 15/23 (65%), Positives = 15/23 (65%)
Frame = -3
Query: 22 DDVEKKMKNKKKKKNKNKKESNL 44
D E K KNKKKKK K KK S L
Sbjct: 73 DFEENKNKNKKKKKKKKKKRSRL 5
>TC78462 similar to GP|21592780|gb|AAM64729.1 nucleic acid binding
protein-like {Arabidopsis thaliana}, partial (85%)
Length = 1281
Score = 30.4 bits (67), Expect = 1.7
Identities = 27/113 (23%), Positives = 46/113 (39%), Gaps = 2/113 (1%)
Frame = +3
Query: 130 PIVMNTAVSANKEPVESPPKEKWYCEYCRNKLQKDKNVEHKENVVTTQKIIESDPSEQIA 189
P + A K+ V+ ++ + +K + E V K +E +++
Sbjct: 594 PSIYEVVTGAAKKQVK---EKSSVSNHSGSKSKSSSKARAPEPQVKQTKPLELPKDDEVE 764
Query: 190 KICTLSVKHKEVEHSS--CALCSERHFNNGEFSPWTVMICDQCEKDYHVGCLK 240
++ + E EH C C E H+ EF W + CD CEK +H C+K
Sbjct: 765 ELD----EEDEDEHGETLCGACGE-HYGTDEF--W--ICCDICEKWFHGKCVK 896
>TC82987 similar to GP|15991740|gb|AAL13047.1 WRKY transcription factor 71
{Arabidopsis thaliana}, partial (33%)
Length = 975
Score = 30.4 bits (67), Expect = 1.7
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +1
Query: 19 KDGDDVEKKMKNKKKKKNKNKKE 41
+DGDD + K +NK KKK K KE
Sbjct: 568 EDGDDEKSKKENKAKKKEKKVKE 636
>BE239386 similar to GP|1747310|dbj| Myb-like DNA binding protein
{Arabidopsis thaliana}, partial (2%)
Length = 521
Score = 30.4 bits (67), Expect = 1.7
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +1
Query: 251 PKHYWFCGVDCYDIHMKLKNFMARGDVLLS 280
P WFC + CY ++KL + G L+S
Sbjct: 328 PLQSWFCNIKCYSPNLKLLQLLIYGSQLIS 417
>AW560664 similar to GP|23495077|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 545
Score = 30.0 bits (66), Expect = 2.3
Identities = 17/40 (42%), Positives = 22/40 (54%)
Frame = -3
Query: 25 EKKMKNKKKKKNKNKKESNLRSNESKNLQPSPKHTTQTSN 64
EKK K+KKKK+ K KKE R + + P+ TT N
Sbjct: 369 EKKEKSKKKKEKKKKKE---RKEKKTSKAALPRGTTLREN 259
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,337,281
Number of Sequences: 36976
Number of extensions: 285286
Number of successful extensions: 2206
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 2094
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2179
length of query: 525
length of database: 9,014,727
effective HSP length: 101
effective length of query: 424
effective length of database: 5,280,151
effective search space: 2238784024
effective search space used: 2238784024
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126007.11


