

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126006.11 + phase: 0
(325 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
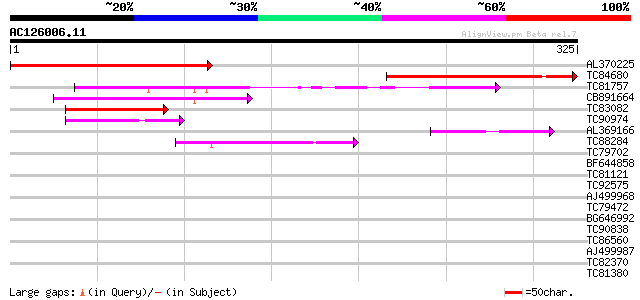
Score E
Sequences producing significant alignments: (bits) Value
AL370225 homologue to GP|21536636|gb| unknown {Arabidopsis thali... 236 7e-63
TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsi... 199 1e-51
TC81757 similar to GP|3341693|gb|AAC27475.1| unknown protein {Ar... 67 9e-12
CB891664 similar to GP|3341693|gb| unknown protein {Arabidopsis ... 64 8e-11
TC83082 similar to GP|18149189|dbj|BAB83610. 6b-interacting prot... 48 4e-06
TC90974 weakly similar to GP|9294038|dbj|BAB01995.1 gene_id:MOB2... 45 5e-05
AL369166 similar to GP|3341693|gb|A unknown protein {Arabidopsis... 43 1e-04
TC88284 similar to PIR|T03793|T03793 calmodulin-binding protein ... 41 7e-04
TC79702 MtN14 36 0.023
BF644858 weakly similar to GP|5679842|emb l1332.6 {Oryza sativa}... 33 0.11
TC81121 homologue to GP|18182311|gb|AAL65125.1 GT-2 factor {Glyc... 33 0.11
TC92575 weakly similar to GP|21428434|gb|AAM49877.1 LD13350p {Dr... 33 0.15
AJ499968 weakly similar to GP|23495961|gb| hypothetical protein ... 33 0.20
TC79472 similar to PIR|C96685|C96685 unknown protein F15E12.10 [... 32 0.26
BG646992 similar to PIR|T06133|T061 hypothetical protein F23E12.... 31 0.57
TC90838 homologue to GP|2281647|gb|AAC49777.1| AP2 domain contai... 31 0.74
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 31 0.74
AJ499987 30 0.97
TC82370 similar to GP|15028377|gb|AAK76665.1 putative heat shock... 30 1.3
TC81380 similar to GP|19310773|gb|AAL85117.1 unknown protein {Ar... 30 1.3
>AL370225 homologue to GP|21536636|gb| unknown {Arabidopsis thaliana},
partial (28%)
Length = 408
Score = 236 bits (603), Expect = 7e-63
Identities = 116/116 (100%), Positives = 116/116 (100%)
Frame = +3
Query: 1 MNEKIENKDQIQQNQESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRA 60
MNEKIENKDQIQQNQESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRA
Sbjct: 60 MNEKIENKDQIQQNQESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRA 239
Query: 61 KLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNKIESMKKRYRSESASSDVSSWPLY 116
KLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNKIESMKKRYRSESASSDVSSWPLY
Sbjct: 240 KLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKNKIESMKKRYRSESASSDVSSWPLY 407
>TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsis
thaliana}, partial (26%)
Length = 431
Score = 199 bits (506), Expect = 1e-51
Identities = 106/109 (97%), Positives = 106/109 (97%)
Frame = +1
Query: 217 EQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEA 276
EQRSKKRRMRTE NNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEA
Sbjct: 1 EQRSKKRRMRTETNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEA 180
Query: 277 KRGEMEIKRTEIIANTQLEIARIFANVNNKDKDKDKDVDVDSSLRIGRS 325
KRGEMEIKRTEIIANTQLEIARIFANVNN KDKDKDVDVDSSLRIGRS
Sbjct: 181 KRGEMEIKRTEIIANTQLEIARIFANVNN--KDKDKDVDVDSSLRIGRS 321
>TC81757 similar to GP|3341693|gb|AAC27475.1| unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 975
Score = 67.0 bits (162), Expect = 9e-12
Identities = 68/254 (26%), Positives = 105/254 (40%), Gaps = 10/254 (3%)
Frame = +3
Query: 38 WSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRS--NSTKSPKTQTQCKNKIE 95
W+ L+ AY+ KW R L+G WE+VA V++R + KT QC++K+E
Sbjct: 135 WTHQETINLIRAYQDKWYSLKRGPLRGSQWEEVAVVVAARCGYDYNHPSKTALQCRHKME 314
Query: 96 SMKKRYRSE----SASSDVS---SWPLYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSS 148
+++R+RSE +A+S V+ SW +D L RG +S L++N
Sbjct: 315 KLRQRHRSEKRRLTATSSVASSRSWQYLRLMDDLERGPLPISVRPLSHNH---------- 464
Query: 149 LVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIAN-KNPLETDSS 207
PI S S+G ++S +I N K ETD
Sbjct: 465 -----------------PI-----SDDSDGA-------AARSRSIHNILNQKQRDETD-- 551
Query: 208 TPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEI 267
EK++ +K G+ +R AE ++ E +ME +KE
Sbjct: 552 -----EEKEDVMAK-------------------GLTAELRSFAERIIGLENMKMEMMKET 659
Query: 268 EKMRVEAEAKRGEM 281
E+ R+E E KR M
Sbjct: 660 ERFRLEMENKRIRM 701
>CB891664 similar to GP|3341693|gb| unknown protein {Arabidopsis thaliana},
partial (26%)
Length = 580
Score = 63.9 bits (154), Expect = 8e-11
Identities = 35/121 (28%), Positives = 58/121 (47%), Gaps = 7/121 (5%)
Frame = +1
Query: 26 IGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPK 85
I RL WS L+++Y KW R LK W++VA VS+R + K
Sbjct: 124 IPPSTRRLPPPCWSPEETLALIDSYREKWYSLGRGNLKATHWQEVADSVSNRCPNVTPAK 303
Query: 86 TQTQCKNKIESMKKRYRSE-------SASSDVSSWPLYSRLDLLLRGTGQLSTTTLNNNQ 138
T QC++K+E ++KRYR+E S S+W + +D + +G + + +++
Sbjct: 304 TPVQCRHKMEKLRKRYRTEIQRARSLPHSRFNSAWVHFKLMDSMEKGPSPVKSENNDSDS 483
Query: 139 A 139
A
Sbjct: 484 A 486
>TC83082 similar to GP|18149189|dbj|BAB83610. 6b-interacting protein 1
{Nicotiana tabacum}, partial (19%)
Length = 374
Score = 48.1 bits (113), Expect = 4e-06
Identities = 22/60 (36%), Positives = 40/60 (66%), Gaps = 1/60 (1%)
Frame = +2
Query: 33 LKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSS-RSNSTKSPKTQTQCK 91
++ D WSE A +TL++A+ +++ NR L+ +DW+DVA V++ ++ K+ +T QCK
Sbjct: 194 VREDCWSEDASSTLVDAWGRRYLELNRGNLRQKDWQDVADAVNALHGHTKKTHRTDVQCK 373
>TC90974 weakly similar to GP|9294038|dbj|BAB01995.1 gene_id:MOB24.1~unknown
protein {Arabidopsis thaliana}, partial (15%)
Length = 690
Score = 44.7 bits (104), Expect = 5e-05
Identities = 22/68 (32%), Positives = 38/68 (55%)
Frame = +1
Query: 33 LKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVSSRSNSTKSPKTQTQCKN 92
L +EW+E A LLE + K++ R L+ ++W DVA+ V S K + QC++
Sbjct: 496 LGSEEWTEHATFVLLEIWGEKFLQIGRNSLRSEEWHDVAEKV---SEELKVERNVAQCRS 666
Query: 93 KIESMKKR 100
++ +K+R
Sbjct: 667 VLDKVKRR 690
>AL369166 similar to GP|3341693|gb|A unknown protein {Arabidopsis thaliana},
partial (13%)
Length = 505
Score = 43.1 bits (100), Expect = 1e-04
Identities = 26/71 (36%), Positives = 39/71 (54%)
Frame = +2
Query: 242 IAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIARIFA 301
+ +I+ + + VR EQ +ME +EIE MR+E ME+KRTE+I +Q I FA
Sbjct: 5 MVNAIKVLRDGFVRMEQMKMEMAREIETMRME-------MEMKRTEMILESQQRIVEAFA 163
Query: 302 NVNNKDKDKDK 312
++ K K
Sbjct: 164 KAISEKNKKRK 196
>TC88284 similar to PIR|T03793|T03793 calmodulin-binding protein TCB60 -
common tobacco, partial (21%)
Length = 1044
Score = 40.8 bits (94), Expect = 7e-04
Identities = 34/111 (30%), Positives = 49/111 (43%), Gaps = 6/111 (5%)
Frame = +1
Query: 96 SMKKRYRSESASSDVSSWP-----LYSRLDLLLRGTGQLSTTTLNNNQAGLVLLEPSSLV 150
SM RY S++ + SS LY D LL Q ++ +++ GL L P S
Sbjct: 106 SMVTRYPSQALIGNSSSRSHFDDSLYLSNDHLLGNAHQSQSSRNDHSTVGLALGPPQSST 285
Query: 151 MS-HAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKN 200
HA QP P P N+ GVD EDEI +S++ + N++
Sbjct: 286 SGFHAGSSSMQPPAPNPFDDWSNNR-DKGVDDFFSEDEIRVRSNEILENED 435
>TC79702 MtN14
Length = 973
Score = 35.8 bits (81), Expect = 0.023
Identities = 36/158 (22%), Positives = 64/158 (39%), Gaps = 1/158 (0%)
Frame = +2
Query: 142 VLLEPSSLVMSHAQEEVHQPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNP 201
++ P +L S +EE P + A+ V +K+ E ETK + K P
Sbjct: 149 IVESPITLAASETKEESSAEEAKP-LVKAETKEVVEEVVVPVKDTEEETKKEEQTETKEP 325
Query: 202 L-ETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTR 260
+ ET + +L E+ ++ + ++ ET+ + + VAE +
Sbjct: 326 VEETKENGNSLNVEETKENGDS--VVEAVQEKPAEESETVNVVKDENVVAEPETKDNVKT 499
Query: 261 METLKEIEKMRVEAEAKRGEMEIKRTEIIANTQLEIAR 298
ET +E + +VE E E K E+I N +I +
Sbjct: 500 EETSEEKNEEKVEKEDAMDEK--KEEEVITNNDAKIEK 607
>BF644858 weakly similar to GP|5679842|emb l1332.6 {Oryza sativa}, partial
(49%)
Length = 692
Score = 33.5 bits (75), Expect = 0.11
Identities = 31/131 (23%), Positives = 59/131 (44%), Gaps = 21/131 (16%)
Frame = +2
Query: 186 DEIETKSSDHIANKNPLETDSSTPALYSEKDEQRSKKRRMRTENNN-----NKRQKKETM 240
DEI + I+ E + A K EQ++KKR++ E + ++Q KE
Sbjct: 212 DEILARHRKEISQLQKKEVEMKKQAARGSKAEQKAKKRQVEEEVSQLSAKLKEKQAKELS 391
Query: 241 GIAES-------------IRWVAEVVVRS--EQTRMETLKEIEKMRVEAEAKRGE-MEIK 284
+ S ++ +A V V S E T++ K+ R + EA+R + ++ +
Sbjct: 392 ALGYSSGIGNEKSNMDNLVKAIAGVSVSSQPENTKVSKAKQRRDKRAQQEAEREQRIQAE 571
Query: 285 RTEIIANTQLE 295
+ +II++ +E
Sbjct: 572 QNDIISDRTVE 604
>TC81121 homologue to GP|18182311|gb|AAL65125.1 GT-2 factor {Glycine max},
partial (37%)
Length = 794
Score = 33.5 bits (75), Expect = 0.11
Identities = 25/107 (23%), Positives = 44/107 (40%)
Frame = +3
Query: 16 ESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLEAYESKWVLRNRAKLKGQDWEDVAKHVS 75
E K+ NI G G R R E LL+ + + LKG WE+V++ ++
Sbjct: 246 EEDDKMNININGGGGNRWPRQE-----TLALLKIRSDMDGVFRDSSLKGPLWEEVSRKLA 410
Query: 76 SRSNSTKSPKTQTQCKNKIESMKKRYRSESASSDVSSWPLYSRLDLL 122
S K + + +N + K+ S S+ ++ + +L L
Sbjct: 411 DLGYHRSSKKCKEKFENVYKYHKRTKEGRSGKSEGKTYRFFDQLQAL 551
>TC92575 weakly similar to GP|21428434|gb|AAM49877.1 LD13350p {Drosophila
melanogaster}, partial (3%)
Length = 557
Score = 33.1 bits (74), Expect = 0.15
Identities = 24/111 (21%), Positives = 54/111 (48%), Gaps = 4/111 (3%)
Frame = -1
Query: 180 DKLIKEDEIETKSSDHIANKNPLETDSSTPALYSEKDE----QRSKKRRMRTENNNNKRQ 235
DKL +E E+ + + + K P ++ KDE ++S+K R ++ + +
Sbjct: 452 DKLSEETELSKEERESVEAKKPELDAMDEESILEGKDEGSEKEKSQKDREDDKDKVDNKS 273
Query: 236 KKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEAKRGEMEIKRT 286
+E + + + ++ + V E + + +KE++K+ EAK +E +R+
Sbjct: 272 NEENIKVEKRLKKRGKGKVNGENVK-KKMKELKKIEDIPEAKDESIEKERS 123
>AJ499968 weakly similar to GP|23495961|gb| hypothetical protein {Plasmodium
falciparum 3D7}, partial (19%)
Length = 342
Score = 32.7 bits (73), Expect = 0.20
Identities = 22/68 (32%), Positives = 34/68 (49%), Gaps = 4/68 (5%)
Frame = -3
Query: 214 EKDEQ----RSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEK 269
E++EQ + K+ + R E +R + E + + R E R E+ R+E + IEK
Sbjct: 307 EREEQERIAKEKEEQERKEREERERLELERIAKEKEERERKEKEEREERERLEAVVRIEK 128
Query: 270 MRVEAEAK 277
R EAE K
Sbjct: 127 ERKEAEEK 104
>TC79472 similar to PIR|C96685|C96685 unknown protein F15E12.10 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1001
Score = 32.3 bits (72), Expect = 0.26
Identities = 30/126 (23%), Positives = 48/126 (37%), Gaps = 20/126 (15%)
Frame = +2
Query: 15 QESPRKLQNNIIGSGNERLKRDEWSEGAVTTLLE--------------AYESKWVLRN-- 58
+E+ K + G GN+ D + + T LE +Y +L+N
Sbjct: 530 EEADYKATKELFGGGNDEKNLDTFIPKSETDFLEYAELISHRLRAFEKSYHYMGLLKNVM 709
Query: 59 ---RAKLKGQDWEDVAKHVSSRSN-STKSPKTQTQCKNKIESMKKRYRSESASSDVSSWP 114
LKG D +D+A V++ +N K+ K K K KK+ + D +
Sbjct: 710 RISMTSLKGADAKDIASSVTAIANEKIKAEKEANAGKKKTGGKKKQLTVDKPDEDFVAAD 889
Query: 115 LYSRLD 120
Y LD
Sbjct: 890 RYDALD 907
>BG646992 similar to PIR|T06133|T061 hypothetical protein F23E12.200 -
Arabidopsis thaliana, partial (11%)
Length = 761
Score = 31.2 bits (69), Expect = 0.57
Identities = 38/168 (22%), Positives = 70/168 (41%), Gaps = 2/168 (1%)
Frame = +1
Query: 156 EEVH--QPLVPPPITTAQNSHGSNGVDKLIKEDEIETKSSDHIANKNPLETDSSTPALYS 213
++VH Q LVPPP + N H +G +EDE E + H+ K + D
Sbjct: 157 KQVHGDQKLVPPPNSNHSNDHHDHG-HLEDEEDESEVEYEVHVVEKKVVNDDG------- 312
Query: 214 EKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVE 273
K + +SK N N + +A I+ + + S + + + E+ K+R
Sbjct: 313 -KPKSKSKPNSAFRPGNRN------PLEVAREIQILFQRASDS-GSHIADILEVGKLRYH 468
Query: 274 AEAKRGEMEIKRTEIIANTQLEIARIFANVNNKDKDKDKDVDVDSSLR 321
+A +M ++A + ++ N + D + D+DV+ + R
Sbjct: 469 HKAATSKM----LHLVAPSLSVVSSASRNAQSGDAN-SVDLDVELTTR 597
>TC90838 homologue to GP|2281647|gb|AAC49777.1| AP2 domain containing
protein RAP2.11 {Arabidopsis thaliana}, partial (22%)
Length = 521
Score = 30.8 bits (68), Expect = 0.74
Identities = 33/103 (32%), Positives = 49/103 (47%), Gaps = 11/103 (10%)
Frame = +2
Query: 222 KRRMRTENNNNKRQKKETMGIAE--SIRWVAEVVVRSEQTRM-----ETLKEIEKMRVEA 274
K R R NN NK +G+ + S RWVAE+ +++ RM ET +E + EA
Sbjct: 128 KGRNRNSNNTNK-----FVGVRQRPSGRWVAEIKDTTQKIRMWLGTFETAEEAARAYDEA 292
Query: 275 EA-KRGEMEIKRTEIIANTQLE---IARIFANVNNKDKDKDKD 313
RG RT I + L+ +RI +NN+ DK ++
Sbjct: 293 ACLLRGSN--TRTNFITHVSLDSPLASRIRNLLNNRKGDKKQE 415
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 30.8 bits (68), Expect = 0.74
Identities = 43/172 (25%), Positives = 63/172 (36%), Gaps = 24/172 (13%)
Frame = +2
Query: 173 SHGSNGVDKLIKEDEI--------ETKSSDHIANKNPLETDSSTPALYSEKDEQRSKKRR 224
S G NG K E+ + E ++ K E DSS+ A S +
Sbjct: 710 SDGVNGAWKEELENAVQRYASIMTELDAAKQDLRKTRQEYDSSSDARVSAVKRTEEAENA 889
Query: 225 MRTENNNNKRQKKETMGIAESIRWVAEVVVRSEQTRMETLKEIEKMRVEAEA-----KRG 279
M+ KE + ESI V S+Q + L E + +R +A K+
Sbjct: 890 MKENTERVSELSKEISAVKESIEQTKLAYVESQQQQALVLTEKDALRQSYKATLEQSKKK 1069
Query: 280 EMEIKR---TEI-------IANTQLEIARIFANVNNK-DKDKDKDVDVDSSL 320
+ +K+ EI +A T EIA + + NK D D V S L
Sbjct: 1070LLALKKEFNPEITKSLEAQLAETMNEIAALHTELENKRSSDLDSVKTVTSEL 1225
>AJ499987
Length = 501
Score = 30.4 bits (67), Expect = 0.97
Identities = 16/64 (25%), Positives = 33/64 (51%)
Frame = +2
Query: 190 TKSSDHIANKNPLETDSSTPALYSEKDEQRSKKRRMRTENNNNKRQKKETMGIAESIRWV 249
T+S + NK+ T S S+++ + SKK+ ++ +K+QKK+ + +
Sbjct: 74 TRSFGNNTNKDRQRTSGSRQTDKSKQNSKNSKKKPQEAKSEKHKKQKKDKSHKTSRSKVL 253
Query: 250 AEVV 253
AE++
Sbjct: 254 AEIL 265
>TC82370 similar to GP|15028377|gb|AAK76665.1 putative heat shock factor
protein hsf8 {Arabidopsis thaliana}, partial (7%)
Length = 645
Score = 30.0 bits (66), Expect = 1.3
Identities = 16/52 (30%), Positives = 30/52 (56%), Gaps = 1/52 (1%)
Frame = +2
Query: 272 VEAEAKRGEMEIKRTEIIANTQLEIARIFA-NVNNKDKDKDKDVDVDSSLRI 322
VE E K+ +++ + + + I + ++ N+DKD D+DVD+ S LR+
Sbjct: 290 VEMEIKKPNQALRKPVNVTKSNVPIVILRKPSLYNEDKDGDEDVDMSSRLRM 445
>TC81380 similar to GP|19310773|gb|AAL85117.1 unknown protein {Arabidopsis
thaliana}, partial (57%)
Length = 1640
Score = 30.0 bits (66), Expect = 1.3
Identities = 30/101 (29%), Positives = 47/101 (45%), Gaps = 20/101 (19%)
Frame = +3
Query: 56 LRNRAKLKGQDWEDVAKHVSSRS----------NSTKSPKTQTQ------CKNKIESMKK 99
LRN+ KL+G+ W+D+ S S +S + K+Q + ++ I +
Sbjct: 879 LRNKVKLEGELWDDLVSSSSQNSHLVSKATHLWDSLLARKSQHEVLASGPIEDLIAHREH 1058
Query: 100 RYR----SESASSDVSSWPLYSRLDLLLRGTGQLSTTTLNN 136
RYR S A+ D SS YS D+L +G LS+ +N
Sbjct: 1059RYRISGSSLLAAMDQSSQAPYS--DVLSGDSGDLSSVQTDN 1175
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.306 0.123 0.328
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,834,379
Number of Sequences: 36976
Number of extensions: 95188
Number of successful extensions: 494
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 487
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 490
length of query: 325
length of database: 9,014,727
effective HSP length: 96
effective length of query: 229
effective length of database: 5,465,031
effective search space: 1251492099
effective search space used: 1251492099
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126006.11


