

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
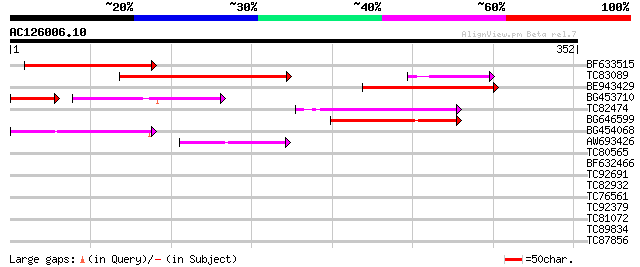
Query= AC126006.10 - phase: 0 /pseudo
(352 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF633515 172 2e-43
TC83089 139 2e-33
BE943429 103 1e-22
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 72 2e-20
TC82474 72 2e-13
BG646599 68 6e-12
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 64 7e-11
AW693426 homologue to GP|23307580|dbj similar to serine/threonin... 44 7e-05
TC80565 weakly similar to GP|14335072|gb|AAK59800.1 AT5g20050/F2... 30 1.4
BF632466 27 1.8
TC92691 weakly similar to GP|12231951|gb|AAG49320.1 JD1 {Nicotia... 29 2.4
TC82932 similar to PIR|T00538|T00538 probable serine proteinase ... 28 4.1
TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 28 4.1
TC92379 homologue to GP|6642635|gb|AAF20216.1| putative translat... 28 4.1
TC81072 similar to PIR|T05401|T05401 hypothetical protein F10M6.... 28 7.0
TC89834 28 7.0
TC87856 similar to GP|19699067|gb|AAL90901.1 AT5g20250/F5O24_140... 27 9.1
>BF633515
Length = 249
Score = 172 bits (435), Expect = 2e-43
Identities = 82/82 (100%), Positives = 82/82 (100%)
Frame = +2
Query: 10 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 69
LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 181
Query: 70 RDFYTTYLGRANLLASTEDPVE 91
RDFYTTYLGRANLLASTEDPVE
Sbjct: 182RDFYTTYLGRANLLASTEDPVE 247
>TC83089
Length = 887
Score = 139 bits (349), Expect = 2e-33
Identities = 67/107 (62%), Positives = 84/107 (77%)
Frame = +3
Query: 69 LRDFYTTYLGRANLLASTEDPVELEELARVRTYCVRCYLLYLIGCLLFGDRSNKRIELIY 128
LR++Y YL A L+ ++ + +EL ++RT CV+ YLLYL+GCLLFGD+SNKRIEL+Y
Sbjct: 264 LREYYEGYLDSATRLSDPQNLGDPQELEKIRTTCVKWYLLYLVGCLLFGDKSNKRIELVY 443
Query: 129 LTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDKALGGSVTLLT 175
LTTMEDGYAGMRN+SWGGMTLAYLY L++ PG AL SVTL+T
Sbjct: 444 LTTMEDGYAGMRNHSWGGMTLAYLYHCLSEVSLPGGGALERSVTLVT 584
Score = 49.7 bits (117), Expect = 2e-06
Identities = 24/54 (44%), Positives = 32/54 (58%)
Frame = +3
Query: 248 SGWIMCGVRRVYRHLPERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFVDFRTH 301
+GW +C HLPER LR YGY+QT+P PT + L +V AF++F H
Sbjct: 585 AGWFLC-------HLPERGLR*YGYVQTVPIPPTTIEPLEPAEVVTAFLEFALH 725
>BE943429
Length = 384
Score = 103 bits (256), Expect = 1e-22
Identities = 45/84 (53%), Positives = 58/84 (68%)
Frame = +2
Query: 220 DRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIPRH 279
D ++ V WRPYE R++ F+DV WYSGWIM G +++ RHLPERVLRQYGY+QT+PR
Sbjct: 2 DSMEHCHVIWRPYEHRRDVTPFQDVCWYSGWIMAGKQKMVRHLPERVLRQYGYVQTVPRP 181
Query: 280 PTDVVELPLPYIVQAFVDFRTHTL 303
PT +V L + AF +F H L
Sbjct: 182 PTTIVPLAPAEVATAFFEFVVHVL 253
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 72.4 bits (176), Expect(2) = 2e-20
Identities = 44/98 (44%), Positives = 58/98 (58%), Gaps = 3/98 (3%)
Frame = -3
Query: 40 LLMRHLGVSQQEAEKICGQEYGGYISYPRLRDFYTTYLGRANLLASTEDPV---ELEELA 96
L+ R LG+S A E G+ISYP L+ Y +L A L EDP+ EL+E A
Sbjct: 298 LMTRLLGMSDANAMAEIRTESAGHISYPTLKRVYEDHLIEARRL---EDPLTREELQERA 128
Query: 97 RVRTYCVRCYLLYLIGCLLFGDRSNKRIELIYLTTMED 134
R R +CVR LLYL+ C+LF ++N+ I+LIYL M D
Sbjct: 127 RRRQWCVRSLLLYLVDCVLFTYKTNRHIDLIYLDCMAD 14
Score = 44.3 bits (103), Expect(2) = 2e-20
Identities = 19/31 (61%), Positives = 24/31 (77%)
Frame = -1
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKK 31
+PMGEMT+TLDDV L H+PI G ML H ++
Sbjct: 414 LPMGEMTITLDDVYNLLHIPIHGCMLDHDEQ 322
>TC82474
Length = 992
Score = 72.4 bits (176), Expect = 2e-13
Identities = 37/103 (35%), Positives = 59/103 (56%)
Frame = -2
Query: 178 KLIFYFFVNVDPNTDYMENYPVAARWKLQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHRE 237
+++F F DP Y E+ P A R+ L GH+ Y + LDR+++DDV + Y+ H +
Sbjct: 910 EILFVF----DPK--YTEDRPHATRFSLAMGHKSPNNYMTFLDRVEVDDVSFNQYDFHHQ 749
Query: 238 IQDFEDVFWYSGWIMCGVRRVYRHLPERVLRQYGYMQTIPRHP 280
F+D+ + WI C + Y HL E V++ YG++Q+ RHP
Sbjct: 748 TCPFDDIS*FFRWIECDRKMNYHHLLELVMK*YGHIQSTLRHP 620
>BG646599
Length = 711
Score = 67.8 bits (164), Expect = 6e-12
Identities = 33/81 (40%), Positives = 50/81 (60%)
Frame = +3
Query: 200 AARWKLQKGHEEGIMYRSLLDRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMCGVRRVY 259
A RWK +G+ + R ++D + +DV W P++ HR++ F+D+ SG+I V
Sbjct: 153 AMRWKYPRGNLKVDQIRRMIDDLTPNDVIWCPFDSHRQVIPFDDICLSSGYIR-WCSNVV 329
Query: 260 RHLPERVLRQYGYMQTIPRHP 280
+LPER LRQ+GY+Q IPR P
Sbjct: 330 PYLPERCLRQFGYIQYIPRPP 392
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 64.3 bits (155), Expect = 7e-11
Identities = 33/94 (35%), Positives = 54/94 (57%), Gaps = 3/94 (3%)
Frame = +3
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEY 60
+P+GEMT+ LDD++CL H+P+ G +L H + + H+G L+ +LG+ +E +
Sbjct: 387 LPVGEMTIALDDISCLLHIPVGGNLLFH-ESLSIHQGTEYLVNYLGLEFEEIAAETKRLK 563
Query: 61 GGYISYPRLRDFYTTYLGRANLLAS---TEDPVE 91
+I+Y L YT+YL A A+ ED +E
Sbjct: 564 SAHITYDTLLSIYTSYLTEAKSYANQPGEEDSME 665
>AW693426 homologue to GP|23307580|dbj similar to serine/threonine
phosphatase PP7 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 209
Score = 44.3 bits (103), Expect = 7e-05
Identities = 25/69 (36%), Positives = 36/69 (51%)
Frame = +1
Query: 106 YLLYLIGCLLFGDRSNKRIELIYLTTMEDGYAGMRNYSWGGMTLAYLYGELADACRPGDK 165
+LL L+GC LF D+S + + YL D Y+WG LA+LY L A K
Sbjct: 1 FLLCLVGCTLFSDKSTFVVNVAYLEFFRD-LDSCGGYAWGVAALAHLYDNLRYASFHHMK 177
Query: 166 ALGGSVTLL 174
++ G +TL+
Sbjct: 178 SISGYLTLI 204
>TC80565 weakly similar to GP|14335072|gb|AAK59800.1 AT5g20050/F28I16_200
{Arabidopsis thaliana}, partial (81%)
Length = 1581
Score = 30.0 bits (66), Expect = 1.4
Identities = 17/42 (40%), Positives = 22/42 (51%), Gaps = 3/42 (7%)
Frame = +2
Query: 81 NLLASTEDPVELEELARVRTYC---VRCYLLYLIGCLLFGDR 119
+LLA EDP +L LAR+ T V C Y +G +F R
Sbjct: 1325 DLLADDEDPTDLNNLARLLTPVSSNVECTSTYSLGSTIFSGR 1450
>BF632466
Length = 579
Score = 26.6 bits (57), Expect(2) = 1.8
Identities = 20/65 (30%), Positives = 30/65 (45%), Gaps = 1/65 (1%)
Frame = -1
Query: 34 KHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRLRDFY-TTYLGRANLLASTEDPVEL 92
KH L +RHL +++ K+C + + LR F T + ++ LL S D L
Sbjct: 309 KHVSLHLQLRHLDEARKLVAKLCSGKLAESVQLWELRVFIEITCITQSFLLPSDTDLSSL 130
Query: 93 EELAR 97
EL R
Sbjct: 129 FELLR 115
Score = 21.6 bits (44), Expect(2) = 1.8
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = -2
Query: 101 YCVRCYLLYLIGCLL 115
Y V YLL+L+ CL+
Sbjct: 62 YVVTTYLLFLLPCLV 18
>TC92691 weakly similar to GP|12231951|gb|AAG49320.1 JD1 {Nicotiana
tabacum}, partial (47%)
Length = 696
Score = 29.3 bits (64), Expect = 2.4
Identities = 16/38 (42%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Frame = +3
Query: 37 GAALLMRHLGVSQ--QEAEKICGQEYGGYISYPRLRDF 72
G+A +M+HLG +Q + A CG GYI LRD+
Sbjct: 492 GSAHVMKHLGFNQTCEAAFAECGGSVKGYIVEDELRDY 605
>TC82932 similar to PIR|T00538|T00538 probable serine proteinase At2g19170
[imported] - Arabidopsis thaliana, partial (32%)
Length = 816
Score = 28.5 bits (62), Expect = 4.1
Identities = 18/70 (25%), Positives = 32/70 (45%), Gaps = 5/70 (7%)
Frame = -3
Query: 264 ERVLRQYGYMQTIPRHPTDVVELPLPYIVQAFVDFRTHTLKAAIGVSRQESRHG-----G 318
+++L Q Y Q R + + PLP+++ +F+D H I + S +
Sbjct: 538 DQILHQGKYFQH--REQIQLHQGPLPFLILSFIDAVPHNKVERISAEKNISLYQFLV**F 365
Query: 319 WLMAICYGTL 328
W +AIC+ L
Sbjct: 364 WSLAICWAVL 335
>TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 1054
Score = 28.5 bits (62), Expect = 4.1
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 7/63 (11%)
Frame = +3
Query: 94 ELARVRTYCVRCYLLYLIGCLLFGD-------RSNKRIELIYLTTMEDGYAGMRNYSWGG 146
+L V Y + CYL+YL+ LL D +N+ + L+ T +ED +R +G
Sbjct: 213 DLRSVLCYLLLCYLIYLLLLLLLYDLILLLLTHTNRYVHLLL*TKVEDRSVTIRF*IYGS 392
Query: 147 MTL 149
+ L
Sbjct: 393 VFL 401
>TC92379 homologue to GP|6642635|gb|AAF20216.1| putative translation
initiation factor EIF-2B beta subunit {Arabidopsis
thaliana}, partial (20%)
Length = 690
Score = 28.5 bits (62), Expect = 4.1
Identities = 30/124 (24%), Positives = 46/124 (36%), Gaps = 2/124 (1%)
Frame = +2
Query: 153 YGELADACRPGDKALGGSVTLLTIRKLIFYFFVNVDPNTDYMENYPVAARWKLQKGHEEG 212
+GE +D D A GGS+ + V+P DY+ V+ GH
Sbjct: 89 FGEFSDCM---DSASGGSLHV-------------VNPTFDYVPPKLVSLFITDTGGHNPS 220
Query: 213 IMYRSLLDRIQLDDVCWRPYEEHREIQDFEDVFWYSGWIMCGVRRV--YRHLPERVLRQY 270
MYR + D DD+ + Y+G + C ++R R V+ +
Sbjct: 221 YMYRLIADYYSADDLVVKRRPATTS*VQLAQFLSYNGCLCCPLKRYVKMREKMNLVVLRI 400
Query: 271 GYMQ 274
G MQ
Sbjct: 401 GRMQ 412
>TC81072 similar to PIR|T05401|T05401 hypothetical protein F10M6.90 -
Arabidopsis thaliana, partial (13%)
Length = 745
Score = 27.7 bits (60), Expect = 7.0
Identities = 11/39 (28%), Positives = 25/39 (63%), Gaps = 3/39 (7%)
Frame = +2
Query: 80 ANLLASTEDPVELEELARV---RTYCVRCYLLYLIGCLL 115
A+++ S++ P + R+ +T+C+RC L+Y++ C +
Sbjct: 302 ASIIGSSQKPNR*QTFQRLCYDQTWCLRCNLVYVLRCFV 418
>TC89834
Length = 771
Score = 27.7 bits (60), Expect = 7.0
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = +2
Query: 305 AAIGVSRQESRHGGWLMAICYGTLGCLTLILCHLSLEIF 343
A + +S G+ + + Y LGCLT++L + LE+F
Sbjct: 554 AIMSLSFSRQSQCGFEVDLLYFFLGCLTVLLMKIKLELF 670
>TC87856 similar to GP|19699067|gb|AAL90901.1 AT5g20250/F5O24_140
{Arabidopsis thaliana}, partial (45%)
Length = 1402
Score = 27.3 bits (59), Expect = 9.1
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 2/36 (5%)
Frame = +2
Query: 102 CVRCYLL-YLIGCLLFGDR-SNKRIELIYLTTMEDG 135
C+ C ++ L+G L+ G S KRI L+ L T+EDG
Sbjct: 662 CLECIIVKVLLGVLMRGRMLSMKRILLLLLATLEDG 769
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.143 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,427,243
Number of Sequences: 36976
Number of extensions: 186776
Number of successful extensions: 1155
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1153
length of query: 352
length of database: 9,014,727
effective HSP length: 97
effective length of query: 255
effective length of database: 5,428,055
effective search space: 1384154025
effective search space used: 1384154025
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126006.10


