

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
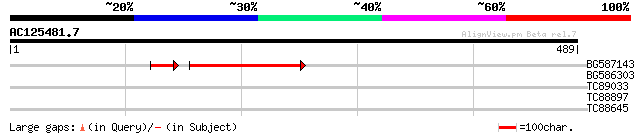
Query= AC125481.7 + phase: 0 /pseudo
(489 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F2... 100 2e-21
BG586303 weakly similar to PIR|A96586|A96 hypothetical protein F... 36 0.029
TC89033 similar to GP|11994412|dbj|BAB02414. chloroplast nucleoi... 30 2.7
TC88897 similar to PIR|S62699|S62699 photoassimilate-responsive ... 29 4.7
TC88645 similar to PIR|S62700|S62700 photoassimilate-responsive ... 28 8.0
>BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (12%)
Length = 717
Score = 99.8 bits (247), Expect = 2e-21
Identities = 47/102 (46%), Positives = 69/102 (67%), Gaps = 2/102 (1%)
Frame = -3
Query: 156 HVPCGGARPFSPCMDRGRCTKYYRKNYKGTTTIDE-GYPRYKRR-DFGIHVDKQRVLLDN 213
H PCG P SPCM+ CTK Y + Y G+T+ID+ GY Y+RR + HV K +L+N
Sbjct: 415 HGPCGVINPKSPCMENNVCTKKYPRPYNGSTSIDKSGYVLYRRR*NETEHVVKNGAILNN 236
Query: 214 RYVVPYNPHLLIRYGGHVNVECCNKSNSIKYLFKYVNKVPDK 255
++VP+N LL +Y H+N+E CN+++++KYLFKY+ K D+
Sbjct: 235 TFIVPHNIKLLKKYEAHINMEWCNRTSAVKYLFKYITKGVDR 110
Score = 42.0 bits (97), Expect = 5e-04
Identities = 17/24 (70%), Positives = 22/24 (90%)
Frame = -3
Query: 122 SDRPDIVCRVFKMKLDKLMDDFEK 145
+D+PDI CRVFKMKLD+L+ DF+K
Sbjct: 691 NDKPDIECRVFKMKLDELLKDFKK 620
>BG586303 weakly similar to PIR|A96586|A96 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (3%)
Length = 698
Score = 36.2 bits (82), Expect = 0.029
Identities = 16/24 (66%), Positives = 19/24 (78%)
Frame = -3
Query: 122 SDRPDIVCRVFKMKLDKLMDDFEK 145
+DR D+ RVFKMKLD+LM DF K
Sbjct: 666 NDRHDLKSRVFKMKLDELMSDFNK 595
>TC89033 similar to GP|11994412|dbj|BAB02414. chloroplast nucleoid DNA
binding protein-like {Arabidopsis thaliana}, partial
(11%)
Length = 1051
Score = 29.6 bits (65), Expect = 2.7
Identities = 11/21 (52%), Positives = 17/21 (80%)
Frame = -2
Query: 307 LKLLFHLHNEQGNIDHVSRMV 327
L+L+FHLH GN+D++S +V
Sbjct: 222 LRLIFHLHFHMGNLDYLSLIV 160
>TC88897 similar to PIR|S62699|S62699 photoassimilate-responsive protein
PAR-1b precursor - common tobacco, partial (77%)
Length = 920
Score = 28.9 bits (63), Expect = 4.7
Identities = 10/34 (29%), Positives = 20/34 (58%)
Frame = +1
Query: 116 ERNLKASDRPDIVCRVFKMKLDKLMDDFEKEECL 149
E+++K + CR +++ DKL D E ++C+
Sbjct: 262 EKHVKRNGEEKYTCRTSEIEADKLKDHIESDQCI 363
>TC88645 similar to PIR|S62700|S62700 photoassimilate-responsive protein
PAR-1c precursor - common tobacco, partial (90%)
Length = 1164
Score = 28.1 bits (61), Expect = 8.0
Identities = 9/34 (26%), Positives = 20/34 (58%)
Frame = +2
Query: 116 ERNLKASDRPDIVCRVFKMKLDKLMDDFEKEECL 149
E+++K + C+ +++ DKL D E ++C+
Sbjct: 254 EKHVKRTGEEAYTCKTLEIEADKLKDHIESDQCI 355
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.342 0.150 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,110,612
Number of Sequences: 36976
Number of extensions: 234456
Number of successful extensions: 1454
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1437
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1453
length of query: 489
length of database: 9,014,727
effective HSP length: 100
effective length of query: 389
effective length of database: 5,317,127
effective search space: 2068362403
effective search space used: 2068362403
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC125481.7


