

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.3 + phase: 0
(476 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
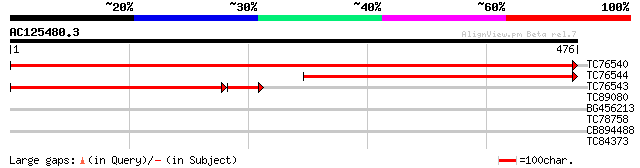
TC76540 homologue to GP|6581004|gb|AAF18411.1| putative integral... 930 0.0
TC76544 homologue to GP|6581004|gb|AAF18411.1| putative integral... 456 e-129
TC76543 homologue to GP|6581004|gb|AAF18411.1| putative integral... 355 e-110
TC89080 GP|9087331|dbj|BAA99475.1 orf190 {Beta vulgaris}, partia... 29 4.5
BG456213 28 5.9
TC78758 similar to PIR|T49962|T49962 hypothetical protein F8M21.... 28 5.9
CB894488 similar to GP|6437560|gb| hypothetical protein {Arabido... 28 5.9
TC84373 similar to GP|15636781|gb|AAL02122.1 copper-transporting... 28 5.9
>TC76540 homologue to GP|6581004|gb|AAF18411.1| putative integral membrane
protein {Phaseolus vulgaris}, complete
Length = 2365
Score = 930 bits (2404), Expect = 0.0
Identities = 474/476 (99%), Positives = 476/476 (99%)
Frame = +3
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 75 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 254
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 255 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 434
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILII+QLFFAGIIVICLDELLQKGYGLGS
Sbjct: 435 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIILQLFFAGIIVICLDELLQKGYGLGS 614
Query: 181 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 240
GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ
Sbjct: 615 GISLFIATNICENIIWKAFSPTTINSGRGAEFEGAVIALFHLLITRTDKVRALREAFYRQ 794
Query: 241 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 300
NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL
Sbjct: 795 NLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSAL 974
Query: 301 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 360
VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP
Sbjct: 975 VSNLYFISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANP 1154
Query: 361 FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 420
FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP
Sbjct: 1155FHALFYLVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIP 1334
Query: 421 TAAAFGGICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
TAAAFGG+CIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF
Sbjct: 1335TAAAFGGMCIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 1502
>TC76544 homologue to GP|6581004|gb|AAF18411.1| putative integral membrane
protein {Phaseolus vulgaris}, partial (48%)
Length = 844
Score = 456 bits (1173), Expect = e-129
Identities = 230/230 (100%), Positives = 230/230 (100%)
Frame = +2
Query: 247 NLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSALVSNLYF 306
NLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSALVSNLYF
Sbjct: 5 NLLATVLIFLIVIYFQGFRVVLPVRSKNARGQQGSYPIKLFYTSNMPIILQSALVSNLYF 184
Query: 307 ISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANPFHALFY 366
ISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANPFHALFY
Sbjct: 185 ISQLLHRKYSGNFIVNLLGKWKESEYGGGHSIPVGGIAYYITAPSSLADMAANPFHALFY 364
Query: 367 LVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFG 426
LVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFG
Sbjct: 365 LVFMLSACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFG 544
Query: 427 GICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 476
GICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF
Sbjct: 545 GICIGALTVLADFMGAIGSGTGILLAVTIIYQYFETFEKERASELGFFGF 694
>TC76543 homologue to GP|6581004|gb|AAF18411.1| putative integral membrane
protein {Phaseolus vulgaris}, partial (44%)
Length = 773
Score = 355 bits (910), Expect(2) = e-110
Identities = 182/182 (100%), Positives = 182/182 (100%)
Frame = +2
Query: 1 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 60
MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG
Sbjct: 104 MGGGFRVLHLVRPFLSFLPEVQTADRKVPFREKVIYTVISLFIFLVCSQLPLYGIHSTTG 283
Query: 61 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 120
ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK
Sbjct: 284 ADPFYWMRVILASNRGTVMELGITPIVTSGLVMQLLAGSKIIEVDNNVREDRALLNGAQK 463
Query: 121 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 180
LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS
Sbjct: 464 LLGILIAVGEAVAYVLSGMYGSVGQLGVGNAILIIVQLFFAGIIVICLDELLQKGYGLGS 643
Query: 181 GI 182
GI
Sbjct: 644 GI 649
Score = 63.5 bits (153), Expect(2) = e-110
Identities = 29/30 (96%), Positives = 29/30 (96%)
Frame = +3
Query: 184 LFIATNICENIIWKAFSPTTINSGRGAEFE 213
LFIATNICENIIW AFSPTTINSGRGAEFE
Sbjct: 654 LFIATNICENIIWXAFSPTTINSGRGAEFE 743
>TC89080 GP|9087331|dbj|BAA99475.1 orf190 {Beta vulgaris}, partial (5%)
Length = 619
Score = 28.9 bits (63), Expect = 4.5
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Frame = +1
Query: 220 FHLLITRTDK-VRALREAFYRQNLPNVTNLLATVLIFLIVIYFQGFRVVLPVRSK 273
FHL+++ +K + L YR+ LP TNLL+ +LI I R+ L + K
Sbjct: 175 FHLILSFVEKMILLLFPEHYRKKLPMNTNLLSDLLILPERIMVPILRIYLQLNPK 339
>BG456213
Length = 628
Score = 28.5 bits (62), Expect = 5.9
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +2
Query: 376 LFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIPTAA 423
+++ W+ SGSSAR L Q + P S L +R +PTA+
Sbjct: 398 VYTVLWVMSSGSSARSPRTSLFRSQPIFP---RSTLSAFTDRLLPTAS 532
>TC78758 similar to PIR|T49962|T49962 hypothetical protein F8M21.160 -
Arabidopsis thaliana, partial (65%)
Length = 2065
Score = 28.5 bits (62), Expect = 5.9
Identities = 19/59 (32%), Positives = 27/59 (45%)
Frame = +3
Query: 373 ACALFSKTWIEVSGSSARDVAKQLKEQQMVMPGHRESNLQKELNRYIPTAAAFGGICIG 431
ACAL S +++SG +A V K+ Q H S Q L +P+ GG +G
Sbjct: 591 ACALRSDELVQISGDAA--VVKKALHQIASRLHHNPSRTQHLLGSAVPSVYPSGGSLMG 761
>CB894488 similar to GP|6437560|gb| hypothetical protein {Arabidopsis
thaliana}, partial (37%)
Length = 830
Score = 28.5 bits (62), Expect = 5.9
Identities = 32/119 (26%), Positives = 53/119 (43%), Gaps = 8/119 (6%)
Frame = +2
Query: 152 ILIIVQLFFAGIIVICLDELLQKGYGLGSGISLFIATNICENIIWKAFSPTTINSGRGAE 211
+ + + L II + + LL+ GL I L + ++C +II N R A
Sbjct: 305 VFVALVLGVGSIIPVTVPRLLELLLGLLGSI-LNVLDSLCASII*---GQDLANLVRLAS 472
Query: 212 F------EGAVIALFHLLITRTDKVRALREAFY--RQNLPNVTNLLATVLIFLIVIYFQ 262
E ++ALFHL+I VR R +Q+ ++ L+ ++ FL+V FQ
Sbjct: 473 IITQNRQEPLMLALFHLIIMDIRYVRVKRNVHIL*KQDNASLVQLVNLIIQFLLVYKFQ 649
>TC84373 similar to GP|15636781|gb|AAL02122.1 copper-transporting P-type
ATPase {Brassica napus}, partial (23%)
Length = 734
Score = 28.5 bits (62), Expect = 5.9
Identities = 25/104 (24%), Positives = 44/104 (42%), Gaps = 3/104 (2%)
Frame = +1
Query: 37 TVISLFIFLVCSQLPLYGIHSTTGADPFYWMRVILASN---RGTVMELGITPIVTSGLVM 93
T+ LFI +C +PL+ + PF + ++ G ++ + ++ G+
Sbjct: 385 TIFRLFISSLCLSVPLFLMKVVCPHIPFMYSLLLWRCGPFLMGDWLKWALVSVIQFGIGK 564
Query: 94 QLLAGSKIIEVDNNVREDRALLNGAQKLLGILIAVGEAVAYVLS 137
+ V RAL NG+ + +LIAVG +YV S
Sbjct: 565 RFY-----------VAAGRALRNGSTN-MDVLIAVGTTASYVYS 660
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.143 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,380,770
Number of Sequences: 36976
Number of extensions: 182101
Number of successful extensions: 1181
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1170
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1181
length of query: 476
length of database: 9,014,727
effective HSP length: 100
effective length of query: 376
effective length of database: 5,317,127
effective search space: 1999239752
effective search space used: 1999239752
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC125480.3


