

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
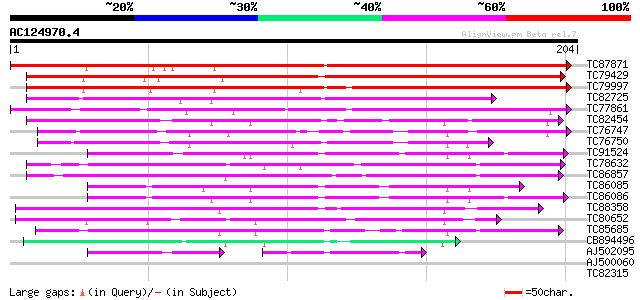
Score E
Sequences producing significant alignments: (bits) Value
TC87871 similar to PIR|T03803|T03803 tumor-related protein clon... 186 5e-48
TC79429 similar to PIR|T03803|T03803 tumor-related protein clon... 182 6e-47
TC79997 similar to PIR|T03803|T03803 tumor-related protein clon... 171 2e-43
TC82725 weakly similar to PIR|T07871|T07871 miraculin homolog r... 132 9e-32
TC77861 weakly similar to GP|13899081|gb|AAK48962.1 Unknown prot... 129 1e-30
TC82454 weakly similar to GP|3551243|dbj|BAA32820.1 Water-Solubl... 80 4e-16
TC76747 similar to GP|20269067|emb|CAD29731. protease inhibitor ... 75 2e-14
TC76750 similar to GP|20269067|emb|CAD29731. protease inhibitor ... 75 2e-14
TC91524 similar to GP|15217238|gb|AAK92582.1 Hypothetical protei... 70 7e-13
TC78632 similar to GP|9367042|gb|AAF87095.1| trypsin inhibitor {... 70 7e-13
TC86857 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {C... 68 3e-12
TC86085 65 2e-11
TC86086 similar to GP|15155442|gb|AAK86324.1 AGR_C_899p {Agrobac... 61 3e-10
TC88358 proteinase inhibitor 20 [Medicago truncatula] 57 5e-09
TC80652 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {C... 51 3e-07
TC85685 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {C... 46 1e-05
CB894496 weakly similar to GP|9367042|gb|A trypsin inhibitor {Gl... 44 4e-05
AJ502095 32 7e-04
AJ500060 32 0.22
TC82315 similar to GP|4038594|emb|CAA10993.1 tDET1 protein {Lyco... 30 0.83
>TC87871 similar to PIR|T03803|T03803 tumor-related protein clone NF34 -
common tobacco, partial (36%)
Length = 905
Score = 186 bits (472), Expect = 5e-48
Identities = 102/214 (47%), Positives = 138/214 (63%), Gaps = 12/214 (5%)
Frame = +3
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQP--EEVRDTSGNLVRNSINYFILPSSI-QCGT--R 55
MK F L T +++PL GA + E+V DT G +R NY+I+P I +CG +
Sbjct: 15 MKNTLLAFFFLFTFLSSQPLLGAAEASNEQVVDTLGKKLRADANYYIIPVPIYKCGPYGK 194
Query: 56 CE-----MALLNTNKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNC 108
C +AL + KTCPLDVV + +A+ +F+P N KKGVIRVSTDLN+ S C
Sbjct: 195 CRSSGSSLALASIGKTCPLDVVVVDRYQALPLTFIPVNPKKGVIRVSTDLNIKFSSRATC 374
Query: 109 STSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRE 168
S+ VWK+D+ +V+ Q F+T GGV GNPG ET++NWFKIE++ YKLVFCP+V +
Sbjct: 375 LHHSM-VWKLDRFNVSKRQWFITIGGVAGNPGWETINNWFKIEKYGDAYKLVFCPSVVQS 551
Query: 169 CEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
+ +CKD+G+F+DEN N R LSD P VKFQ+A
Sbjct: 552 FKHMCKDVGVFVDENGNKRLALSDVPLKVKFQQA 653
>TC79429 similar to PIR|T03803|T03803 tumor-related protein clone NF34 -
common tobacco, partial (42%)
Length = 807
Score = 182 bits (463), Expect = 6e-47
Identities = 98/207 (47%), Positives = 130/207 (62%), Gaps = 13/207 (6%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILP-SSIQC--------GTR 55
+L +L +T+PL G PE+V DT G VR ++Y+I P + C G+
Sbjct: 38 ILLAVLFALSTQPLLGEADASPEQVVDTEGKKVRAGVDYYIRPVPTTPCDGRGPCVVGSG 217
Query: 56 CEMALLNTNKTCPLDVVEEE--EAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
+ + N+TCPL+VV E +F P N KKGVIRVSTDLN+ S T+C S
Sbjct: 218 FVLIARSPNETCPLNVVVVEGFRGQGVTFTPVNPKKGVIRVSTDLNIKTSLNTSCEES-- 391
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
T+W +D D +T Q FVTTGGV GNPG++TVDNWFKIE++E YK VFCPTVC C+V+C
Sbjct: 392 TIWTLDDFDSSTGQWFVTTGGVLGNPGKDTVDNWFKIEKYEDDYKFVFCPTVCNFCKVMC 571
Query: 174 KDIGIFLDENRNTRFVLSDFPFGVKFQ 200
+++GIF D N N R L+D P+ V+FQ
Sbjct: 572 RNVGIFRDSNGNQRVALTDVPYKVRFQ 652
>TC79997 similar to PIR|T03803|T03803 tumor-related protein clone NF34 -
common tobacco, partial (48%)
Length = 801
Score = 171 bits (433), Expect = 2e-43
Identities = 92/210 (43%), Positives = 130/210 (61%), Gaps = 14/210 (6%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAEQ--PEEVRDTSGNLVRNSINYFILPSS-------IQCGTRCE 57
L FLLL +++PL G+ + P++V DTSG +R NY+I+P+ + C
Sbjct: 56 LAFLLLFALSSQPLLGSAEASPDQVIDTSGKKLRADTNYYIIPAKPFTTCGFVSCFNSGG 235
Query: 58 MALLNTNKTCPLDVV--EEEEAMQFSFVPFNFKKGVIRVSTDLNVIHS---FPTNCSTSS 112
+AL ++CPLDVV + + + F P N KKGV+RVSTDLN+ S + + C S
Sbjct: 236 IALETVGESCPLDVVVVKHNQGLPLRFTPVNNKKGVVRVSTDLNIKFSNDAYDSRCPNHS 415
Query: 113 VTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVV 172
+ VWK+D + + FVTT GV GNPG T+ NWF+IE++E YKLV+CP VC C+ V
Sbjct: 416 L-VWKIDPF--SKEETFVTTNGVLGNPGSNTIHNWFQIEKYEDAYKLVYCPNVCPSCKHV 586
Query: 173 CKDIGIFLDENRNTRFVLSDFPFGVKFQRA 202
CKDIGI++ + R R L++ PF VKFQ+A
Sbjct: 587 CKDIGIYVYKYREMRLALTNVPFKVKFQKA 676
>TC82725 weakly similar to PIR|T07871|T07871 miraculin homolog root-knot
nematode-induced - tomato, partial (51%)
Length = 818
Score = 132 bits (332), Expect = 9e-32
Identities = 73/174 (41%), Positives = 103/174 (58%), Gaps = 5/174 (2%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL---LNT 63
L FLLL T++PL + E V D +G +R + Y +L S +R L N
Sbjct: 74 LAFLLLIALTSQPLLSSSL-EHVVDITGKNLRANAYYNVLLSMPYTNSRSPEGLGLSNNI 250
Query: 64 NKTCPLDV--VEEEEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKV 121
+ CPLDV V +++ F P N KKGVIRVS+DLN++ ++C + TVWK+D+
Sbjct: 251 GQPCPLDVIVVSRYQSLPIRFTPLNLKKGVIRVSSDLNIMFRSNSSCPYHT-TVWKLDRF 427
Query: 122 DVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKD 175
D + + FVTT G GNPG +++ NWFKIE++ GYKLV+CP VC C+ CK+
Sbjct: 428 DASKGKSFVTTDGFIGNPGPQSISNWFKIEKYVEGYKLVYCPIVCPSCKHECKN 589
Score = 37.0 bits (84), Expect = 0.005
Identities = 19/43 (44%), Positives = 26/43 (60%)
Frame = +3
Query: 159 LVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQR 201
L+F P V + K +G+F DEN N R LSD P+ VKF++
Sbjct: 552 LLFVPLV----NMSAKMVGLFEDENGNKRLALSDVPYQVKFRK 668
>TC77861 weakly similar to GP|13899081|gb|AAK48962.1 Unknown protein
{Arabidopsis thaliana}, partial (39%)
Length = 871
Score = 129 bits (323), Expect = 1e-30
Identities = 82/211 (38%), Positives = 113/211 (52%), Gaps = 9/211 (4%)
Frame = +3
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK F +L F K + PE V D SG + + Y+ILP + G + +
Sbjct: 51 MKTSFLAFSILCLAFICKTIAA---PEPVLDISGKQLTTGVKYYILP--VIRGKGGGLTV 215
Query: 61 LN---TNKTCPLDVVEEEEAMQ----FSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
N N+TCPL VV+E+ ++ +F P+N K+GVI STDLN I SF T
Sbjct: 216 ANHGENNQTCPLYVVQEKLEVKNGEAVTFTPYNAKQGVILTSTDLN-IKSFVTKTKCPQT 392
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVC 173
VWK+ K T F+ TGGV+GNP TV NWFKIE+ + Y L FCP +C+ +C
Sbjct: 393 QVWKLLKE--LTGVWFLATGGVEGNPSMATVGNWFKIEKADKDYVLSFCPAEACKCQTLC 566
Query: 174 KDIGIFLDENRNTRFVLSDF--PFGVKFQRA 202
+++G+F+D+ N LSD F V F+RA
Sbjct: 567 RELGLFVDDKGNKHLALSDQIPSFRVVFKRA 659
>TC82454 weakly similar to GP|3551243|dbj|BAA32820.1 Water-Soluble
Chlorophyll Protein {Brassica oleracea}, partial (17%)
Length = 940
Score = 80.5 bits (197), Expect = 4e-16
Identities = 68/202 (33%), Positives = 96/202 (46%), Gaps = 9/202 (4%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
L FLL L T PL + E+V+D +GN + S YFI P+ G LN +
Sbjct: 62 LPFLLFALTTYFPLPFTQGVEQVKDKNGNPILVSKKYFIWPADGSGGG----LRLNETEQ 229
Query: 67 CPLDV----VEEEEAMQFSFVPF-NFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKV 121
CPL V E+ +++ F+P N + T L+++ T C+ SS W V V
Sbjct: 230 CPLVVQQAFSEDVKSLPLKFIPTENINDFIFTGYTSLDIVFEKKTKCAESS--KWVV--V 397
Query: 122 DVATSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDIGIF 179
+ ++ GG G G+ +D FKIE S GYKLVFCPT+ + C +IG F
Sbjct: 398 KGGFMEPWIGIGG--GVNGKSVIDGLFKIETIRSFRGYKLVFCPTI-SDPTGQCNNIGRF 568
Query: 180 LDENRNTRFVLSD--FPFGVKF 199
D R ++S+ PF V F
Sbjct: 569 FDNENGLRLIMSENFKPFEVVF 634
>TC76747 similar to GP|20269067|emb|CAD29731. protease inhibitor {Sesbania
rostrata}, partial (86%)
Length = 1189
Score = 75.1 bits (183), Expect = 2e-14
Identities = 68/205 (33%), Positives = 102/205 (49%), Gaps = 13/205 (6%)
Frame = +3
Query: 11 LLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT-NKTCPL 69
LL LF TKPLQGAE PE V D GN ++ Y++ P G + L +T NKTCPL
Sbjct: 135 LLVLFNTKPLQGAE-PEAVVDKQGNPLKPGEGYYVFPLWADNG---GITLGHTRNKTCPL 302
Query: 70 DVVEEEEA----MQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVAT 125
DV+ +A + FS ++ + ++ ++ ++ S P N VW++ K V +
Sbjct: 303 DVIRNPDAIGTPVYFSASGLDYIPTLTDLTIEIPILGS-PCN----EPKVWRLLK--VGS 461
Query: 126 SQRFVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCPTVCRECEVVCKDIGIFL 180
FV+TGG G+ + + FKIER Y FCP+V V+C +G F+
Sbjct: 462 GFWFVSTGGAAGD-----LVSKFKIERLAGEHAYEIYSFKFCPSV---PGVLCAPVGTFV 617
Query: 181 DENRNTRFVLSD---FPFGVKFQRA 202
D + + D P+ V+FQ+A
Sbjct: 618 DTDGTKVMAVGDGIEEPYYVRFQKA 692
>TC76750 similar to GP|20269067|emb|CAD29731. protease inhibitor {Sesbania
rostrata}, partial (77%)
Length = 651
Score = 74.7 bits (182), Expect = 2e-14
Identities = 61/175 (34%), Positives = 81/175 (45%), Gaps = 11/175 (6%)
Frame = +3
Query: 11 LLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNT-NKTCPL 69
L LF TK LQG ++PE V D GN ++ Y++ P G + L T NKTCPL
Sbjct: 30 LSALFNTKALQG-DKPEAVVDKQGNPLKPGEGYYVFPLWADNGG---ITLGQTRNKTCPL 197
Query: 70 DVVEEEEAMQFSFVPFNFKKGVIRVSTDLNV-IHSFPTNCSTSSVTVWKVDKVDVATSQR 128
DV+ EA+ + ++ I TDL V I + CS VWK+ K
Sbjct: 198 DVIRNPEAIGSPVYFYEYEHDYIPTLTDLTVEIPILGSPCSERK--VWKISKEGTRARFW 371
Query: 129 FVTTGGVQGNPGRETVDNWFKIERFESG-----YKLVFCP----TVCRECEVVCK 174
FV+TGG GN + + FKIER E Y ++CP T+C CK
Sbjct: 372 FVSTGGFPGN-----LFSQFKIERLEGEHAYEIYSFLYCPSVPGTLCAPVGTFCK 521
>TC91524 similar to GP|15217238|gb|AAK92582.1 Hypothetical protein {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (5%)
Length = 824
Score = 69.7 bits (169), Expect = 7e-13
Identities = 58/181 (32%), Positives = 86/181 (47%), Gaps = 8/181 (4%)
Frame = +3
Query: 29 VRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFSFV---PF 85
V DTSG V N +YFI P+ G + T +CP +V + +A Q V PF
Sbjct: 102 VIDTSGEPVENDEDYFIRPAITGNGGSLTLV---TRNSCPFNVGLDPDAPQGFAVLLSPF 272
Query: 86 --NFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRET 143
N ++ +R+ DL VI T+C S T W++ + D T +RF+ TG G +
Sbjct: 273 VSNREEDEVRLGRDLRVIFQAGTSCGQS--TEWRLGERDATTGRRFIITGRDDSTVG--S 440
Query: 144 VDNWFKIERFESG--YKLVFCPT-VCRECEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQ 200
N+F+I + S + + +CPT VC C+ C +GI + EN L V FQ
Sbjct: 441 YGNFFRIVQTPSRGIFNIQWCPTEVCPSCKFECGTVGI-VRENGKILLALDGSALPVAFQ 617
Query: 201 R 201
+
Sbjct: 618 K 620
>TC78632 similar to GP|9367042|gb|AAF87095.1| trypsin inhibitor {Glycine
max}, partial (44%)
Length = 803
Score = 69.7 bits (169), Expect = 7e-13
Identities = 59/199 (29%), Positives = 93/199 (46%), Gaps = 5/199 (2%)
Frame = +2
Query: 7 LLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
LLF L T F PL EQ VRD++GN + S +++ PS N +
Sbjct: 41 LLFALATYF---PLAFTEQ---VRDSNGNPIFFSSRFYVKPSIFGAAGGGVKLGETGNSS 202
Query: 67 CPLDVVEEEEAMQFSFVPFNFKKG----VIRVSTDLNVIHS-FPTNCSTSSVTVWKVDKV 121
CPL V+++ + + +P F + +STD + + FP + + W + +
Sbjct: 203 CPLTVLQDYSEV-VNGLPVKFSTDAEIFIDLISTDTSRVDIVFPEKPECAESSKWLLIED 379
Query: 122 DVATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLD 181
D + +V GG++ G+ +D FKI + GYKLVFCPT +C DIG + D
Sbjct: 380 DFP--RPWVGIGGIEDYIGKHIIDGKFKIVKHGFGYKLVFCPTFTAP-PGLCHDIGRYDD 550
Query: 182 ENRNTRFVLSDFPFGVKFQ 200
+N + D P+ V F+
Sbjct: 551 KNGRRLILTEDDPYEVVFE 607
>TC86857 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {Cicer
arietinum}, partial (41%)
Length = 860
Score = 67.8 bits (164), Expect = 3e-12
Identities = 55/197 (27%), Positives = 89/197 (44%), Gaps = 4/197 (2%)
Frame = +1
Query: 7 LLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT 66
+LF L F+ L A+ E+V DT+GN + ++I+PS N
Sbjct: 109 ILFALTICFS---LAFAQVSEQVFDTNGNPIFPGGTFYIMPSIFGAAGGGLRLGKTKNSK 279
Query: 67 CPLDVVEEEE----AMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVD 122
CPL V+++ + F +I +T L++ + +C+ SS V VD +
Sbjct: 280 CPLTVLQDYSEVVNGLPVKFTRLEAGHDIISTNTALDIAFTTKPDCAESSKWVL-VDDFN 456
Query: 123 VATSQRFVTTGGVQGNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDE 182
T +V GG + N G+ ++ +FKI++ GYKLVFCP + C DIG D
Sbjct: 457 KLTGP-WVGIGGTEDNEGKHIINGFFKIQKHGFGYKLVFCPDITAP-PGACYDIGRHDDF 630
Query: 183 NRNTRFVLSDFPFGVKF 199
+ ++ P+ V F
Sbjct: 631 TGRLLVLANNDPYEVVF 681
>TC86085
Length = 884
Score = 65.1 bits (157), Expect = 2e-11
Identities = 49/166 (29%), Positives = 77/166 (45%), Gaps = 9/166 (5%)
Frame = +3
Query: 29 VRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCP----LDVVEEEEAMQFSFVP 84
V DTSG V + YFI P+ G L+ N CP LD E + F P
Sbjct: 138 VIDTSGEPVEDDEEYFIRPAI--TGNGGGSTLVTGNGPCPLHVGLDNTEGTLGVAVKFTP 311
Query: 85 F--NFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRE 142
F +R++ DL V T+C S T W++ + D + +R + TG + G
Sbjct: 312 FAPQHDDDDVRLNRDLRVTFLTSTSCGQS--TDWRLGEKDATSGRRLIVTG---RDNGAG 476
Query: 143 TVDNWFKIERFESG--YKLVFCPT-VCRECEVVCKDIGIFLDENRN 185
+ N+F+I + ++G Y + +CPT C C+V C +G+ + +N
Sbjct: 477 SQGNFFRIVQTQTGGTYNIQWCPTEACPSCKVQCGTVGVIRENGKN 614
>TC86086 similar to GP|15155442|gb|AAK86324.1 AGR_C_899p {Agrobacterium
tumefaciens str. C58 (Cereon)}, partial (2%)
Length = 1329
Score = 61.2 bits (147), Expect = 3e-10
Identities = 54/184 (29%), Positives = 82/184 (44%), Gaps = 11/184 (5%)
Frame = -3
Query: 29 VRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDV----VEEEEAMQFSFVP 84
V DTSG V + YFI P+ G L+ N CPL V E + F P
Sbjct: 1186 VIDTSGEPVEDDEEYFIRPAI--TGNGGGSILVTRNGPCPLHVGLGNSEGTLGLAVKFTP 1013
Query: 85 F----NFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPG 140
F + +R++ DL V T C S T W++ + D + +R + TG + G
Sbjct: 1012 FAPRHDDDDDDVRLNRDLRVTFQGFTGCGQS--TDWRLGEKDATSGRRLIVTG---RDNG 848
Query: 141 RETVDNWFKIERFESG--YKLVFCPT-VCRECEVVCKDIGIFLDENRNTRFVLSDFPFGV 197
+ N+F+I + ++G Y + +CPT C C+V C +G+ + EN L V
Sbjct: 847 AGSHGNFFRIVQTQTGGIYNIQWCPTEACPSCKVQCGTVGV-IRENGKILLALDGGALPV 671
Query: 198 KFQR 201
FQ+
Sbjct: 670 VFQK 659
>TC88358 proteinase inhibitor 20 [Medicago truncatula]
Length = 806
Score = 57.0 bits (136), Expect = 5e-09
Identities = 50/196 (25%), Positives = 82/196 (41%), Gaps = 6/196 (3%)
Frame = +2
Query: 3 LHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLN 62
L L F + T L + E+V D +GN + Y+ILP+ G
Sbjct: 50 LSLTLSFFIFVFITNPSLATSNDVEQVLDINGNPIFPGGQYYILPALRGPGGGGVRLGRT 229
Query: 63 TNKTCPLDVVEE----EEAMQFSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKV 118
+ CP+ V+++ + + F G+I T L + ++ +C+ S T W +
Sbjct: 230 GDLKCPVTVLQDRREVKNGLPVKFTIPGISPGIIFTGTPLEIEYTKKPSCAAS--TKWLI 403
Query: 119 DKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEVVCKDI 176
VD + + GG + PG +T+ F I++ S GY L FC T C DI
Sbjct: 404 -FVDNVIGKACIGIGGPENYPGVQTLKGKFNIQKHASGFGYNLGFCVT----GSPTCLDI 568
Query: 177 GIFLDENRNTRFVLSD 192
G F ++ R L++
Sbjct: 569 GRFDNDEAGRRLNLTE 616
>TC80652 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {Cicer
arietinum}, partial (81%)
Length = 788
Score = 51.2 bits (121), Expect = 3e-07
Identities = 54/186 (29%), Positives = 81/186 (43%), Gaps = 11/186 (5%)
Frame = +2
Query: 3 LHFPLLFLLLTLFTTKPLQGAEQP-EEVRDTSGNLVRNSINYFILPS---SIQCGTRCEM 58
L L FLL T L + E+V D +GN + Y+ILP+ + G R
Sbjct: 11 LSLTLSFLLFVFTTNLSLAFSNDAVEQVLDINGNPIFPGGKYYILPAIRGPLGGGLRLGK 190
Query: 59 ALLNTNKTCPLDVVEEEEAMQFSFVPFNF-----KKGVIRVSTDLNVIHSFPTNCSTSSV 113
++N C + VV++ + + VP F G+I T +++ + NC SS
Sbjct: 191 ---SSNSDCEVTVVQDYNEV-INGVPVKFSIPEISPGIIFTGTPIDIEFTKKPNCVESSK 358
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES--GYKLVFCPTVCRECEV 171
+ VD V + V GG + PG T+ F IE+ ES GY+L +C +
Sbjct: 359 WLIFVDSV---IQKACVGIGGPENYPGFRTLSGTFNIEKHESGFGYRLGYCV----KDSP 517
Query: 172 VCKDIG 177
C DIG
Sbjct: 518 TCLDIG 535
>TC85685 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {Cicer
arietinum}, partial (42%)
Length = 870
Score = 45.8 bits (107), Expect = 1e-05
Identities = 58/196 (29%), Positives = 82/196 (41%), Gaps = 6/196 (3%)
Frame = +1
Query: 10 LLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPL 69
LLL FT PL E V D +GN V Y+I P + G + CP+
Sbjct: 82 LLLFSFTYFPLAFTET---VEDINGNPVFPGGKYYIAPLISKGGGGGLKLGKTGDSECPV 252
Query: 70 DVVEE-EEAMQFSFVPFNF--KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATS 126
V+++ E ++ V F K+GVI + ++++ C+ S+ W + D TS
Sbjct: 253 TVLQDFSEVVRGLPVRFTIIVKRGVIFTTDEVDIEFVKKPKCAESAK--WVLAHDDFPTS 426
Query: 127 QRFVTTGGVQGNPGRETVDNWFKIERFESG---YKLVFCPTVCRECEVVCKDIGIFLDEN 183
G+ N + FKIE SG YKLV+CP C DIG + DEN
Sbjct: 427 WV-----GIGDNI--DAFQGKFKIETLGSGSGAYKLVYCPLFSAP-PGACSDIGRYRDEN 582
Query: 184 RNTRFVLSDFPFGVKF 199
+ PF V F
Sbjct: 583 GWRLVPTENDPFRVVF 630
>CB894496 weakly similar to GP|9367042|gb|A trypsin inhibitor {Glycine max},
partial (29%)
Length = 799
Score = 43.9 bits (102), Expect = 4e-05
Identities = 42/164 (25%), Positives = 65/164 (39%), Gaps = 7/164 (4%)
Frame = +3
Query: 6 PLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNK 65
P F L +F++ P + V+D GN V S YFI P + G + N
Sbjct: 6 PTYFPLPFVFSSFPSSPTDADLIVKDIHGNPVVPSGRYFIWPEFLVSGGGFRLGETE-NS 182
Query: 66 TCPLDVVEEEE----AMQFSFVPFNFKKG--VIRVSTDLNVIHSFPTNCSTSSVTVWKVD 119
TCP V+++ + P N I L++ + +C+ SS W +
Sbjct: 183 TCPFTVLQDYSNLGHGLPVKLTPQNQPSSDDPITRGLRLDIAFDYKPDCAESSK--WLM- 353
Query: 120 KVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFE-SGYKLVFC 162
V F T G+ D WF++ R++ SGY + FC
Sbjct: 354 ---VEAENEFPTPWLAIDGTGKNVHDGWFELTRYKRSGYLIFFC 476
>AJ502095
Length = 578
Score = 32.3 bits (72), Expect(2) = 7e-04
Identities = 20/59 (33%), Positives = 32/59 (53%)
Frame = +3
Query: 92 IRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKI 150
+R+S DL V +NC SS W++ + D + +R +TTG G G + N+F+I
Sbjct: 378 VRLSRDLRV-QFLASNCGQSSE--WRLGERDATSGRRLITTGRDDGTVG--SFGNFFRI 539
Score = 26.6 bits (57), Expect(2) = 7e-04
Identities = 18/49 (36%), Positives = 23/49 (46%)
Frame = +1
Query: 29 VRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEA 77
V DT+G V YFI P G L+ N +CPLDV E+ +
Sbjct: 181 VIDTNGEPVEAGDEYFIRPIITGNGGT---TLVTRNGSCPLDVGLEQNS 318
>AJ500060
Length = 345
Score = 31.6 bits (70), Expect = 0.22
Identities = 22/62 (35%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Frame = +2
Query: 29 VRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDV----VEEEEAMQFSFVP 84
V DTSG V + YFI P+ G L+ N TCPL+V E +F+P
Sbjct: 164 VIDTSGEPVEDDEEYFIRPAI--TGNGGGFTLITGNGTCPLNVGLDNTEGTLGAPVTFIP 337
Query: 85 FN 86
F+
Sbjct: 338 FS 343
>TC82315 similar to GP|4038594|emb|CAA10993.1 tDET1 protein {Lycopersicon
esculentum}, partial (34%)
Length = 680
Score = 29.6 bits (65), Expect = 0.83
Identities = 12/43 (27%), Positives = 24/43 (54%)
Frame = +3
Query: 149 KIERFESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLS 191
K RF+S + ++C + E++CKD ++++ N+ F S
Sbjct: 387 KARRFDSFFTQLYCVPLASCNELICKDFFLYMESNQFGLFATS 515
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,475,177
Number of Sequences: 36976
Number of extensions: 107742
Number of successful extensions: 780
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 759
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 763
length of query: 204
length of database: 9,014,727
effective HSP length: 92
effective length of query: 112
effective length of database: 5,612,935
effective search space: 628648720
effective search space used: 628648720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC124970.4


