

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124967.9 - phase: 0
(122 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
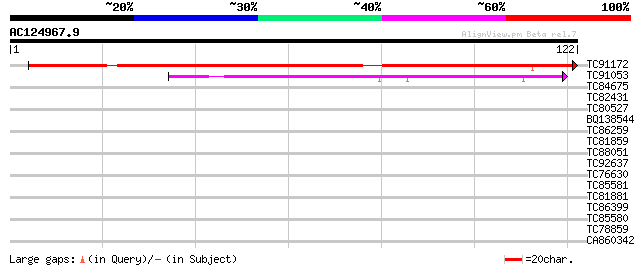
Score E
Sequences producing significant alignments: (bits) Value
TC91172 similar to GP|21618190|gb|AAM67240.1 unknown {Arabidopsi... 110 1e-25
TC91053 weakly similar to PIR|G84766|G84766 hypothetical protein... 39 5e-04
TC84675 similar to GP|21553798|gb|AAM62891.1 unknown {Arabidopsi... 33 0.021
TC82431 weakly similar to GP|8777399|dbj|BAA96989.1 emb|CAB86483... 30 0.17
TC80527 similar to GP|11994425|dbj|BAB02427. auxin-regulated pro... 30 0.30
BQ138544 similar to GP|8777399|dbj| emb|CAB86483.1~gene_id:MFB16... 29 0.51
TC86259 PIR|A25642|HSZM4 histone H4 - maize, complete 28 1.1
TC81859 similar to GP|18491135|gb|AAL69536.1 At1g04290/F19P19_27... 26 3.3
TC88051 similar to GP|15982866|gb|AAL09780.1 At1g27210/T7N9_27 {... 26 4.3
TC92637 similar to PIR|T39903|T39903 serine-rich protein - fissi... 25 5.6
TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule m... 25 5.6
TC85581 weakly similar to PIR|T14329|T14329 dermal glycoprotein ... 25 9.6
TC81881 similar to GP|20161745|dbj|BAB90661. hypothetical protei... 25 9.6
TC86399 similar to PIR|T00710|T00710 thioredoxin homolog F22O13.... 25 9.6
TC85580 similar to GP|13272443|gb|AAK17160.1 unknown protein {Ar... 25 9.6
TC78859 homologue to PIR|T39903|T39903 serine-rich protein - fis... 25 9.6
CA860342 GP|12311751|emb putative vacuolar polyphosphatase {Schi... 25 9.6
>TC91172 similar to GP|21618190|gb|AAM67240.1 unknown {Arabidopsis
thaliana}, partial (55%)
Length = 577
Score = 110 bits (276), Expect = 1e-25
Identities = 62/120 (51%), Positives = 81/120 (66%), Gaps = 2/120 (1%)
Frame = +1
Query: 5 GKLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVV 64
GKL KLKSA+K+W S K SR++ S + +LH V+VGKSRR Y V SDV+
Sbjct: 46 GKLTKLKSAIKRWPSLT--KLSRNNSSVSSSTKQHEHEQELHAVYVGKSRRQYLVNSDVI 219
Query: 65 DNPVFRELVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQPE--SLDELVDFYAC 122
+PVF+ELV+RS +D V VV+CEVVLFEHLLWMLE+ + + + S+ ELV+FY C
Sbjct: 220 QHPVFQELVDRS----SCDDGVVVVSCEVVLFEHLLWMLESVEGETQLGSMAELVEFYNC 387
>TC91053 weakly similar to PIR|G84766|G84766 hypothetical protein At2g35290
[imported] - Arabidopsis thaliana, partial (47%)
Length = 698
Score = 38.9 bits (89), Expect = 5e-04
Identities = 29/100 (29%), Positives = 51/100 (51%), Gaps = 14/100 (14%)
Frame = +2
Query: 35 DEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRE---TEQQND------- 84
D+ S SR TV VGK ++++ V ++ F+ L++ S + T+++ D
Sbjct: 233 DQQDSDSRV---TVVVGKEKKVFLVDPIILQENPFKVLMDISMKKDPTKKKKDHFHFSSS 403
Query: 85 --TVNVVACEVVLFEHLLWMLENADPQ--PESLDELVDFY 120
V V + +LFEH+LW++ N +L ++VDFY
Sbjct: 404 QQRVIFVDVDDILFEHMLWLMNNDTSSLFQLNLKDIVDFY 523
>TC84675 similar to GP|21553798|gb|AAM62891.1 unknown {Arabidopsis
thaliana}, partial (46%)
Length = 599
Score = 33.5 bits (75), Expect = 0.021
Identities = 23/90 (25%), Positives = 45/90 (49%), Gaps = 8/90 (8%)
Frame = +1
Query: 24 KQSRHSISAVADEDSSSSRSDLH-------TVFVGKSRRLYRVTSDVVDNPVFRELVERS 76
K+ +S +DEDS +S H V+VG R + + + + + +F+ L+E++
Sbjct: 79 KRLNSLMSFDSDEDSCNSPKAPHDVPKGYLAVYVGPELRRFIIPTSYLSHSLFKMLLEKA 258
Query: 77 RETEQQNDTVNV-VACEVVLFEHLLWMLEN 105
+ N + + CE+ F++LL +EN
Sbjct: 259 ADEFGFNQCGGLTIPCEIETFKYLLSCMEN 348
>TC82431 weakly similar to GP|8777399|dbj|BAA96989.1
emb|CAB86483.1~gene_id:MFB16.16~similar to unknown
protein {Arabidopsis thaliana}, partial (33%)
Length = 717
Score = 30.4 bits (67), Expect = 0.17
Identities = 18/61 (29%), Positives = 34/61 (55%), Gaps = 1/61 (1%)
Frame = +1
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRELVER-SRETEQQNDTVNVVACEVVLFEHLLWMLENA 106
V+VG R+ + + + ++P+F+ L+E E +ND + C+V LF L +E+A
Sbjct: 283 VYVGPERQRFVIKIKIFNHPLFKTLLEDVENEYGYRNDGPLWLPCDVDLFCEALVEIESA 462
Query: 107 D 107
+
Sbjct: 463 E 465
>TC80527 similar to GP|11994425|dbj|BAB02427. auxin-regulated protein-like
{Arabidopsis thaliana}, partial (62%)
Length = 632
Score = 29.6 bits (65), Expect = 0.30
Identities = 16/58 (27%), Positives = 35/58 (59%), Gaps = 1/58 (1%)
Frame = +1
Query: 48 VFVGKSRRLYRVTSDVVDNPVFRELV-ERSRETEQQNDTVNVVACEVVLFEHLLWMLE 104
++VG + V ++++++PVF +L+ E ++E + V + C V++FE +L L+
Sbjct: 280 IYVGDEMERFVVCAELLNHPVFIKLLNESAQEYGYEQKGVLRLPCHVLVFERVLEALK 453
>BQ138544 similar to GP|8777399|dbj| emb|CAB86483.1~gene_id:MFB16.16~similar
to unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 395
Score = 28.9 bits (63), Expect = 0.51
Identities = 27/86 (31%), Positives = 40/86 (46%), Gaps = 13/86 (15%)
Frame = +2
Query: 5 GKLMKLKSALKKWNSFG-NGKQSRHSISAVADEDSSSSRSD------------LHTVFVG 51
GK K K LK W S G G S+ S + ++ SS S S+ TV+VG
Sbjct: 83 GKCKKNK-ILKAWRSLGRGGDNSKLLRSLLLNKSSSKSFSENAKGRIVKIPNGCFTVYVG 259
Query: 52 KSRRLYRVTSDVVDNPVFRELVERSR 77
+ + V + V++P F+ L+ SR
Sbjct: 260 LQSQRFVVKTKFVNHPKFKMLLG*SR 337
>TC86259 PIR|A25642|HSZM4 histone H4 - maize, complete
Length = 597
Score = 27.7 bits (60), Expect = 1.1
Identities = 15/29 (51%), Positives = 18/29 (61%)
Frame = +1
Query: 71 ELVERSRETEQQNDTVNVVACEVVLFEHL 99
E VER E E+QNDTV C V+ F+ L
Sbjct: 67 EKVERDWEREEQNDTVR---CFVITFKEL 144
>TC81859 similar to GP|18491135|gb|AAL69536.1 At1g04290/F19P19_27
{Arabidopsis thaliana}, partial (65%)
Length = 960
Score = 26.2 bits (56), Expect = 3.3
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +1
Query: 72 LVERSRETEQQNDTVNVVACEVVLFEHLLWMLENADPQPESLD 114
+V S+ T ++ V C VVL LLW + A+PQ LD
Sbjct: 148 VVSSSQ*TYPHVYSIQVSTCTVVL--SLLWWISQAEPQFPQLD 270
>TC88051 similar to GP|15982866|gb|AAL09780.1 At1g27210/T7N9_27 {Arabidopsis
thaliana}, partial (16%)
Length = 1597
Score = 25.8 bits (55), Expect = 4.3
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +1
Query: 16 KWNSFGNGKQSRHSISAVADEDSS 39
KWNSFGN + IS +E S+
Sbjct: 382 KWNSFGNKQSG*KQISKARNESST 453
>TC92637 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (7%)
Length = 671
Score = 25.4 bits (54), Expect = 5.6
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = -2
Query: 17 WNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYR 58
W S + + RH S + + SSSS+S T F+ RRL+R
Sbjct: 436 WASSASFHRHRHRFSPLPPQLSSSSKSS--TCFLILPRRLHR 317
>TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule membrane
protein {Medicago sativa}, complete
Length = 2605
Score = 25.4 bits (54), Expect = 5.6
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +2
Query: 77 RETEQQNDTVNVVACEVVLFEH 98
R E + D + ACE++LF H
Sbjct: 1502 RAVESKRDNGQLTACEILLFHH 1567
>TC85581 weakly similar to PIR|T14329|T14329 dermal glycoprotein precursor
extracellular - carrot (fragment), partial (72%)
Length = 1570
Score = 24.6 bits (52), Expect = 9.6
Identities = 10/22 (45%), Positives = 16/22 (72%)
Frame = +2
Query: 94 VLFEHLLWMLENADPQPESLDE 115
++F +L+W+L N D P SLD+
Sbjct: 1232 IIFCNLIWLLLNWDLAPCSLDD 1297
>TC81881 similar to GP|20161745|dbj|BAB90661. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (14%)
Length = 651
Score = 24.6 bits (52), Expect = 9.6
Identities = 15/62 (24%), Positives = 30/62 (48%)
Frame = +3
Query: 15 KKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVE 74
K W G+++R VA E S V+VG + + + ++ ++P+F+ L+E
Sbjct: 492 KSWPGLPRGEENRRK--KVAPEGCFS-------VYVGPQMQRFVIKTEYANHPLFKMLLE 644
Query: 75 RS 76
+
Sbjct: 645 EA 650
>TC86399 similar to PIR|T00710|T00710 thioredoxin homolog F22O13.5 -
Arabidopsis thaliana, partial (61%)
Length = 1568
Score = 24.6 bits (52), Expect = 9.6
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = -2
Query: 4 GGKLMKLKSALKKWNSFGNGKQSRHS 29
GG+ +KL+ L++W +F G ++ S
Sbjct: 322 GGRSVKLRFQLQRWKTFHTGVKATSS 245
>TC85580 similar to GP|13272443|gb|AAK17160.1 unknown protein {Arabidopsis
thaliana}, partial (87%)
Length = 1548
Score = 24.6 bits (52), Expect = 9.6
Identities = 10/22 (45%), Positives = 16/22 (72%)
Frame = +1
Query: 94 VLFEHLLWMLENADPQPESLDE 115
++F +L+W+L N D P SLD+
Sbjct: 1228 IIFCNLIWLLLNWDLAPCSLDD 1293
>TC78859 homologue to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (3%)
Length = 1204
Score = 24.6 bits (52), Expect = 9.6
Identities = 16/80 (20%), Positives = 32/80 (40%)
Frame = -2
Query: 6 KLMKLKSALKKWNSFGNGKQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVD 65
K + S K++S RHS E+ + D + V + + T + +
Sbjct: 888 KKFLMSSISDKFHSVWGSHSHRHSKEDGLIEEGGGNCDDHNKALVVYNNNAHHRTYEALQ 709
Query: 66 NPVFRELVERSRETEQQNDT 85
NP ++V+ ++ E D+
Sbjct: 708 NPSETQVVDEQQDQEDGYDS 649
>CA860342 GP|12311751|emb putative vacuolar polyphosphatase
{Schizosaccharomyces pombe}, partial (1%)
Length = 245
Score = 24.6 bits (52), Expect = 9.6
Identities = 17/61 (27%), Positives = 26/61 (41%)
Frame = +1
Query: 24 KQSRHSISAVADEDSSSSRSDLHTVFVGKSRRLYRVTSDVVDNPVFRELVERSRETEQQN 83
K+ + + V ED R+D+ F +S D P LVE + ETE+ N
Sbjct: 67 KKKKKKKNKVVFEDEVEQRADIEFQFEDQS-----------DYPAPVPLVEENNETEENN 213
Query: 84 D 84
+
Sbjct: 214 E 216
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.130 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,536,572
Number of Sequences: 36976
Number of extensions: 37950
Number of successful extensions: 206
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 204
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 204
length of query: 122
length of database: 9,014,727
effective HSP length: 84
effective length of query: 38
effective length of database: 5,908,743
effective search space: 224532234
effective search space used: 224532234
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC124967.9


