

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124967.1 - phase: 0 /pseudo
(462 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
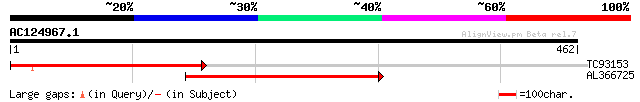
Score E
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 147 1e-35
AL366725 104 8e-23
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 147 bits (370), Expect = 1e-35
Identities = 87/162 (53%), Positives = 103/162 (62%), Gaps = 2/162 (1%)
Frame = +1
Query: 1 CCCT*GCSSSCSTATAS--W*WRSEDAGDLFEKSSPSIQRKV*SRWCPVMVEGSGKDFQS 58
CCC *G SSSCSTAT W* ++DA D E+SS +IQ V S WC + EG K+ S
Sbjct: 28 CCCA*GGSSSCSTAT*G*YW**WNQDARDFLEESSTNIQG*VCS*WCLEVAEGD*KNIPS 207
Query: 59 HAVL*SAKGAVRDAHVG*GGRRLVGKFVTSVGTEWSCSDLGSVQKGISE*VFPGRCSR*E 118
HA+ * +GAV DAHV * GR LV FV G C +LG VQ+G+S VF C *E
Sbjct: 208 HAMF*DTEGAVWDAHVS*RGR*LVD*FVACSGAG*CCGNLGHVQEGVSGQVFSRGCQG*E 387
Query: 119 GD*IPGVEAG*YVCC*VCCKICGACKVLSPLCPGDS*ILQVH 160
D*I GVE G*+VC VCCK+CG C +L L GDS*+LQV+
Sbjct: 388 RD*ISGVETG*HVCHRVCCKVCGTCHILPSLQCGDS*VLQVY 513
>AL366725
Length = 485
Score = 104 bits (259), Expect = 8e-23
Identities = 76/161 (47%), Positives = 98/161 (60%)
Frame = +1
Query: 144 KVLSPLCPGDS*ILQVHKV*EWLES*HQESHRIPAD*NFL*FGESL*DLRGGHQSSL*SD 203
KVLS LC D *+L++H+V EW E+ HQE +RIP +F *F E L +LRGG+ S
Sbjct: 1 KVLSALCC*DC*VLEMHQVREWFEARHQEGNRIPTAPSFP*FSEYLQNLRGGY*GS*QGC 180
Query: 204 E*AKR*GTSESS*TV*CPD**G*AET***EEAE*EGCSC*DSVL*VWRERSQE*RLYQG* 263
E*A+ G ES * +*CP **G E * + EGCSC D + +WRER QE* L
Sbjct: 181 E*AEDQGPIESP*AL*CPC**GQTENGR**AS*EEGCSCGDCLFQLWRERPQE*CLS*RD 360
Query: 264 EEVFQMWTEGSHASRLQAR*CCVL*L**RGPHQYSVHTAEE 304
+E+ + EGS+ S LQA+* CVL L RG + + + A E
Sbjct: 361 QEMCPV*QEGSYCS*LQAK*YCVLQLQRRGSYWFPMQAA*E 483
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.376 0.169 0.742
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,950,217
Number of Sequences: 36976
Number of extensions: 290445
Number of successful extensions: 5845
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 1777
Number of HSP's successfully gapped in prelim test: 188
Number of HSP's that attempted gapping in prelim test: 3768
Number of HSP's gapped (non-prelim): 2338
length of query: 462
length of database: 9,014,727
effective HSP length: 99
effective length of query: 363
effective length of database: 5,354,103
effective search space: 1943539389
effective search space used: 1943539389
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124967.1


