

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.3 + phase: 0
(470 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
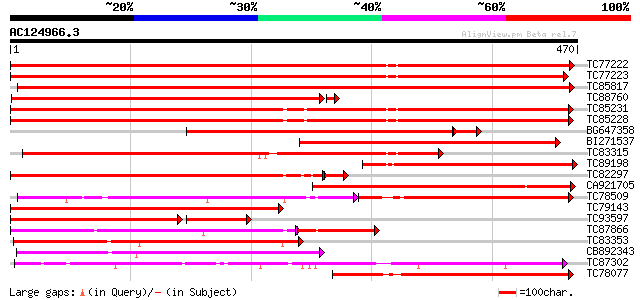
Score E
Sequences producing significant alignments: (bits) Value
TC77222 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 527 e-150
TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 524 e-149
TC85817 weakly similar to PIR|B85014|B85014 probable flavonol gl... 514 e-146
TC88760 similar to GP|19911209|dbj|BAB86931. glucosyltransferase... 477 e-136
TC85231 similar to GP|13508844|emb|CAC35167. arbutin synthase {R... 438 e-123
TC85228 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 431 e-121
BG647358 similar to GP|19911209|db glucosyltransferase-13 {Vigna... 379 e-113
BI271537 similar to GP|19911209|db glucosyltransferase-13 {Vigna... 386 e-108
TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 330 5e-91
TC89198 similar to GP|19911209|dbj|BAB86931. glucosyltransferase... 270 7e-73
TC82297 similar to GP|19911209|dbj|BAB86931. glucosyltransferase... 245 3e-69
CA921705 weakly similar to GP|13508844|em arbutin synthase {Rauv... 256 2e-68
TC78509 similar to GP|19911193|dbj|BAB86923. glucosyltransferase... 157 2e-66
TC79143 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 247 8e-66
TC93597 similar to GP|19911209|dbj|BAB86931. glucosyltransferase... 189 8e-66
TC87866 similar to GP|19911195|dbj|BAB86924. glucosyltransferase... 152 3e-55
TC83353 similar to GP|19911195|dbj|BAB86924. glucosyltransferase... 200 1e-51
CB892343 similar to GP|19911195|dbj glucosyltransferase-6 {Vigna... 198 4e-51
TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase... 181 4e-46
TC78077 similar to GP|4115536|dbj|BAA36411.1 UDP-glycose:flavono... 179 3e-45
>TC77222 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (76%)
Length = 1643
Score = 527 bits (1358), Expect = e-150
Identities = 259/469 (55%), Positives = 344/469 (73%), Gaps = 1/469 (0%)
Frame = +3
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
M KT+HIAV+P G+SHLVPI++FSKRLV HPNFHVTC IP+LGSP +++KS L+ +P
Sbjct: 21 MAKTIHIAVIPSPGFSHLVPIVEFSKRLVTNHPNFHVTCIIPSLGSPPDSSKSYLEKIPP 200
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
NIN FLPP+N DLPQG I T+T SLP +HQ LKSL+ + PLVA++ D+F+ E
Sbjct: 201 NINSIFLPPINKQDLPQGVYPAILIQQTVTLSLPSIHQALKSLSSKAPLVAIIADSFAFE 380
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
L+ KE N LSY YFP++A L++ +LPKLDEE SCE++D+ EPIK+ GCVP++G D
Sbjct: 381 ALDFAKEFNSLSYFYFPTSANVLSFILHLPKLDEEVSCEFKDLQEPIKLQGCVPINGIDL 560
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPI 240
+ +DRSS+AY+ FL K DG+L+NSF E+E + A+K++G N +PVGPI
Sbjct: 561 PTPTKDRSSEAYRMFLQRAKSFYFVDGILINSFYELESSAVEALKQKGYGNISYFPVGPI 740
Query: 241 IETETKSGD-DANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFL 299
+ + + D + ECL WL Q SVLYVSFGSGGTLSQ QI E+A GLELS +F+
Sbjct: 741 TQIGSSNNDVVGDEHECLKWLKNQPQNSVLYVSFGSGGTLSQRQINEIAFGLELSGQRFI 920
Query: 300 WVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGG 359
W +RAPS S ++A YL + N+ D L+FLP GF ERTKEKGF++ SWAPQ++IL +SVGG
Sbjct: 921 WGVRAPSDSVNAA-YLESTNE-DPLKFLPEGFQERTKEKGFILPSWAPQVEILKHSSVGG 1094
Query: 360 FLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVA 419
FL+HCGWNS LES+ GVP++ WPLFAEQ MNAV+L +GLKV +R ++ IVE+ + A
Sbjct: 1095FLSHCGWNSVLESMQEGVPIVAWPLFAEQAMNAVMLCDGLKVAIRLKFEDDEIVEKDKTA 1274
Query: 420 KVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRN 468
VIK LMEG+EG+ +R+ MK LK+ A NAVK++GSS + +SQ+A +W +
Sbjct: 1275NVIKCLMEGEEGKTMRDRMKSLKDYAVNAVKDEGSSIQNLSQLASQWES 1421
>TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (50%)
Length = 1628
Score = 524 bits (1350), Expect = e-149
Identities = 258/464 (55%), Positives = 340/464 (72%), Gaps = 1/464 (0%)
Frame = +3
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
M KT+HIAV+P G+SHLVPI++F+KRLV HPNFHVTC IP+LGSP +++KS L+T+P
Sbjct: 12 MAKTIHIAVIPSPGFSHLVPIVEFTKRLVTNHPNFHVTCIIPSLGSPPDSSKSYLETIPP 191
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
NIN FLPP+N DLPQG I LT+T+SLP +HQ L+SL + PLVA++ D F+ E
Sbjct: 192 NINSIFLPPINKQDLPQGVHPGVLIQLTVTHSLPSIHQALESLTSKTPLVAIIADTFAFE 371
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
L+ KE N LSY+YFP ++ L+ +LPKLDEE SCEY+D+ EPIK+ GCVP++G D
Sbjct: 372 ALDFAKEFNSLSYLYFPCSSFVLSLLLHLPKLDEEFSCEYKDLQEPIKLQGCVPINGIDL 551
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPI 240
+ +DRS++ YK ++ K + DG+L+NSF+E+E + A++ +G +PVGPI
Sbjct: 552 PAATKDRSNEGYKMYIQRAKSMYFVDGILINSFIELESSAIKALELKGYGKIDFFPVGPI 731
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
+T + D + LECL WL Q SVLYVSFGSGGTLSQ QI ELA GLELS +F+W
Sbjct: 732 TQTGLSNNDVGDELECLKWLKNQPQNSVLYVSFGSGGTLSQTQINELAFGLELSGQRFIW 911
Query: 301 VLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
VLRAPS S S+A YL A N+ D L+FLP GFLERTKEKG ++ SWAPQ+QIL SVGGF
Sbjct: 912 VLRAPSDSVSAA-YLEATNE-DPLKFLPKGFLERTKEKGLILPSWAPQVQILKEKSVGGF 1085
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK 420
L+HCGWNS LES+ GVP++ WPLFAEQ MNAV+LS LKV +R ++ IVE+ ++A
Sbjct: 1086LSHCGWNSVLESMQEGVPIVAWPLFAEQAMNAVMLSNDLKVAIRLKFEDDEIVEKDKIAN 1265
Query: 421 VIKYLMEGDEGEKLRNNMKELKEAASNAVK-EDGSSTKTISQIA 463
VIK LMEG+EG+ +R+ MK L++ A+ A+ +DGSS +T+S +A
Sbjct: 1266VIKCLMEGEEGKAMRDRMKSLRDYATKALNVKDGSSIQTLSHLA 1397
>TC85817 weakly similar to PIR|B85014|B85014 probable flavonol
glucosyltransferase [imported] - Arabidopsis thaliana,
partial (83%)
Length = 1736
Score = 514 bits (1324), Expect = e-146
Identities = 244/462 (52%), Positives = 333/462 (71%)
Frame = +3
Query: 7 IAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTF 66
+ ++P G HL+P+++F+KR++ L+ N +T FIPT G PS A K++LQ+LP I+HTF
Sbjct: 162 VVMLPSPGMGHLIPMIEFAKRIIILNQNLQITFFIPTEGPPSKAQKTVLQSLPKFISHTF 341
Query: 67 LPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGK 126
LPPV+ +DLP + +E+ I LT+ SLP L Q +L++ + A+VVD F + ++ +
Sbjct: 342 LPPVSFSDLPPNSGIETIISLTVLRSLPSLRQNFNTLSETHTITAVVVDLFGTDAFDVAR 521
Query: 127 ELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQD 186
E N+ Y+++PS A L+ YLP+LDEE CE+R++ EP+KIPGC+P+HG+ L QD
Sbjct: 522 EFNVPKYVFYPSTAMALSLFLYLPRLDEEVHCEFRELTEPVKIPGCIPIHGKYLLDPLQD 701
Query: 187 RSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETK 246
R + AY+ K ADG++ NSFLE+E GP+ + +E P YPVGP+++ E +
Sbjct: 702 RKNDAYQSVFRNAKRYREADGLIENSFLELEPGPIKELLKEEPGKPKFYPVGPLVKREVE 881
Query: 247 SGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPS 306
G E L WLD Q SVL+VSFGSGGTLS +QIVELALGLE+S +FLWV+R+P+
Sbjct: 882 VGQIGPNSESLKWLDNQPHGSVLFVSFGSGGTLSSKQIVELALGLEMSEQRFLWVVRSPN 1061
Query: 307 SSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGW 366
++A Y SAE D D FLP+GFLERTK +G V++SWAPQ Q+L+ S GGFLTHCGW
Sbjct: 1062DKVANASYFSAETDSDPFDFLPNGFLERTKGRGLVVSSWAPQPQVLAHGSTGGFLTHCGW 1241
Query: 367 NSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLM 426
NS LESVV+GVPL+ WPL+AEQKMNAV+L+E +KVGLR +V ENG+VER+E+A V+K LM
Sbjct: 1242NSVLESVVNGVPLVVWPLYAEQKMNAVMLTEDVKVGLRPNVGENGLVERLEIASVVKCLM 1421
Query: 427 EGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRN 468
EG+EG+KLR MK+LKEAAS + E+G+ST IS +ALKW N
Sbjct: 1422EGEEGKKLRYQMKDLKEAASKTLGENGTSTNHISNLALKWTN 1547
>TC88760 similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13 {Vigna
angularis}, partial (34%)
Length = 862
Score = 477 bits (1228), Expect(2) = e-136
Identities = 231/260 (88%), Positives = 242/260 (92%)
Frame = +3
Query: 2 EKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSN 61
EKTVHIAVVPGVGYSHLVPILQFSKRLVQLHP+FHVTCFIPTLGSPSNATKSILQTLPSN
Sbjct: 48 EKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPDFHVTCFIPTLGSPSNATKSILQTLPSN 227
Query: 62 INHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEV 121
INHTFLPPVNPNDLPQGTTMESQ+ LTL NSLPYLH LKSLA E PLVALVVD+F+VEV
Sbjct: 228 INHTFLPPVNPNDLPQGTTMESQMFLTLNNSLPYLHDALKSLAIESPLVALVVDSFAVEV 407
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
LNIGKELNMLSY+YFP+AATTLAW YLPKLDEETSCEYRDI EPIKIPGCVP+HGRD L
Sbjct: 408 LNIGKELNMLSYVYFPAAATTLAWSIYLPKLDEETSCEYRDIPEPIKIPGCVPIHGRDLL 587
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
S+AQDRSSQ YKHFLP KLLS ADGV VNSFLE+EMGP+SAMKEEG DNPPVYPVGPII
Sbjct: 588 SVAQDRSSQVYKHFLPLFKLLSFADGVFVNSFLELEMGPISAMKEEGSDNPPVYPVGPII 767
Query: 242 ETETKSGDDANGLECLAWLD 261
+TET SGDDAN LECLAWLD
Sbjct: 768 QTETSSGDDANALECLAWLD 827
Score = 27.3 bits (59), Expect(2) = e-136
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = +2
Query: 263 QQPCSVLYVSF 273
QQPCSVLYVSF
Sbjct: 830 QQPCSVLYVSF 862
>TC85231 similar to GP|13508844|emb|CAC35167. arbutin synthase {Rauvolfia
serpentina}, partial (34%)
Length = 1602
Score = 438 bits (1127), Expect = e-123
Identities = 233/471 (49%), Positives = 322/471 (67%), Gaps = 4/471 (0%)
Frame = +3
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
MEK IA+VP G SHL+P+++F+K L+Q H +H+T IPTLG + + +SIL TLP
Sbjct: 48 MEKKTCIAMVPSPGLSHLIPLVEFAKLLLQNHNEYHITFLIPTLGPLTPSMQSILNTLPP 227
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
N+N LP VN DLP +Q+ L + +S+P+L++ +KSL + LVALV FS +
Sbjct: 228 NMNFIVLPQVNIEDLPHNLDPATQMKLIVKHSIPFLYEEVKSLLSKTRLVALVFSMFSTD 407
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEY-RDILEPIKIPG-CVPLHGR 178
++ K N+LSY++F S A + +P LDE S ++ E + IPG +PLH +
Sbjct: 408 AHDVAKHFNLLSYLFFSSGAVLFSLFLTIPNLDEAASTQFLGSSYETVNIPGFSIPLHIK 587
Query: 179 DFLS-IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPV 237
+ +RSS AYK L + LS DGV++N+F ++E + +++ + P VYPV
Sbjct: 588 ELPDPFICERSSDAYKSILDVCQKLSLFDGVIMNTFTDLEPEVIRVLQDR--EKPSVYPV 761
Query: 238 GPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTK 297
GP+I E S ++AN CL WL+ QQP SVL+VSFGSGGTLSQ+Q+ E+A GLELS K
Sbjct: 762 GPMIRNE--SNNEANMSMCLRWLENQQPSSVLFVSFGSGGTLSQDQLNEIAFGLELSGHK 935
Query: 298 FLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
FLWV+RAPS +SSSA Y S +N+ D L++LP+GFLERTKE G V+ SWAPQ++IL S+
Sbjct: 936 FLWVVRAPSKNSSSA-YFSGQNN-DPLEYLPNGFLERTKENGLVVASWAPQVEILGHGSI 1109
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASV-NENGIVERV 416
GGFL+HCGW+STLESVV+GVPLI WPLFAEQ+MNA LL++ LKV +R V +E GI+++
Sbjct: 1110GGFLSHCGWSSTLESVVNGVPLIAWPLFAEQRMNAKLLTDVLKVAVRPKVDDETGIIKQE 1289
Query: 417 EVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
EVAK IK +M+GDE ++R +KEL A+ + E GSS K +S +ALKW+
Sbjct: 1290EVAKAIKRIMKGDESFEIRKKIKELSVGAATVLSEHGSSRKALSSLALKWQ 1442
>TC85228 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (31%)
Length = 3139
Score = 431 bits (1108), Expect = e-121
Identities = 228/471 (48%), Positives = 319/471 (67%), Gaps = 4/471 (0%)
Frame = -1
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
ME IA+VP G SHL+P ++F+K L+Q H H+T IPTLG + + +SIL TLP
Sbjct: 2788 MENKTCIAMVPSPGLSHLIPQVEFAKLLLQHHNESHITFLIPTLGPLTPSMQSILNTLPP 2609
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
N+N T LP VN DLP +Q+ L + +S+P+LH+ +KSL + LVALV FS +
Sbjct: 2608 NMNFTVLPQVNIEDLPHNLEPSTQMKLIVKHSIPFLHEEVKSLLSKTNLVALVCSMFSTD 2429
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEY-RDILEPIKIPG-CVPLHGR 178
++ K N+LSY++F S A ++ LP LD+ S ++ E + +PG +P H +
Sbjct: 2428 AHDVAKHFNLLSYLFFSSGAVLFSFFLTLPNLDDAASTQFLGSSYEMVNVPGFSIPFHVK 2249
Query: 179 DFLS-IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPV 237
+ +RSS YK L + S DGV++N+F +E+ + +++ + P V+PV
Sbjct: 2248 ELPDPFNCERSSDTYKSILDVCQKSSLFDGVIINTFSNLELEAVRVLQDR--EKPSVFPV 2075
Query: 238 GPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTK 297
GPII E S ++AN CL WL+ Q P SV++VSFGSGGTLSQ+Q+ ELA GLELS K
Sbjct: 2074 GPIIRNE--SNNEANMSVCLRWLENQPPSSVIFVSFGSGGTLSQDQLNELAFGLELSGHK 1901
Query: 298 FLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
FLWV+RAPS SSSA Y + +N+ + L++LP+GF+ERTKEKG V+TSWAPQ++IL S+
Sbjct: 1900 FLWVVRAPSKHSSSA-YFNGQNN-EPLEYLPNGFVERTKEKGLVVTSWAPQVEILGHGSI 1727
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVN-ENGIVERV 416
GGFL+HCGW+STLESVV+GVPLI WPLFAEQ+MNA LL++ LKV +R V+ E GI++R
Sbjct: 1726 GGFLSHCGWSSTLESVVNGVPLIAWPLFAEQRMNAKLLTDVLKVAVRPKVDGETGIIKRE 1547
Query: 417 EVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
EV+K +K +MEGDE ++R +KEL +A+ + E GSS K +S +ALKW+
Sbjct: 1546 EVSKALKRIMEGDESFEIRKKIKELSVSAATVLSEHGSSRKALSTLALKWQ 1394
>BG647358 similar to GP|19911209|db glucosyltransferase-13 {Vigna angularis},
partial (33%)
Length = 738
Score = 379 bits (974), Expect(2) = e-113
Identities = 184/225 (81%), Positives = 204/225 (89%)
Frame = +3
Query: 147 FYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVKLLSSAD 206
FYLPKLDEET+CEYRD+ EPIK+PGCVPLHGRD L+I QDRSSQAYK+FL VK LS AD
Sbjct: 3 FYLPKLDEETTCEYRDLPEPIKVPGCVPLHGRDLLTIVQDRSSQAYKYFLQHVKSLSFAD 182
Query: 207 GVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETKSGDDANGLECLAWLDKQQPC 266
GVLVNSFLE+EMGP++A+ EEG NP VYPVGPII+T T S DDANGLECL+WLDKQQ C
Sbjct: 183 GVLVNSFLEMEMGPINALTEEGSGNPSVYPVGPIIQTVTGSVDDANGLECLSWLDKQQSC 362
Query: 267 SVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQF 326
SVLYVSFGSGGTLS EQIVELALGLELSN KFLWV+RAPSSSSS+A YLSA+ND+D LQF
Sbjct: 363 SVLYVSFGSGGTLSHEQIVELALGLELSNQKFLWVVRAPSSSSSNAAYLSAQNDVDALQF 542
Query: 327 LPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLE 371
LPSGFLERTKE+GFVITSWAPQIQILS +SVGGFL+HCGW+S L+
Sbjct: 543 LPSGFLERTKEEGFVITSWAPQIQILSHSSVGGFLSHCGWSSHLK 677
Score = 48.1 bits (113), Expect(2) = e-113
Identities = 21/23 (91%), Positives = 22/23 (95%)
Frame = +2
Query: 369 TLESVVHGVPLITWPLFAEQKMN 391
TLESVVHGVPLITWP+FAEQ MN
Sbjct: 668 TLESVVHGVPLITWPMFAEQGMN 736
>BI271537 similar to GP|19911209|db glucosyltransferase-13 {Vigna angularis},
partial (34%)
Length = 652
Score = 386 bits (992), Expect = e-108
Identities = 197/216 (91%), Positives = 201/216 (92%)
Frame = +2
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
I T SGD +GLECL WLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSN FLW
Sbjct: 2 IPTIESSGDANHGLECLTWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNKIFLW 181
Query: 301 VLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
VLRAPSSSSSSAGY SA+ND DT QFLPSGFLERTKEKGFVITSW PQIQILS NSVGGF
Sbjct: 182 VLRAPSSSSSSAGYFSAQNDADTWQFLPSGFLERTKEKGFVITSWVPQIQILSHNSVGGF 361
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK 420
LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK
Sbjct: 362 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK 541
Query: 421 VIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSST 456
VIK LMEG+EGEKLRNNMKELKE+ASNAVKEDGSST
Sbjct: 542 VIKCLMEGEEGEKLRNNMKELKESASNAVKEDGSST 649
>TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (42%)
Length = 1048
Score = 330 bits (847), Expect = 5e-91
Identities = 179/357 (50%), Positives = 238/357 (66%), Gaps = 8/357 (2%)
Frame = +2
Query: 11 PGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTFLPPV 70
P G+SHLVPI++FSKRLV HPNFHVTC IP+LGSP +++KS L+T+P NIN FLPP+
Sbjct: 2 PSPGFSHLVPIVEFSKRLVTNHPNFHVTCIIPSLGSPPDSSKSYLETIPPNINSIFLPPI 181
Query: 71 NPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNM 130
N DLPQG I T+T SLP +HQ LKSL + PLVA++ D F+ E L+ KE N
Sbjct: 182 NKQDLPQGVYPAILIQQTVTLSLPSIHQALKSLNSKAPLVAIIADIFAQETLDFAKEFNS 361
Query: 131 LSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSSQ 190
L Y+YFPS+A L+ ++P LDEE SCEY+D+ EPIK+ GC+P++G D + +DRS++
Sbjct: 362 LFYLYFPSSAFVLSLVLHIPNLDEEVSCEYKDLKEPIKLQGCLPINGIDLPTPTKDRSNE 541
Query: 191 AYKHFLPFVKLLSSA---DGVLV----NSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIET 243
+ ++ K + G++ N L+ A++++G +PVG I +
Sbjct: 542 SLQNASSTCKKYAFG*WNLGLIAS*N*NQVLQ------KALEQKGYGKIGFFPVGTITQI 703
Query: 244 ETKSGD-DANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVL 302
+ + D + ECL WL Q SVLYVSFGSGGTLSQ QI ELA GLELS +F+WV+
Sbjct: 704 GSSNNDVVGDEHECLKWLKNQPQNSVLYVSFGSGGTLSQTQINELAFGLELSGQRFIWVV 883
Query: 303 RAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGG 359
RAPS S S+A YL + N+ D L+FLP G LERTKEKG + SWAPQ++IL +SVGG
Sbjct: 884 RAPSDSVSAA-YLESTNE-DPLKFLPIGXLERTKEKGXYLASWAPQVEILKHSSVGG 1048
>TC89198 similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13 {Vigna
angularis}, partial (31%)
Length = 828
Score = 270 bits (691), Expect = 7e-73
Identities = 134/178 (75%), Positives = 161/178 (90%)
Frame = +1
Query: 293 LSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQIL 352
LSN KFLWV+R+PS+++++A YLSA +D+D LQFLPSGFLER KE+G VI SWAPQIQIL
Sbjct: 13 LSNHKFLWVVRSPSNTANAA-YLSA-SDVDPLQFLPSGFLERKKEQGMVIPSWAPQIQIL 186
Query: 353 SPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGI 412
+SVGGFLTHCGWNSTLESV+HGVPLITWPLFAEQ+ NAVLLSEGLKVGLR +N+NGI
Sbjct: 187 RHSSVGGFLTHCGWNSTLESVLHGVPLITWPLFAEQRTNAVLLSEGLKVGLRPKINQNGI 366
Query: 413 VERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNLV 470
VE+V++A++IK LMEG+EG KLR NMKELKE+A++A K+DGS TKT+SQ+ALKWRNLV
Sbjct: 367 VEKVQIAELIKCLMEGEEGGKLRKNMKELKESANSAHKDDGSFTKTLSQLALKWRNLV 540
>TC82297 similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13 {Vigna
angularis}, partial (13%)
Length = 875
Score = 245 bits (626), Expect(2) = 3e-69
Identities = 131/263 (49%), Positives = 171/263 (64%)
Frame = +2
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
ME+ HI VVP G++HLVPIL+FSKRLV LHP FH+TC IP++GSP +++KS LQTLP
Sbjct: 44 MEEKPHIVVVPSAGFTHLVPILEFSKRLVNLHPQFHITCIIPSIGSPPSSSKSYLQTLPP 223
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
I+ FLPP+N + +P + QI L++ +SLPY+ Q LKSL +VA+V D F+ +
Sbjct: 224 TISSIFLPPINVDQVPDAKILAVQISLSVKHSLPYIEQELKSLCSRSKVVAVVADVFAHD 403
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
VL+I K+ N+L YIY P AA L+ FY KLDE S E RD EPIK+PGCV +D
Sbjct: 404 VLDIAKDFNLLCYIYLPQAAMVLSTYFYSSKLDEILSDESRDPNEPIKVPGCVAFDLKDL 583
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPI 240
+ RS+ Y FL + DGV VNSFLE E + +KEE P VYPVGPI
Sbjct: 584 PLPFRFRSNIGYTKFLERAEKYHLFDGVFVNSFLEFEEDAIKGLKEE-KKKPMVYPVGPI 760
Query: 241 IETETKSGDDANGLECLAWLDKQ 263
I+ + GD+ N ++CL WL+KQ
Sbjct: 761 IQ-KVSIGDE-NEVKCLTWLEKQ 823
Score = 35.0 bits (79), Expect(2) = 3e-69
Identities = 16/20 (80%), Positives = 18/20 (90%)
Frame = +3
Query: 262 KQQPCSVLYVSFGSGGTLSQ 281
K +P SVL+VSFGSGGTLSQ
Sbjct: 816 KNRPKSVLFVSFGSGGTLSQ 875
>CA921705 weakly similar to GP|13508844|em arbutin synthase {Rauvolfia
serpentina}, partial (34%)
Length = 787
Score = 256 bits (653), Expect = 2e-68
Identities = 120/219 (54%), Positives = 173/219 (78%), Gaps = 1/219 (0%)
Frame = -2
Query: 252 NGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSS 311
N +CL+WLD+Q+P SV++VSFGSGGT+SQ Q+ ELALGLELS+ KFLWV+R P+ +S+
Sbjct: 753 NESQCLSWLDEQKPNSVVFVSFGSGGTISQNQMNELALGLELSSQKFLWVVREPNDIASA 574
Query: 312 AGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLE 371
+ + + D L FLP GFLERT ++GF++++WAPQ++ILS ++GGF+THCGW STLE
Sbjct: 573 IYFDVSNSKKDPLSFLPKGFLERTNKQGFLVSNWAPQVEILSHKAIGGFVTHCGWFSTLE 394
Query: 372 SVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASV-NENGIVERVEVAKVIKYLMEGDE 430
VV+GVP++ WPLFAEQ+MNA +L++G+K+ +R ++ N +G+VE+VE+ V+K L+ DE
Sbjct: 393 CVVNGVPIVAWPLFAEQRMNATILADGIKIAIRPTIDNVSGVVEKVEIVNVLKRLIV-DE 217
Query: 431 GEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNL 469
G ++R MK LK+AA+NA+K DGSS T+SQ+ KW +
Sbjct: 216 GIEIRRRMKVLKDAAANAMKVDGSSIITMSQLVTKWTKM 100
>TC78509 similar to GP|19911193|dbj|BAB86923. glucosyltransferase-5 {Vigna
angularis}, partial (52%)
Length = 1655
Score = 157 bits (398), Expect(2) = 2e-66
Identities = 80/178 (44%), Positives = 115/178 (63%)
Frame = +2
Query: 290 GLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQI 349
GLE S +FLW++R+ S E +D L LP GFLERTKEKG V+ +WAPQ
Sbjct: 959 GLEKSEQRFLWIVRSDMESE--------ELSLDEL--LPEGFLERTKEKGMVVRNWAPQG 1108
Query: 350 QILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNE 409
IL +SVGGF+THCGWNS LE++ GVP+ITWPL+AEQKMN ++L + KV L + ++
Sbjct: 1109 SILRHSSVGGFVTHCGWNSVLEAICEGVPMITWPLYAEQKMNRLILVQEWKVALELNESK 1288
Query: 410 NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+G V E+ + +K LME ++G+++R + ++K +A A GSS + ++ WR
Sbjct: 1289 DGFVSENELGERVKELMESEKGKEVRETILKMKISAKEARGGGGSSLVDLKKLGDSWR 1462
Score = 113 bits (283), Expect(2) = 2e-66
Identities = 85/299 (28%), Positives = 144/299 (47%), Gaps = 16/299 (5%)
Frame = +3
Query: 7 IAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLG---------SPSNATKSILQT 57
I + P G HL+ +++ K ++ HP+F + I T S S S+
Sbjct: 90 IVLYPAFGSGHLMSMVELGKLILTHHPSFSIKILILTPPNQDTNTINVSTSQYISSVSNK 269
Query: 58 LPSNINHTFLPPVNPN-DLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDA 116
PS IN ++P ++ LP Q L S ++H L+S+AK L A+++D
Sbjct: 270 FPS-INFHYIPSISFTFTLPP----HLQTLELSPRSNHHVHHILQSIAKTSNLKAVMLDF 434
Query: 117 FSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRD--ILEPIKIPGCVP 174
+ + L + +Y Y+ S A+ L P + + +D + PI++PG
Sbjct: 435 LNYSASQVTNNLEIPTYFYYTSGASLLCLFLNFPTFHKNATIPIKDYNMHTPIELPGLPR 614
Query: 175 LHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE----EGGD 230
L D+ +D SS +Y+ L K L +DG++VN+F IE + A++ G
Sbjct: 615 LSKEDYPDEGKDPSSPSYQVLLQSAKSLRESDGIIVNTFDAIEKKAIKALRNGLCVPDGT 794
Query: 231 NPPVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELAL 289
P ++ +GP++ T + +D +G CL+WLD Q SV+ +SFGS G S+ QI ++A+
Sbjct: 795 TPLLFCIGPVVSTSCE--EDKSG--CLSWLDSQPGQSVVLLSFGSLGRFSKAQINQIAM 959
>TC79143 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (38%)
Length = 698
Score = 247 bits (630), Expect = 8e-66
Identities = 117/227 (51%), Positives = 160/227 (69%)
Frame = +3
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
M KT HIAV+ G+SH+ PI++FSKRLV HPNFHVTC IP+LGS +++KS L+T+P
Sbjct: 18 MAKTSHIAVISSPGFSHIAPIVEFSKRLVTNHPNFHVTCIIPSLGSLQDSSKSYLETVPP 197
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
NIN FLPP+N DLPQG I LT+T SLP +HQ LKS+ + PLVA++ D F+ E
Sbjct: 198 NINLVFLPPINKQDLPQGVYPGILIQLTVTRSLPSIHQALKSINSKAPLVAIIADNFAWE 377
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
L+ KE N LSY+YFP +A L++ + PKLDEE SC+Y+D+ EPIK+ GCVP++G D
Sbjct: 378 ALDFAKEFNSLSYVYFPCSAFVLSFYLHWPKLDEEVSCKYKDLQEPIKLQGCVPINGIDL 557
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEE 227
++ +DRS QAYK +L K + DG+L NSF +E + A++++
Sbjct: 558 PTVTKDRSGQAYKMYLQRAKDMCFVDGILFNSFFALESSAIKALEQK 698
>TC93597 similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13 {Vigna
angularis}, partial (32%)
Length = 660
Score = 189 bits (479), Expect(2) = 8e-66
Identities = 93/143 (65%), Positives = 111/143 (77%)
Frame = +1
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
MEKTVHIAVVPGVGY HL PILQFSK LVQLHP FHVTCFIP++ S +K+I+QTLPS
Sbjct: 49 MEKTVHIAVVPGVGYGHLFPILQFSKLLVQLHPYFHVTCFIPSIESLPTDSKTIIQTLPS 228
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
NIN TFLP V+ DLPQG + QI LT+ +SLP +HQ LKSL P VALVVD+ +++
Sbjct: 229 NINCTFLPSVSSKDLPQGIALVLQIQLTVIHSLPSIHQALKSLTLRTPFVALVVDSLAID 408
Query: 121 VLNIGKELNMLSYIYFPSAATTL 143
L+ KE NMLSY+YFPS+ T+L
Sbjct: 409 ALDFAKEFNMLSYVYFPSSVTSL 477
Score = 80.1 bits (196), Expect(2) = 8e-66
Identities = 35/54 (64%), Positives = 46/54 (84%)
Frame = +3
Query: 147 FYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSSQAYKHFLPFVK 200
FYL KL++ETSC+Y+D+LEPI+IPGCVP+HG+D + AQDRSSQ+YK L V+
Sbjct: 489 FYLLKLNKETSCQYKDLLEPIQIPGCVPIHGQDLVDQAQDRSSQSYKFLLERVE 650
>TC87866 similar to GP|19911195|dbj|BAB86924. glucosyltransferase-6 {Vigna
angularis}, partial (12%)
Length = 960
Score = 152 bits (384), Expect(2) = 3e-55
Identities = 91/247 (36%), Positives = 143/247 (57%), Gaps = 6/247 (2%)
Frame = +2
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
M K +IA+VP G SHL+P ++F+K+L+Q H F VT IPTL + + + +L TLP
Sbjct: 29 MNKKTYIAMVPSPGLSHLIPQVEFAKQLLQQHNEFIVTFIIPTLCPLTPSMQQVLNTLPP 208
Query: 61 NINHTFLPPVNPNDLPQ-GTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSV 119
NI LP V LP+ +Q+ L + +S+P+L + +KSL + LV+L+ FS
Sbjct: 209 NIEFIVLPQV--KHLPEINLEPATQMKLIVKHSIPFLQEEVKSLISKTNLVSLIFGLFST 382
Query: 120 EVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCE---YRDILEPIKIPGCV-PL 175
+ + K+ N+ SY+++ S A +L++ LP LD+ S E E + IPGCV P
Sbjct: 383 DAHEVAKQFNLSSYLFYASGALSLSFFLTLPNLDDSVSSEAEFLESAYETVNIPGCVIPF 562
Query: 176 HGRDFLSIAQ-DRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPV 234
H +D I +RS+ YK FL + L+ DG ++++F ++E + ++E+ + P V
Sbjct: 563 HIKDIPEIILCERSNVNYKIFLEVCQKLTLVDGFIISTFTDLEPDVIRVLQEK--EKPCV 736
Query: 235 YPVGPII 241
YPVGPII
Sbjct: 737 YPVGPII 757
Score = 81.3 bits (199), Expect(2) = 3e-55
Identities = 40/68 (58%), Positives = 48/68 (69%)
Frame = +3
Query: 239 PIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKF 298
P+ E+ D+ N + CL WL+ Q P SVLYV FGSGGT+S EQ+ ELA GLELS KF
Sbjct: 750 PLFINESNCEDNINSM-CLRWLENQPPNSVLYVCFGSGGTVSHEQLNELAFGLELSGKKF 926
Query: 299 LWVLRAPS 306
LWV+R PS
Sbjct: 927 LWVVRVPS 950
>TC83353 similar to GP|19911195|dbj|BAB86924. glucosyltransferase-6 {Vigna
angularis}, partial (5%)
Length = 789
Score = 200 bits (508), Expect = 1e-51
Identities = 111/245 (45%), Positives = 155/245 (62%), Gaps = 5/245 (2%)
Frame = +1
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNIN 63
T HI V +SH +++F KRL+Q+H +F +TC PT+ +P AT +L++LPS I+
Sbjct: 46 TSHIVVTSIPLFSHESSVIEFCKRLIQVHNHFQITCIFPTIDAPIPATLKLLESLPSTIH 225
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKE--IPLVALVVDAFSVEV 121
TFLPP+ DLPQ TM Q+ L +T S+P + L L P+VA+VVD F+ +
Sbjct: 226 CTFLPPIKKQDLPQEVTM--QLELGVTKSMPSFRESLSLLCSTSTTPVVAIVVDPFANQA 399
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF- 180
L I KE N+LS++YFP +A T + +LP LDE+ S EY D +EPI+IPGC P+ G+D
Sbjct: 400 LEIAKEFNILSFMYFPVSAMTTSLHLHLPILDEQVSGEYMDHVEPIEIPGCTPIRGQDLP 579
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAM--KEEGGDNPPVYPVG 238
+ +DRSS AY+ L K S ADGVL+NSF E+E + A+ KE+ + VY VG
Sbjct: 580 RTFFEDRSSIAYETILRQTKRFSLADGVLINSFSEMEESTVRALMEKEQSNNKQLVYLVG 759
Query: 239 PIIET 243
PII+T
Sbjct: 760 PIIQT 774
>CB892343 similar to GP|19911195|dbj glucosyltransferase-6 {Vigna angularis},
partial (5%)
Length = 811
Score = 198 bits (503), Expect = 4e-51
Identities = 112/261 (42%), Positives = 158/261 (59%), Gaps = 5/261 (1%)
Frame = +1
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
HIAVV +SH I++F KRL+ LH + H+TC T+ +P AT +L++LPS+IN T
Sbjct: 19 HIAVVSIPLFSHQSSIIEFCKRLIHLHHHIHITCIFSTIDAPIPATLKLLESLPSSINCT 198
Query: 66 FLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSL---AKEIPLVALVVDAFSVEVL 122
FLPP+N DLP+ +E I LT S+P + L SL + P+VALVVD ++ + L
Sbjct: 199 FLPPINKQDLPRDFVLE--IELTTAQSMPSFRKSLLSLCSSSTSSPVVALVVDPYASQAL 372
Query: 123 NIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF-L 181
I K+ N+LS++YFP +A T ++ Y P L E+ SCEY+D + I+ PGC+P+ +D
Sbjct: 373 EIAKD*NILSFVYFPLSAMTTSFNLYFPALHEQVSCEYKDHTDLIQFPGCLPICSQDLPP 552
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPP-VYPVGPI 240
DRSS Y FL K LS A G LVNSF ++E A++EE VY VG I
Sbjct: 553 EFFHDRSSVRYSFFLLHSKNLSLAHGFLVNSFSKMEASTGRALQEEHNKTTKLVYMVGTI 732
Query: 241 IETETKSGDDANGLECLAWLD 261
I++ + D++NG CL WL+
Sbjct: 733 IQSGSNCSDESNGSICLKWLE 795
>TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase-8 {Vigna
angularis}, partial (53%)
Length = 1787
Score = 181 bits (460), Expect = 4e-46
Identities = 143/486 (29%), Positives = 224/486 (45%), Gaps = 28/486 (5%)
Frame = +1
Query: 5 VHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINH 64
+HI P + H++P + ++ V VT L P + NI
Sbjct: 55 LHIIFFPFLANGHIIPCVDLAR--VFSSRGLKVTIVTTHLNVP--LISRTIGKAKINIKT 222
Query: 65 TFLPPVNPNDLPQGTTMESQIL-----LTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSV 119
P LP+G L + S L + L+ + ++ LV D F
Sbjct: 223 IKFPSPEETGLPEGCENSESALAPDKFIKFMKSTLLLREPLEHVLEQEKPDCLVADMFFP 402
Query: 120 EVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRD 179
+ + N+ ++ L C + + EP +P L G
Sbjct: 403 WSTDSAAKFNIPRIVFHGLGFFPL--CVLACTRQYKPQDKVSSYTEPFVVPN---LPGE- 564
Query: 180 FLSIAQDRSSQAYKHFLPFVKLLSSAD-------GVLVNSFLEIEMGPLSAMKEEGGDNP 232
+++ + + Q +H F KLL ++ GV+ N+F E+E + E G
Sbjct: 565 -ITLTKMQLPQLPQHDKVFTKLLEESNESEVKSFGVIANTFYELEPVYADHYRNELGRK- 738
Query: 233 PVYPVGPII----ETETKS--GDDAN--GLECLAWLDKQQPCSVLYVSFGSGGTLSQEQI 284
+ +GP+ +TE K+ G +A+ ECL WL ++P SV+YV FGS S Q+
Sbjct: 739 -AWHLGPVSLCNRDTEEKACRGREASIDEHECLKWLQSKEPNSVIYVCFGSMTVFSDAQL 915
Query: 285 VELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKE--KGFVI 342
E+A+GLE S F+WV+R SA+++ + L++LP GF ER + KG +I
Sbjct: 916 KEIAMGLEASEVPFIWVVRK-----------SAKSEGENLEWLPEGFEERIEGSGKGLII 1062
Query: 343 TSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVG 402
WAPQ+ IL SVGGF+THCGWNSTLE V G+P++TWP++ EQ NA LS+ +K+G
Sbjct: 1063RGWAPQVMILDHESVGGFVTHCGWNSTLEGVSAGLPMVTWPMYGEQFYNAKFLSDIVKIG 1242
Query: 403 LRASVNE------NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSST 456
+ V V++ + K ++ +M GDE E++R+ KE + A AV+ GSS
Sbjct: 1243VGVGVQTWIGMGGGEPVKKDVIEKAVRRIMVGDEAEEMRSRAKEFGKMARRAVEVGGSSY 1422
Query: 457 KTISQI 462
S +
Sbjct: 1423NDFSNL 1440
>TC78077 similar to GP|4115536|dbj|BAA36411.1 UDP-glycose:flavonoid
glycosyltransferase {Vigna mungo}, partial (66%)
Length = 865
Score = 179 bits (453), Expect = 3e-45
Identities = 89/200 (44%), Positives = 130/200 (64%)
Frame = +1
Query: 268 VLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFL 327
V+ +SFGS G S+ Q+ E+A+GLE S +FLWV+R+ S LS + +
Sbjct: 1 VVLLSFGSMGRFSRAQLNEIAIGLEKSEQRFLWVVRSEPDSDK----LSLD------ELF 150
Query: 328 PSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAE 387
P GFLERTK+KG V+ +WAPQ+ ILS NSVGGF+THCGWNS LE++ GVP+I WPLFAE
Sbjct: 151 PEGFLERTKDKGMVVRNWAPQVAILSHNSVGGFVTHCGWNSVLEAICEGVPMIAWPLFAE 330
Query: 388 QKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASN 447
Q++N ++L + +KV L+ + +EN V E+ + +K LME D G+ ++ + ++K +A
Sbjct: 331 QRLNRLVLVDEMKVALKVNQSENRFVSGTELGERVKELMESDRGKDIKERILKMKISAKE 510
Query: 448 AVKEDGSSTKTISQIALKWR 467
A GSS + ++ WR
Sbjct: 511 ARGGGGSSLVDLKKLGDSWR 570
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,136,308
Number of Sequences: 36976
Number of extensions: 227573
Number of successful extensions: 1500
Number of sequences better than 10.0: 162
Number of HSP's better than 10.0 without gapping: 1360
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1372
length of query: 470
length of database: 9,014,727
effective HSP length: 100
effective length of query: 370
effective length of database: 5,317,127
effective search space: 1967336990
effective search space used: 1967336990
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124966.3


