

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
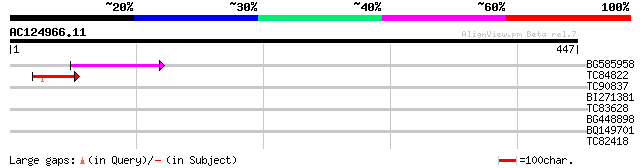
Query= AC124966.11 - phase: 0
(447 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 65 5e-11
TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsi... 54 1e-07
TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical... 37 0.015
BI271381 similar to GP|21397269|gb Unknown protein {Oryza sativa... 33 0.29
TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_18... 32 0.38
BG448898 similar to GP|19715622|gb| AT4g37080/C7A10_280 {Arabido... 28 5.5
BQ149701 similar to PIR|G86160|G86 protein F10O3.17 [imported] -... 28 9.4
TC82418 similar to PIR|C86204|C86204 hypothetical protein [impor... 28 9.4
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 65.1 bits (157), Expect = 5e-11
Identities = 36/75 (48%), Positives = 44/75 (58%), Gaps = 1/75 (1%)
Frame = +2
Query: 49 RKELEEGSGSRS-RKYLNRDHAAANQRLIDDYFANEPTYGDAMFRRRYRMQKHVFLRIVG 107
++E S RS R+ + R+ +RL +DYF+ P Y D FRRRYRM KHVFLRIV
Sbjct: 440 QEEFASSSRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVFLRIVE 619
Query: 108 DLSITDNYFTQRVDA 122
L D YF VDA
Sbjct: 620 ALGQHDEYFQLTVDA 664
>TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsis
thaliana}, partial (47%)
Length = 636
Score = 53.9 bits (128), Expect = 1e-07
Identities = 25/41 (60%), Positives = 34/41 (81%), Gaps = 4/41 (9%)
Frame = +1
Query: 19 RLSMDP----FDIEAYKQKRDIEDTYIVNRFIQRRKELEEG 55
++SMDP FDIEAYK+K +IE+ YI+NRF +RRK++EEG
Sbjct: 31 KISMDPSKFRFDIEAYKRKSEIEERYILNRFRERRKQIEEG 153
>TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical
protein~similar to Oryza sativa chromosome10
OSJNBa0079H13.17, partial (7%)
Length = 596
Score = 37.0 bits (84), Expect = 0.015
Identities = 30/97 (30%), Positives = 45/97 (45%), Gaps = 11/97 (11%)
Frame = +1
Query: 295 YYLADGIYPS---YPTFVKSIRLPQSE--------PDKLFAKHQESCRKDIERAFGVLQA 343
YYL D YP Y IR QS+ + F + S R IER+FGVL+
Sbjct: 10 YYLVDKGYPDKEGYMVPYPRIRYHQSQFEHEPPTNAQEAFNRAHSSLRSCIERSFGVLKK 189
Query: 344 RFKIIREPARLWDIADLGIIMRSCIILHNMIVEDERD 380
R+KI+ + + + +I+ + LHN I + +D
Sbjct: 190 RWKILNKMPQFSVKTQIDVII-AAFALHNYIRINSQD 297
>BI271381 similar to GP|21397269|gb Unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 407
Score = 32.7 bits (73), Expect = 0.29
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = +2
Query: 319 PDKLFAKHQESCRKDIERAFGVLQARFKIIR 349
P++LF S R IER FGVL+ RF I++
Sbjct: 134 PEELFNYRHSSLRMTIERCFGVLKNRFPILK 226
>TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_180
{Arabidopsis thaliana}, partial (23%)
Length = 773
Score = 32.3 bits (72), Expect = 0.38
Identities = 25/106 (23%), Positives = 47/106 (43%)
Frame = +2
Query: 82 NEPTYGDAMFRRRYRMQKHVFLRIVGDLSITDNYFTQRVDAANKEGISPLAKCTTAMRML 141
N + D FRR +RM K F I +L D+ T++ + ++ I + + L
Sbjct: 431 NREDFPDDEFRRCFRMSKQTFNMICNEL---DSSVTKK-NTTLRDAIPVRQRVAVCIYRL 598
Query: 142 AYGVAADAVDEYIKIGGTTALECLRRFCKGIIRLYEEVYLRAPNQD 187
A G V + +G +T + + C I + + +LR P+++
Sbjct: 599 ATGEPLRLVSKKFGLGISTCHKLVLEVCAAIKSVLMQKFLRWPDEE 736
>BG448898 similar to GP|19715622|gb| AT4g37080/C7A10_280 {Arabidopsis
thaliana}, partial (14%)
Length = 552
Score = 28.5 bits (62), Expect = 5.5
Identities = 23/73 (31%), Positives = 35/73 (47%), Gaps = 6/73 (8%)
Frame = +2
Query: 316 QSEPDKLFAK--HQESCRKDIERAF----GVLQARFKIIREPARLWDIADLGIIMRSCII 369
Q + DKL K H+E+ RK +ERAF G L R P L +A++ ++ +
Sbjct: 200 QQDVDKLKKKLKHEENIRKALERAFNRPLGAL-PRLPPYLPPYILALLAEVAVLEEEIVW 376
Query: 370 LHNMIVEDERDTY 382
L +V +D Y
Sbjct: 377 LEEKVVHFRQDLY 415
>BQ149701 similar to PIR|G86160|G86 protein F10O3.17 [imported] - Arabidopsis
thaliana, partial (26%)
Length = 649
Score = 27.7 bits (60), Expect = 9.4
Identities = 28/95 (29%), Positives = 39/95 (40%), Gaps = 9/95 (9%)
Frame = +1
Query: 329 SCRKDIERAFGVL--QARFKIIREPARLWD-------IADLGIIMRSCIILHNMIVEDER 379
SCR D+ER G+ QA + I P L+ I D I+R I N+ EDE
Sbjct: 256 SCRSDLERRIGLQLDQAILEDILIPTSLYQNNNHSSTIYDTDSIVRIFSIFLNLDEEDED 435
Query: 380 DTYAQRWTDFEQSEGSGSSTAQPYSTEVLPTFANH 414
D+ + T+ S S Q +V N+
Sbjct: 436 DSRMRDETEMVYDFDSPGSPKQSSILKVSKLLDNY 540
>TC82418 similar to PIR|C86204|C86204 hypothetical protein [imported] -
Arabidopsis thaliana, partial (83%)
Length = 731
Score = 27.7 bits (60), Expect = 9.4
Identities = 13/53 (24%), Positives = 25/53 (46%)
Frame = +2
Query: 275 EQGKAPRVNFFVNQRPYNMAYYLADGIYPSYPTFVKSIRLPQSEPDKLFAKHQ 327
++ AP F+ PY + + ++ YP F K +RL +++P F +
Sbjct: 452 QEAAAPNNILFIENLPYETTGRMLEMLFEQYPGF-KEVRLIEAKPGIAFVNFE 607
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,104,864
Number of Sequences: 36976
Number of extensions: 190735
Number of successful extensions: 875
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 873
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 875
length of query: 447
length of database: 9,014,727
effective HSP length: 99
effective length of query: 348
effective length of database: 5,354,103
effective search space: 1863227844
effective search space used: 1863227844
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124966.11


