

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.11 - phase: 0 /pseudo
(168 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
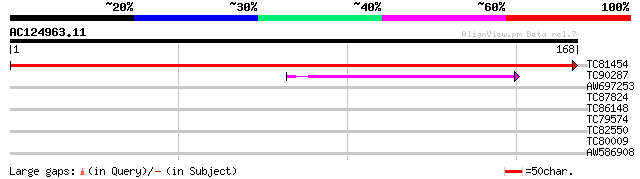
Score E
Sequences producing significant alignments: (bits) Value
TC81454 similar to GP|4371292|gb|AAD18150.1| unknown protein {Ar... 322 3e-89
TC90287 similar to GP|21592380|gb|AAM64331.1 glycoprotein-specif... 39 8e-04
AW697253 similar to GP|12597289|gb| FasL isoform {Homo sapiens},... 30 0.35
TC87824 homologue to GP|16612249|gb|AAL27496.1 AT4g35230/F23E12_... 30 0.35
TC86148 homologue to PIR|T05747|T05747 histone H2A.M4I22.40 - Ar... 27 3.9
TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13... 27 5.1
TC82550 similar to GP|3832528|gb|AAC70787.1| unknown {Glycine ma... 26 8.6
TC80009 weakly similar to PIR|T03747|T03747 glucosyltransferase ... 26 8.6
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 26 8.6
>TC81454 similar to GP|4371292|gb|AAD18150.1| unknown protein {Arabidopsis
thaliana}, partial (51%)
Length = 1003
Score = 322 bits (826), Expect = 3e-89
Identities = 168/168 (100%), Positives = 168/168 (100%)
Frame = +2
Query: 1 MVTKVGQLQHLESIAKKQQQQQQQNCM*NHYLN*SLKDF*S**HQQAQNYLITMFF*EGW 60
MVTKVGQLQHLESIAKKQQQQQQQNCM*NHYLN*SLKDF*S**HQQAQNYLITMFF*EGW
Sbjct: 446 MVTKVGQLQHLESIAKKQQQQQQQNCM*NHYLN*SLKDF*S**HQQAQNYLITMFF*EGW 625
Query: 61 QIL*SLLINHCFGLLSKRKPNQLNCQRY*GKLVSCIDMLFSVKSSWT*KLN*IIKEI*LL 120
QIL*SLLINHCFGLLSKRKPNQLNCQRY*GKLVSCIDMLFSVKSSWT*KLN*IIKEI*LL
Sbjct: 626 QIL*SLLINHCFGLLSKRKPNQLNCQRY*GKLVSCIDMLFSVKSSWT*KLN*IIKEI*LL 805
Query: 121 DTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQRHYYLQTEK 168
DTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQRHYYLQTEK
Sbjct: 806 DTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQRHYYLQTEK 949
>TC90287 similar to GP|21592380|gb|AAM64331.1 glycoprotein-specific
UDP-glucuronyltransferase-like protein {Arabidopsis
thaliana}, partial (69%)
Length = 804
Score = 39.3 bits (90), Expect = 8e-04
Identities = 27/69 (39%), Positives = 38/69 (54%)
Frame = +3
Query: 83 LNCQRY*GKLVSCIDMLFSVKSSWT*KLN*IIKEI*LLDTLNIIDLVGLFTLLAFLMFMI 142
LNC *GK V CI + ++ *++ I+EI LNII+L+GLFTLL +M ++
Sbjct: 39 LNC---*GKRV*CIGIWCVLRIQRM*RIEEFIREIKRWSILNIINLMGLFTLLMMIMCIL 209
Query: 143 FNSSNSLEI 151
EI
Sbjct: 210 LTCFRQSEI 236
>AW697253 similar to GP|12597289|gb| FasL isoform {Homo sapiens}, partial
(16%)
Length = 761
Score = 30.4 bits (67), Expect = 0.35
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -3
Query: 119 LLDTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQRHYY 163
++ T ++ L+GL LL L+ +I SS+S + G HY+
Sbjct: 240 IVATTEVLKLIGLLLLLFLLLLIILTSSSSSSSSQQRGFTTTHYF 106
>TC87824 homologue to GP|16612249|gb|AAL27496.1 AT4g35230/F23E12_210
{Arabidopsis thaliana}, partial (36%)
Length = 895
Score = 30.4 bits (67), Expect = 0.35
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -3
Query: 119 LLDTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILRSLGHGQRHYY 163
++ T ++ L+GL LL L+ +I SS+S + G HY+
Sbjct: 248 IVATTEVLKLIGLLLLLFLLLLIILTSSSSSSSSQQRGFTTTHYF 114
>TC86148 homologue to PIR|T05747|T05747 histone H2A.M4I22.40 - Arabidopsis
thaliana, partial (96%)
Length = 696
Score = 26.9 bits (58), Expect = 3.9
Identities = 15/33 (45%), Positives = 21/33 (63%)
Frame = +3
Query: 121 DTLNIIDLVGLFTLLAFLMFMIFNSSNSLEILR 153
+TLN++ LV LFT L FL + N LE+L+
Sbjct: 207 NTLNVLVLVLLFT*LLFLNISLLRFWNWLEMLQ 305
>TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana, partial (38%)
Length = 1412
Score = 26.6 bits (57), Expect = 5.1
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = +2
Query: 147 NSLEILRSLGHGQRHYYLQ 165
N LEI+R HGQR +LQ
Sbjct: 800 NLLEIIREFDHGQRRAFLQ 856
>TC82550 similar to GP|3832528|gb|AAC70787.1| unknown {Glycine max},
partial (32%)
Length = 875
Score = 25.8 bits (55), Expect = 8.6
Identities = 10/16 (62%), Positives = 14/16 (87%)
Frame = +1
Query: 10 HLESIAKKQQQQQQQN 25
HL S+ ++QQQQQQQ+
Sbjct: 43 HLPSLQQRQQQQQQQH 90
>TC80009 weakly similar to PIR|T03747|T03747 glucosyltransferase IS5a (EC
2.4.1.-) salicylate-induced - common tobacco, partial
(19%)
Length = 1118
Score = 25.8 bits (55), Expect = 8.6
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = +1
Query: 50 YLITMFF*EGWQIL*SLLINHCFGLLSKRKPNQLNCQR 87
YLI + W++ * L+ H +GL KRK ++ ++
Sbjct: 946 YLINNYM--RWRVQ*KHLVTHSYGLFLKRKGKKMRVKK 1053
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 25.8 bits (55), Expect = 8.6
Identities = 11/18 (61%), Positives = 15/18 (83%)
Frame = +2
Query: 7 QLQHLESIAKKQQQQQQQ 24
QLQ L+ + ++QQQQQQQ
Sbjct: 575 QLQLLQKLQQQQQQQQQQ 628
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.362 0.162 0.552
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,245,170
Number of Sequences: 36976
Number of extensions: 73270
Number of successful extensions: 913
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 792
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 872
length of query: 168
length of database: 9,014,727
effective HSP length: 89
effective length of query: 79
effective length of database: 5,723,863
effective search space: 452185177
effective search space used: 452185177
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC124963.11


