

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
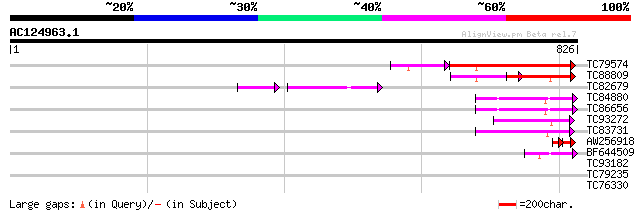
Score E
Sequences producing significant alignments: (bits) Value
TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13... 252 5e-67
TC88809 similar to PIR|T05688|T05688 hypothetical protein F20M13... 108 1e-23
TC82679 similar to PIR|T48309|T48309 hypothetical protein F9G14.... 101 1e-21
TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase h... 88 1e-17
TC86656 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein l... 84 3e-16
TC93272 similar to GP|20260606|gb|AAM13201.1 unknown protein {Ar... 73 4e-13
TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.2... 57 3e-08
AW256918 homologue to PIR|T48309|T48 hypothetical protein F9G14.... 38 3e-07
BF644509 similar to GP|7108521|gb ubiquitin-protein ligase 1 {Ar... 45 1e-04
TC93182 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein l... 41 0.002
TC79235 weakly similar to GP|7297741|gb|AAF52992.1| CG17108 gene... 30 2.9
TC76330 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 29 8.4
>TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana, partial (38%)
Length = 1412
Score = 252 bits (643), Expect = 5e-67
Identities = 127/206 (61%), Positives = 153/206 (73%), Gaps = 22/206 (10%)
Frame = +2
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF---------------------- 678
+FR+S+IEDLCLDF+LPGY L S + + N+ +
Sbjct: 452 SFRDSRIEDLCLDFTLPGYPDIVLASGSDHTMVNMRNLEDYVSLIVDATVKSGISRQVEA 631
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFNQVFPIE+L++F EEELE ILCGE +SW N+L H KFDHGYTA SPPI+NLLE
Sbjct: 632 FKSGFNQVFPIENLQIFYEEELERILCGEDDSWAINELADHIKFDHGYTASSPPIVNLLE 811
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
I++EF++ +RRAF+QFVT +PRLPPGGLASL+PKLT+V+K N DTDLPSVMTCANYL
Sbjct: 812 IIREFDHGQRRAFLQFVTGTPRLPPGGLASLNPKLTIVRKHCSNQADTDLPSVMTCANYL 991
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKERMK+KLLYAITEG+G F
Sbjct: 992 KLPPYSSKERMKEKLLYAITEGQGSF 1069
Score = 81.6 bits (200), Expect = 1e-15
Identities = 51/103 (49%), Positives = 59/103 (56%), Gaps = 18/103 (17%)
Frame = +1
Query: 556 TDFGILYLGL*GVSKSRFRPVERG------------------YSVWSLSSPMVENAR*I* 597
T+FGILYLG+ GV++ F VERG +S+W+LSS MV N R I
Sbjct: 7 TNFGILYLGVSGVTEVWFGHVERGCFFIHHKDKLAG*GNRNPFSLWTLSSSMVVNTRYIW 186
Query: 598 WPKDF*STKEICPFRSSCC*SYSRW*APRSPYLQSLS*TYLWK 640
W *S K I PFRSSCC*S SRW RSP S+ *TY WK
Sbjct: 187 WHTVL*SNK*IFPFRSSCC*STSRWKGLRSPLF*SIL*TYSWK 315
>TC88809 similar to PIR|T05688|T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana, partial (40%)
Length = 1406
Score = 108 bits (269), Expect = 1e-23
Identities = 58/113 (51%), Positives = 74/113 (65%), Gaps = 13/113 (11%)
Frame = +3
Query: 725 GYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT 784
G R+P L++ + + RAF QFVT +P+LPPGGLA L+PKLT+V+K+S
Sbjct: 747 GTLPRAPQ**IYLKLWESSHRSSSRAFCQFVTGAPKLPPGGLAVLNPKLTIVRKLSSTAA 926
Query: 785 DT-------------DLPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
+T DLPSVMTCANYLKLPPYS+KE M +KL+YAI EG+G F
Sbjct: 927 NTTSNGNGPSETADDDLPSVMTCANYLKLPPYSTKEIMHKKLMYAINEGQGSF 1085
Score = 99.0 bits (245), Expect = 7e-21
Identities = 54/127 (42%), Positives = 70/127 (54%), Gaps = 20/127 (15%)
Frame = +2
Query: 642 FRNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLV 681
+R + I DLCLDF+LPGY + L P I + NL + F
Sbjct: 437 YRGAPIADLCLDFTLPGYPEYTLKPGDEIVDLNNLEDYISMVVDATVKTGITRQLEAFRA 616
Query: 682 GFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQ 741
GFNQVF I L++F+ EL+ +LCG W L H KFDHGYTA+SP I+NLLEI+
Sbjct: 617 GFNQVFDISSLQIFTPHELDYLLCGRRELWKTETLADHIKFDHGYTAKSPAIVNLLEIMG 796
Query: 742 EFNNEER 748
EF E++
Sbjct: 797 EFTPEQQ 817
>TC82679 similar to PIR|T48309|T48309 hypothetical protein F9G14.190 -
Arabidopsis thaliana, partial (6%)
Length = 958
Score = 101 bits (252), Expect = 1e-21
Identities = 66/138 (47%), Positives = 83/138 (59%)
Frame = +1
Query: 405 KSSCLFCGGS*NPE*PKITTTFF*TG*VFEQEVDREVGTTNGGSFGIVHWSYALLVLSID 464
K+ C S* *PK ++F T *V EQ VDRE G T+ G +G +HW YA LV S++
Sbjct: 514 KNVCFC*R*S**SR*PKNNSSFCPTK*VREQ*VDREAGATDEGLYGSLHWQYAFLV*SVN 693
Query: 465 DLVSLPIQLCDKMQVL*IGDIWPATEPSSRV**QP*NNERQKIEPQWIAAQEMSSFSRPN 524
++S PIQL KMQVL G IW AT + * *N++ Q EP IA++E+ S RP+
Sbjct: 694 GILSFPIQL*SKMQVLQTGSIWSATNTT----**L*NSK*QATEPWCIASKEVFSLPRPD 861
Query: 525 FGVCCTNDEPECLSKSRS 542
GVC TNDE C S S S
Sbjct: 862 LGVCYTNDETTCQS*SGS 915
Score = 43.1 bits (100), Expect = 4e-04
Identities = 28/61 (45%), Positives = 33/61 (53%)
Frame = +3
Query: 332 TCFLS*WTRIGS*IVAVSDTSMPNAKARQNFFCC*TVESSACIML*KGCRI*HHNPSRIC 391
T FL * T GS*I ++S S PN K + + C VES ACI L* C * HN +
Sbjct: 144 TYFLP*RTTSGS*IDSISGNSWPNNKTKW*WC*CKIVESRACINL*SSCET*GHNAFGLS 323
Query: 392 F 392
F
Sbjct: 324 F 326
>TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase homolog
F17O7.15 - Arabidopsis thaliana, partial (17%)
Length = 801
Score = 88.2 bits (217), Expect = 1e-17
Identities = 55/152 (36%), Positives = 90/152 (59%), Gaps = 4/152 (2%)
Frame = +1
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GF ++ P E + +F+++ELEL++ G P+ +DL + ++ GY+A SP I E
Sbjct: 148 FLEGFGEIIPKELISIFNDKELELLISGLPDI-DLDDLRANTEYS-GYSAASPVIQWFWE 321
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI----SYNHTDTDLPSVMTC 794
++Q F+ E++ +QFVT + ++P G ++L ++ QK +Y +D LPS TC
Sbjct: 322 VVQGFSKEDKARLLQFVTGTSKVPLEGFSALQG-ISGAQKFQIHKAYGSSD-HLPSAHTC 495
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SK+ ++++LL AI E F F
Sbjct: 496 FNQLDLPEYPSKQHLEERLLLAIHEANEGFGF 591
>TC86656 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein ligase 1
{Arabidopsis thaliana}, partial (12%)
Length = 1939
Score = 83.6 bits (205), Expect = 3e-16
Identities = 54/152 (35%), Positives = 89/152 (58%), Gaps = 4/152 (2%)
Frame = +1
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GF+++ P E + +F+++ELEL++ G P+ +DL + ++ GY+A SP I E
Sbjct: 1318 FLEGFSELIPRELISIFNDKELELLISGLPDI-DLDDLRANTEYS-GYSAASPVIQWFWE 1491
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI----SYNHTDTDLPSVMTC 794
++Q + E++ +QFVT + ++P G ++L ++ QK +Y D LPS TC
Sbjct: 1492 VVQGLSKEDKARLLQFVTGTSKVPLEGFSALQG-ISGSQKFQIHKAYGSPD-HLPSAHTC 1665
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SK+ ++++LL AI E F F
Sbjct: 1666 FNQLDLPEYPSKQHLEERLLLAIHEASEGFGF 1761
>TC93272 similar to GP|20260606|gb|AAM13201.1 unknown protein {Arabidopsis
thaliana}, partial (12%)
Length = 652
Score = 73.2 bits (178), Expect = 4e-13
Identities = 45/120 (37%), Positives = 65/120 (53%), Gaps = 3/120 (2%)
Frame = +3
Query: 706 GEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGG 765
G +S +DL +H + Y + I E+L+ F+ E ++ F++FVT R P G
Sbjct: 6 GSLDSLDVDDLREHTNYAGTYHSEHDVIEMFWEVLKGFSMENKKKFLKFVTGCSRGPLLG 185
Query: 766 LASLDPKLTVVQKISYNHTDTDL---PSVMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
L+P L +Q+ N T+ L P+ TC N LKLPPY SK++M+ KLLYAI G
Sbjct: 186 FRYLEP-LFCIQRAGGNATEDALDRLPTAATCMNLLKLPPYRSKDQMESKLLYAINSDAG 362
>TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.20 -
Arabidopsis thaliana, partial (27%)
Length = 954
Score = 57.0 bits (136), Expect = 3e-08
Identities = 42/153 (27%), Positives = 64/153 (41%), Gaps = 9/153 (5%)
Frame = +2
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F G + L++F+ E +L G +D + ++ GY S I E
Sbjct: 479 FYRGLTDLISPSWLKLFNASEFNQLLSGGNYDIDIDDFKSNTRYTGGYNEGSRTIKIFWE 658
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYN---------HTDTDLP 789
+++ F +ER ++FVT R P G L P T + K++ + L
Sbjct: 659 VIKGFEPKERCMLLKFVTSCSRGPLLGFKYLQPPFT-IHKVACDVPLWATIGGQDAERLS 835
Query: 790 SVMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
S TC N LKLP Y ++ K LYAI+ G
Sbjct: 836 SASTCYNTLKLPTYKRPGTLRAKALYAISSNAG 934
>AW256918 homologue to PIR|T48309|T48 hypothetical protein F9G14.190 -
Arabidopsis thaliana, partial (2%)
Length = 622
Score = 38.1 bits (87), Expect(2) = 3e-07
Identities = 16/16 (100%), Positives = 16/16 (100%)
Frame = -3
Query: 791 VMTCANYLKLPPYSSK 806
VMTCANYLKLPPYSSK
Sbjct: 620 VMTCANYLKLPPYSSK 573
Score = 35.0 bits (79), Expect(2) = 3e-07
Identities = 15/19 (78%), Positives = 18/19 (93%)
Frame = -1
Query: 806 KERMKQKLLYAITEGRGCF 824
+ERMK+KLLYAITEG+G F
Sbjct: 463 QERMKEKLLYAITEGQGSF 407
>BF644509 similar to GP|7108521|gb ubiquitin-protein ligase 1 {Arabidopsis
thaliana}, partial (3%)
Length = 554
Score = 44.7 bits (104), Expect = 1e-04
Identities = 30/80 (37%), Positives = 44/80 (54%), Gaps = 4/80 (5%)
Frame = +1
Query: 751 FVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTCANYLKLPPYSSK 806
++QF T + ++P G +L P+ + K +Y D LPS TC N L LP Y+SK
Sbjct: 127 YLQFXTGTSKVPLEGFKALQGISGPQRFQIHK-AYGAPDR-LPSAHTCFNQLDLPEYTSK 300
Query: 807 ERMKQKLLYAITEGRGCFLF 826
E+++ +LL AI E F F
Sbjct: 301 EQLQDRLLLAIHEASEGFGF 360
>TC93182 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein ligase 1
{Arabidopsis thaliana}, partial (1%)
Length = 748
Score = 40.8 bits (94), Expect = 0.002
Identities = 20/39 (51%), Positives = 26/39 (66%)
Frame = +3
Query: 788 LPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
LPS TC+N L LP Y+SKE+++ +LL AI E F F
Sbjct: 48 LPSAHTCSNQLDLPEYTSKEQLQDRLLLAIHEASEGFGF 164
>TC79235 weakly similar to GP|7297741|gb|AAF52992.1| CG17108 gene product
{Drosophila melanogaster}, partial (21%)
Length = 648
Score = 30.4 bits (67), Expect = 2.9
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = -3
Query: 38 HHLGKHLHD*ISVKTLIQELSVHVMHFLMAYLLH 71
HHL +H + + TL +LS ++H M LLH
Sbjct: 391 HHLFSTIHPLLQILTLHHQLSTTIIHHFMLILLH 290
>TC76330 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (72%)
Length = 1082
Score = 28.9 bits (63), Expect = 8.4
Identities = 17/52 (32%), Positives = 23/52 (43%), Gaps = 2/52 (3%)
Frame = +1
Query: 34 ISHFHHLGKHLHD*ISVKTLIQELSVHVMHFL--MAYLLHLQVFEFANLTKI 83
ISH HL H H + + + LS H +H ++ LLHL T I
Sbjct: 331 ISHLPHLSTHHHHQFTNISHLPHLSTHHLHLFTSISLLLHLSTHHHHQFTNI 486
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,449,576
Number of Sequences: 36976
Number of extensions: 469003
Number of successful extensions: 5065
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1558
Number of HSP's successfully gapped in prelim test: 235
Number of HSP's that attempted gapping in prelim test: 3430
Number of HSP's gapped (non-prelim): 1940
length of query: 826
length of database: 9,014,727
effective HSP length: 104
effective length of query: 722
effective length of database: 5,169,223
effective search space: 3732179006
effective search space used: 3732179006
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC124963.1


