

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
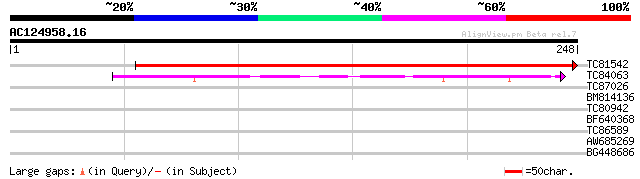
Score E
Sequences producing significant alignments: (bits) Value
TC81542 similar to GP|13877685|gb|AAK43920.1 Unknown protein {Ar... 383 e-107
TC84063 similar to GP|7339697|dbj|BAA92902.1 hypothetical protei... 63 9e-11
TC87026 weakly similar to GP|21592576|gb|AAM64525.1 SOUL-like pr... 39 0.001
BM814136 similar to PIR|G96841|G9684 hypothetical protein F23A5.... 28 3.3
TC80942 GP|21112489|gb|AAM40721.1 conserved hypothetical protein... 28 4.3
BF640368 similar to GP|3668092|gb unknown protein {Arabidopsis t... 27 7.3
TC86589 similar to GP|7576200|emb|CAB87861.1 protein synthesis i... 27 7.3
AW685269 similar to GP|13899097|gb| Unknown protein {Arabidopsis... 27 7.3
BG448686 similar to GP|10177219|dbj contains similarity to AP2 d... 27 7.3
>TC81542 similar to GP|13877685|gb|AAK43920.1 Unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 753
Score = 383 bits (984), Expect = e-107
Identities = 192/193 (99%), Positives = 193/193 (99%)
Frame = +3
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
VETPKYEVTKTTQDYEIR+YAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK
Sbjct: 3 VETPKYEVTKTTQDYEIRMYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 182
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD
Sbjct: 183 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 362
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL
Sbjct: 363 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 542
Query: 236 PMFRTNEVMIPIE 248
PMFRTNEVMIPIE
Sbjct: 543 PMFRTNEVMIPIE 581
>TC84063 similar to GP|7339697|dbj|BAA92902.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (42%)
Length = 599
Score = 63.2 bits (152), Expect = 9e-11
Identities = 62/205 (30%), Positives = 89/205 (43%), Gaps = 7/205 (3%)
Frame = +2
Query: 46 IMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSV-AAEVTYDPSQFKGNKDGGFMVLANY 104
IM M+ E+P+Y V T D+EIR+Y SV + + F+ GF L +
Sbjct: 47 IMAMIRNTTATESPQYTVVHTESDFEIRLYRTSVWMSAPAVNLISFEKATRNGFHRLFQF 226
Query: 105 IGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSY 164
I N +I MT PV+T T P +G + F LP+ +
Sbjct: 227 I----QGANLNFSRIPMTTPVLT--------TTVPGTGPLESQG-----YYVSFYLPTKF 355
Query: 165 EKAEEAPKPTDERVVIREEGERKY--GVVKFSGVASDEVVKEKVEKLRLSLERDG----F 218
++ P P + I+ G + V FSG A DE V ++ EKL LSL R G
Sbjct: 356 QENPPLPLP---ELNIKSYGFESHCVAVRGFSGFAKDEKVVKEAEKLDLSLSRWGGESEV 526
Query: 219 KVIGDFLLGRYNPPWTLPMFRTNEV 243
K G + + +YN P+ + R NEV
Sbjct: 527 KRKGGYSIAQYNGPFRIAK-RRNEV 598
>TC87026 weakly similar to GP|21592576|gb|AAM64525.1 SOUL-like protein
{Arabidopsis thaliana}, partial (64%)
Length = 794
Score = 39.3 bits (90), Expect = 0.001
Identities = 41/160 (25%), Positives = 66/160 (40%), Gaps = 2/160 (1%)
Frame = +1
Query: 56 VETPKYEVTKTTQDYEIRIYAPSV--AAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+E P Y+V + YEIR+Y SV + D S + + GF+ L +YI N Q
Sbjct: 229 IECPNYDVIEAGNGYEIRLYNSSVWISNSPIQDISLVEATRT-GFLRLFDYIQGKNNYQ- 402
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+KI MTAPV+ +E + P S F++ K +A P
Sbjct: 403 ---QKIEMTAPVL----SEVLPSDGPFCESS-------------FVVSFYVPKVNQANPP 522
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSL 213
+ + ++ V +F G D + E+ L+ S+
Sbjct: 523 PAKGLHVQRWKTVYAAVKQFGGFVKDTNIGEEAAALKDSI 642
>BM814136 similar to PIR|G96841|G9684 hypothetical protein F23A5.25
[imported] - Arabidopsis thaliana, partial (46%)
Length = 576
Score = 28.1 bits (61), Expect = 3.3
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +1
Query: 7 IQISYPLLYKGLKATSTTHKNLLLPTSFNSHFTLSHSRSI 46
I+I++ L K + +H LLLP F HF HS S+
Sbjct: 169 IKITWKKLIK--QGAKDSHPVLLLPNDFEKHFIKEHSFSV 282
>TC80942 GP|21112489|gb|AAM40721.1 conserved hypothetical protein
{Xanthomonas campestris pv. campestris str. ATCC 33913},
partial (9%)
Length = 482
Score = 27.7 bits (60), Expect = 4.3
Identities = 37/139 (26%), Positives = 51/139 (36%), Gaps = 5/139 (3%)
Frame = -2
Query: 59 PKYEVTKTTQDYEIRIYAPS-----VAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
P YEV KT +YE YAP+ +EV + P+ K + +
Sbjct: 361 PNYEVPKTKTNYE---YAPAPKIEKPKSEVVHTPNPTKPDYE---------------VPK 236
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+K + P I K +E A+ P TKS E + K Y A + KP
Sbjct: 235 SKTDYGYAPTPKIEKSKSE--AVYTPNPTKSDYEVPKTK-------TDYEYAPAPKIEKP 83
Query: 174 TDERVVIREEGERKYGVVK 192
E V R + Y V K
Sbjct: 82 KSEVVYTRNPTKPDYEVPK 26
>BF640368 similar to GP|3668092|gb unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 630
Score = 26.9 bits (58), Expect = 7.3
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -3
Query: 7 IQISYPLLYKGLKATSTTHKNLLLP 31
IQI +P+LY GL A+ T+ L+P
Sbjct: 142 IQIQWPILYVGLLASQTSAVKKLVP 68
>TC86589 similar to GP|7576200|emb|CAB87861.1 protein synthesis initiation
factor-like {Arabidopsis thaliana}, partial (28%)
Length = 2539
Score = 26.9 bits (58), Expect = 7.3
Identities = 23/101 (22%), Positives = 46/101 (44%), Gaps = 1/101 (0%)
Frame = +3
Query: 106 GALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEG-ERNKMVTMQFILPSSY 164
G LGNP T + G A ++ P+ + ++ G +RN + ++ +++
Sbjct: 69 GVLGNPH---------TPTALPYGGA---ILSGPMQSMVNQSGVQRNSPDSERWQRAANF 212
Query: 165 EKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEK 205
++ P P+ +V + E+KY V K V +E K++
Sbjct: 213 QQRGLIPSPSQSPLVTMHKAEKKYEVGK---VTDEEQAKQR 326
>AW685269 similar to GP|13899097|gb| Unknown protein {Arabidopsis thaliana},
partial (36%)
Length = 634
Score = 26.9 bits (58), Expect = 7.3
Identities = 16/45 (35%), Positives = 24/45 (52%)
Frame = -3
Query: 177 RVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVI 221
RV+++ E E K GV+ F + +KE VE+ L FK+I
Sbjct: 398 RVIVKREIEVKIGVIFFLSDLNFVELKENVERTSLQFFFFFFKII 264
>BG448686 similar to GP|10177219|dbj contains similarity to AP2 domain
transcription factor~gene_id:MPF21.9 {Arabidopsis
thaliana}, partial (31%)
Length = 652
Score = 26.9 bits (58), Expect = 7.3
Identities = 20/74 (27%), Positives = 31/74 (41%), Gaps = 2/74 (2%)
Frame = +1
Query: 60 KYEVTKTTQDYEIRIYAPSVAAEVTYDPS--QFKGNKDGGFMVLANYIGALGNPQNTKPE 117
K+E+ Q+Y + P+ AA YD + +F+GN+ L P+N
Sbjct: 277 KFEIHTKQQEYGLEHLKPAEAAARAYDEAALRFRGNR-----------AKLNFPENVPAS 423
Query: 118 KIAMTAPVITKGSA 131
T PV T S+
Sbjct: 424 ATRPTFPVNTSTSS 465
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,162,241
Number of Sequences: 36976
Number of extensions: 73231
Number of successful extensions: 324
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 320
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 320
length of query: 248
length of database: 9,014,727
effective HSP length: 94
effective length of query: 154
effective length of database: 5,538,983
effective search space: 853003382
effective search space used: 853003382
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC124958.16


